

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
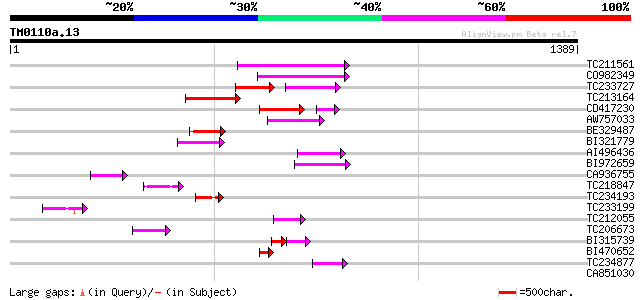
Query= TM0110a.13
(1389 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 209 5e-54
CO982349 148 1e-35
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 92 3e-29
TC213164 110 4e-24
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 84 1e-20
AW757033 93 7e-19
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 92 1e-18
BI321779 90 8e-18
AI496436 85 2e-16
BI972659 85 3e-16
CA936755 64 6e-10
TC218847 60 5e-09
TC234193 58 3e-08
TC233199 50 9e-06
TC212055 49 2e-05
TC206673 49 2e-05
BI315739 37 2e-05
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 45 2e-04
TC234877 43 8e-04
CA851030 43 0.001
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 209 bits (533), Expect = 5e-54
Identities = 105/275 (38%), Positives = 163/275 (59%), Gaps = 1/275 (0%)
Frame = +1
Query: 558 LIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTA 617
LIANEV+D + + K+D+ K YD++ W FL ++ M+F +WI WI CV +A
Sbjct: 13 LIANEVIDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEECVKSA 192
Query: 618 TLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGP-G 676
++S+LVNGS T F ++GLRQGDPL+PFLFN+ GL+ L+ + ++++ + +G G
Sbjct: 193 SISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYLVGANG 372
Query: 677 LALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQR 736
+ ++ LQ+ADDT+ F E + ++ + L SF S L++N KS V + Q
Sbjct: 373 VPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTDQWKQE 552
Query: 737 VAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMK 796
A S+ P +YLG P+G N RR W+ ++ K RK W + IS GR+TL+K
Sbjct: 553 AANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTLIK 732
Query: 797 ATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
+ L S+P++ + F+ P VV ++ KI R+FL GG
Sbjct: 733 SVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGG 837
>CO982349
Length = 795
Score = 148 bits (374), Expect = 1e-35
Identities = 82/225 (36%), Positives = 124/225 (54%), Gaps = 1/225 (0%)
Frame = +3
Query: 608 NWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSN 667
+WIR C+ +A++SILVN S F ++GLRQGDPL+P LFN+ GL+ L+ + +D
Sbjct: 66 HWIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKR 245
Query: 668 FSGVRIGPGLA-LNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIG 726
F+ +G ++ LQ+ADDT+ F E + ++ L F SGLK+N +S
Sbjct: 246 FNSFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGA 425
Query: 727 CNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFI 786
+ A S+ +P YLG P+ NPR +P++ K K W + I
Sbjct: 426 IWKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHI 605
Query: 787 SLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
SL GR+TL+ A L ++P++ + F+AP KVV ++ I R FL GG
Sbjct: 606 SLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGG 740
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 92.4 bits (228), Expect(2) = 3e-29
Identities = 43/97 (44%), Positives = 61/97 (62%), Gaps = 1/97 (1%)
Frame = -3
Query: 553 ISDCILIANEVVDFLNKKQ-GGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIR 611
I DC + +EV++ L+KK GG LK+D K +D +D FL +L + + + NWIR
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIR 655
Query: 612 TCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+ +A LSI VNG GFF ++G+RQGDPLSP L+
Sbjct: 654 VILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
Score = 56.2 bits (134), Expect(2) = 3e-29
Identities = 30/135 (22%), Positives = 66/135 (48%), Gaps = 1/135 (0%)
Frame = -2
Query: 677 LALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQR 736
L +H+ ADD ++FC+ + L N + + G ++ HKS I ++ + +
Sbjct: 460 LTRSHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRV 281
Query: 737 VAGMYGWSIGQLPILYLGAPL-GGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLM 795
++ + + +P Y G P+ G P R + + DK++ K +W +S+ GR+ L+
Sbjct: 280 ISDLLQYFHSDIPFNYFGVPIFKGKPTRRHLY-CVADKIKCKLASWKGSILSIMGRIQLV 104
Query: 796 KATLCSVPLFLMAIF 810
+ + + L+ +++
Sbjct: 103 NSVIHGMLLYSFSVY 59
>TC213164
Length = 446
Score = 110 bits (275), Expect = 4e-24
Identities = 50/135 (37%), Positives = 88/135 (65%)
Frame = +3
Query: 431 LEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKG 490
L +F + E+ + ++PG DG N F K+FW I+K D+++F ++F++ G +K
Sbjct: 18 LVARFEKKEVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLKR 197
Query: 491 LNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQG 550
NA F++LIPK + P ++++++PISLIG YK+++K+L+ R + V P +I E Q F +G
Sbjct: 198 SNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFMEG 377
Query: 551 RQISDCILIANEVVD 565
R ++IANE+++
Sbjct: 378 RHTLHNVVIANEIME 422
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 83.6 bits (205), Expect(2) = 1e-20
Identities = 44/111 (39%), Positives = 70/111 (62%), Gaps = 1/111 (0%)
Frame = -3
Query: 613 CVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVR 672
C+ST T+S+L NG F G+RQ DP++P+LF LC+ LS L+N +++ ++
Sbjct: 668 CMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQ 489
Query: 673 IG-PGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKS 722
+ GL ++HL F DD +LF E + +Q+D++ TL F +SG KVN+ K+
Sbjct: 488 VNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKT 336
Score = 36.2 bits (82), Expect(2) = 1e-20
Identities = 17/56 (30%), Positives = 32/56 (56%)
Frame = -2
Query: 752 YLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLM 807
YLG P+ + ++ ++DK+ ++ W S +S+ GRLTL K + ++P +M
Sbjct: 243 YLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIM 76
>AW757033
Length = 441
Score = 93.2 bits (230), Expect = 7e-19
Identities = 50/142 (35%), Positives = 76/142 (53%), Gaps = 1/142 (0%)
Frame = -2
Query: 631 FEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGP-GLALNHLQFADDTL 689
F +GLRQGDPL+P LFN+ GL+ L+ + + +++ +G ++ LQ+ADDT+
Sbjct: 431 FSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTI 252
Query: 690 LFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLP 749
F E + ++ L F SGLK+N KS +V + A S+ LP
Sbjct: 251 FFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSLLSLP 72
Query: 750 ILYLGAPLGGNPRRASFWEPML 771
YLG P+G NPRR W+P++
Sbjct: 71 FSYLGIPIGANPRRRETWDPII 6
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 92.4 bits (228), Expect = 1e-18
Identities = 40/88 (45%), Positives = 58/88 (65%)
Frame = -2
Query: 440 IWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLI 499
+W+ C ++PG DG N F K FW +K D+++F +FH+ G KG NA+FI+LI
Sbjct: 363 VWQ----CGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALI 196
Query: 500 PKHNCPSTVSDFRPISLIGSAYKLISKV 527
PK P ++DFRPISLIG YK+++K+
Sbjct: 195 PKVKHPQALNDFRPISLIGCVYKIVAKI 112
>BI321779
Length = 421
Score = 89.7 bits (221), Expect = 8e-18
Identities = 42/115 (36%), Positives = 68/115 (58%)
Frame = +2
Query: 412 RPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
RPK+ S + L + F EI +V+ C+ +++PG DG++L K FW ++K
Sbjct: 26 RPKLNGVEFLSFSNEDNAMLIQDFEVEEIKKVI*DCESSKSPGPDGYSLLLIKIFWEVIK 205
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISK 526
E+V+ FF++FH L +G N +FI+LI K + P + + PISL+G YK++ K
Sbjct: 206 EEVINFFKEFHKFDFLPRGTNRSFIALIAKCDNPLDIGQYCPISLVGCLYKIV*K 370
>AI496436
Length = 414
Score = 85.1 bits (209), Expect = 2e-16
Identities = 42/117 (35%), Positives = 68/117 (57%), Gaps = 1/117 (0%)
Frame = +1
Query: 706 LLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRA 764
L F +SGLK+N KS C D+ + A S+ +P +YLG P+G N R +
Sbjct: 7 LRCFELSSGLKINFAKSQFGTICKSDIWRKE-AADYLNSSVLSMPFVYLGIPIGANSRHS 183
Query: 765 SFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVE 821
WEP++ K RK +W K++S GR+TL+ + L ++P++L++ F+ P KVV + +
Sbjct: 184 DVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVVDKAD 354
>BI972659
Length = 453
Score = 84.7 bits (208), Expect = 3e-16
Identities = 42/138 (30%), Positives = 73/138 (52%)
Frame = +1
Query: 697 NQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAP 756
+ I +L + L F SGL++N KS A + +P YLG P
Sbjct: 19 DNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPFHYLGMP 198
Query: 757 LGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKV 816
+ WEP+++K + K WN K++S+ GR+TL+K+ L ++P++L++ FK P ++
Sbjct: 199 IAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFKIPQRI 378
Query: 817 VGEVEKILRSFLQGGKLH 834
V ++ + R+F+ GG H
Sbjct: 379 VDKLVSLQRTFMWGGNQH 432
>CA936755
Length = 424
Score = 63.5 bits (153), Expect = 6e-10
Identities = 32/90 (35%), Positives = 49/90 (53%)
Frame = -2
Query: 199 WLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGPKPFLFYNHWLLDKEFEKLVS 258
+LVSE + + SQ SD PII+ +++GPKPF + WL K +++LV
Sbjct: 294 FLVSEQWLLSWPDSSQYTLPRDFSDHCPIIMQTKKVDWGPKPFRVVDWWLHQKGYQRLVR 115
Query: 259 DWWCSATTVGWSAFVLQQKLKGLKNRIKEW 288
+ W + GW +L+ KL+ LK IK+W
Sbjct: 114 ETWSAEQQPGWGGILLKNKLRMLKLSIKQW 25
>TC218847
Length = 521
Score = 60.5 bits (145), Expect = 5e-09
Identities = 34/98 (34%), Positives = 54/98 (54%)
Frame = -3
Query: 328 LTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSI 387
L I+E W + E RQKAR +W+RE D N+S+FH + ++ +++GG +
Sbjct: 297 LIIME--WTAAQAHESLLRQKARTKWIRERDCNSSYFHKLINYIHRTNAVNGVMIGGSWV 124
Query: 388 EDPNMIKDAIFSHFQSFFTSENRLRPKIRCANLSRLSQ 425
E+PN +K A+ + FQ F+ + RP I AN + Q
Sbjct: 123 EEPNRVK-AVRNFFQIRFSESDYDRP*INGANFRSIGQ 13
>TC234193
Length = 697
Score = 57.8 bits (138), Expect = 3e-08
Identities = 28/68 (41%), Positives = 46/68 (67%)
Frame = -3
Query: 455 TDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPI 514
++G+N NF K+ ++KEDVV+ ++FHS G L +G NA+F++ + P + DFRP+
Sbjct: 197 SNGYNFNFIKKC*EVMKEDVVRAIQEFHSHGCLSRGTNASFLT-----HPP*GLGDFRPV 33
Query: 515 SLIGSAYK 522
+L+GS K
Sbjct: 32 TLVGSLCK 9
>TC233199
Length = 1105
Score = 49.7 bits (117), Expect = 9e-06
Identities = 34/118 (28%), Positives = 55/118 (45%), Gaps = 8/118 (6%)
Frame = +2
Query: 81 MESAVKNVRFIGLLLRALVDSSLLFIMNVYGP---NVDSERYDFFNLLSEALSLNNCFCK 137
+E V FI L D + I+N+Y P N+ + N L A S N +C
Sbjct: 122 LEKKVIGSGFISLEGTWKGDGEKITIVNIYSPCDVNLKRNLWGQINQLRSA-S*NKLWCI 298
Query: 138 IMGGDFNAILNEGERIGIQM-----GVESSFSDFVADSNLIDLPLQSDKFTWYSSRAG 190
+ DFN + + ER+G G+ + F+D++ D + D+P K+TWY ++G
Sbjct: 299 V--DDFNCVRRQSERVGASQREQGGGIMTEFNDWIVDLEVEDMPSVGHKYTWYRLKSG 466
>TC212055
Length = 776
Score = 48.5 bits (114), Expect = 2e-05
Identities = 27/78 (34%), Positives = 44/78 (55%), Gaps = 1/78 (1%)
Frame = +1
Query: 647 LFNLCVNGLSCLLNQLLDDSNFSGVRIGPG-LALNHLQFADDTLLFCENDENQIDLLCNT 705
LFN+ GL+ L+ + LD S +S + +G + +N LQ+ADDT+ E + + +
Sbjct: 1 LFNIVAEGLTGLMREALDKSLYSSLMVGKNKIPVNILQYADDTIFLGEATMQNVMTIKSM 180
Query: 706 LLSFLFTSGLKVNLHKST 723
L F SGLK++ KS+
Sbjct: 181 LRVFELASGLKIHFAKSS 234
>TC206673
Length = 742
Score = 48.5 bits (114), Expect = 2e-05
Identities = 24/93 (25%), Positives = 48/93 (50%)
Frame = +3
Query: 300 INTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDR 359
I+ I+ L D ++ ++ K + + + LW+ E RQK+R +WL+EGD
Sbjct: 21 IHNIKKKLNEVEDLASNRSLSVDEIKAKKELQQQLWEASTAYESLLRQKSRDKWLKEGDN 200
Query: 360 NTSFFHSISKIRKSKKLISQLVVGGVSIEDPNM 392
N+++FH++ R+ + +++ G I P +
Sbjct: 201 NSAYFHTVINFRRHYNGLQGILIQGEWIALPGL 299
>BI315739
Length = 442
Score = 36.6 bits (83), Expect(2) = 2e-05
Identities = 22/59 (37%), Positives = 32/59 (53%), Gaps = 1/59 (1%)
Frame = +2
Query: 679 LNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIG-CNVDVRETQR 736
++HL F D+ LLF + +Q+ L+ + L F T GLK L KS + NV R+ R
Sbjct: 164 ISHLFFPDNCLLFVKATSSQVKLVRDVLQQFCLT*GLKAGLQKSRFLA*KNVSNRKIAR 340
Score = 31.2 bits (69), Expect(2) = 2e-05
Identities = 13/40 (32%), Positives = 26/40 (64%), Gaps = 3/40 (7%)
Frame = +3
Query: 642 PLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRI---GPGLA 678
PLSP+LF +C+ L+ L+ +++ + ++I GPG++
Sbjct: 48 PLSPYLF*ICMEKLAILIQSKVNEETWEPIKISRNGPGIS 167
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 45.1 bits (105), Expect = 2e-04
Identities = 20/36 (55%), Positives = 27/36 (74%)
Frame = +1
Query: 611 RTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPF 646
+ C+S+A++SILVNGS T F+ +GLRQG P PF
Sbjct: 1 KECLSSASISILVNGSPTEEFKPXRGLRQGGPFGPF 108
>TC234877
Length = 534
Score = 43.1 bits (100), Expect = 8e-04
Identities = 21/87 (24%), Positives = 45/87 (51%)
Frame = +1
Query: 742 GWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCS 801
G+S+G LP +YL PL R P+ D+++ + ++W +S G + L+ + +
Sbjct: 19 GFSVGSLPFIYLSVPLFQEKPRPVHLRPIADRIKAELQSWKGNLLSFMGCVQLIISIIDG 198
Query: 802 VPLFLMAIFKAPLKVVGEVEKILRSFL 828
+ L ++ P ++ E++ R+F+
Sbjct: 199 MLL*SFHLYSWPKALLIEMQTWTRNFI 279
>CA851030
Length = 358
Score = 42.7 bits (99), Expect = 0.001
Identities = 23/72 (31%), Positives = 40/72 (54%), Gaps = 2/72 (2%)
Frame = +1
Query: 332 EGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIED 389
E L ++EE W+Q++R W +E D+NT FFH + RK+ I+++ G V +
Sbjct: 145 ESLDSLLKQEEEYWQQRSRAIWFKERDKNTEFFHRKTLQRKASNSITKIQDDEGNVCFK- 321
Query: 390 PNMIKDAIFSHF 401
P+ I+D + +
Sbjct: 322 PDQIEDILIKFY 357
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,007,971
Number of Sequences: 63676
Number of extensions: 1055345
Number of successful extensions: 8313
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 8076
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8291
length of query: 1389
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1280
effective length of database: 5,698,948
effective search space: 7294653440
effective search space used: 7294653440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0110a.13


