

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.3
(169 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
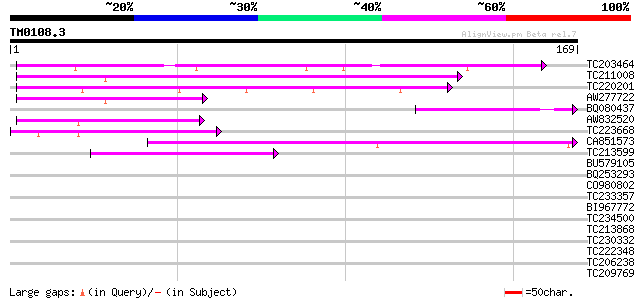
Sequences producing significant alignments: (bits) Value
TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leish... 77 5e-15
TC211008 68 2e-12
TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance ... 54 4e-08
AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa pr... 47 6e-06
BQ080437 homologue to GP|20197033|gb| expressed protein {Arabido... 46 7e-06
AW832520 45 2e-05
TC223668 43 6e-05
CA851573 43 8e-05
TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transpor... 42 2e-04
BU579105 similar to GP|12328468|dbj P0416D03.16 {Oryza sativa (j... 35 0.017
BQ253293 35 0.017
CO980802 33 0.085
TC233357 homologue to UP|Q8VWW2 (Q8VWW2) Two-component response ... 30 0.42
BI967772 27 0.45
TC234500 30 0.55
TC213868 Glycine max mRNA sequence 30 0.72
TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/... 29 1.2
TC222348 29 1.2
TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragmen... 28 1.6
TC209769 28 2.7
>TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leishmania
major;} , partial (8%)
Length = 1386
Score = 76.6 bits (187), Expect = 5e-15
Identities = 59/169 (34%), Positives = 84/169 (48%), Gaps = 11/169 (6%)
Frame = +2
Query: 3 WLLPPVGSWKFNVDGS--VWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLA----EL 56
W P +G K NVDGS +++ AGCGGVLRD W++GF ++L NP A EL
Sbjct: 488 WKKPEIGWVKLNVDGSRDPYKSSSAGCGGVLRDASAKWLRGFAKKL---NPTYAVHQTEL 658
Query: 57 WAIRTAVDLAVQRGCSPIIIETDSAEAHAAV-TSAQPLIPPYS--EIVRHIRRSIPHGMK 113
AI T + +A + +I+E+DS + V +P P Y E++R RR +
Sbjct: 659 EAILTGLKVASEMNVKKLIVESDSDSVVSMVENGVKPNHPDYGVVELIRTKRRRF--DWE 832
Query: 114 VVFDLIPREANMVADCLAQNAH--VFNFGVHLLHHPLMECRYLLYQDSM 160
V F + +AN VAD LA +A + HP C LL +D +
Sbjct: 833 VRFVSVSNKANRVADRLANDARKLACDDVCEEYLHPPANCNQLLLEDEI 979
>TC211008
Length = 516
Score = 67.8 bits (164), Expect = 2e-12
Identities = 44/138 (31%), Positives = 62/138 (44%), Gaps = 5/138 (3%)
Frame = +1
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGC-----GGVLRDCDGNWVQGFCRRL*SSNPLLAELW 57
W P G N DGSV++ GC GG++RD G ++ GF L +++ LAELW
Sbjct: 64 WRCPXNGWVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELW 243
Query: 58 AIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFD 117
+ + LA GC + ++ DS A V P +V I + V F
Sbjct: 244 GVVHGLKLAWDLGCKKVKVDIDSGNALGLVRHGPVANDPAFALVSEINELVRKEWLVEFS 423
Query: 118 LIPREANMVADCLAQNAH 135
+ RE+N AD LA H
Sbjct: 424 HVFRESNRAADKLAHLGH 477
>TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance receptor
PtdSerR (Phosphatidylserine receptor), partial (5%)
Length = 1185
Score = 53.9 bits (128), Expect = 4e-08
Identities = 47/140 (33%), Positives = 68/140 (48%), Gaps = 10/140 (7%)
Frame = +1
Query: 3 WLLPPVGSWKFNVDGSVW--QTREAGCGGVLRDCDGNWVQGFCRRL*SS---NPLLAELW 57
W P G K NVDGS + AGCGGV+RD G W GF ++L + EL
Sbjct: 337 WKKPESGWVKLNVDGSRIHEEPASAGCGGVIRDEWGTWCVGFDQKLDPNICRQAHYTELQ 516
Query: 58 AIRTAVDLAVQR--GCSPIIIETDSAEAHAAVTS--AQPLIPPYSEIVRHIRRSIPHGMK 113
AI T + +A + +++E+DS A V S P ++V+ I R +
Sbjct: 517 AILTGLKVAREDMINVEKLVVESDSEPAVNMVKSRLGYKYHRPEYKVVQDINRLLKDPKW 696
Query: 114 VV-FDLIPREANMVADCLAQ 132
+ D +PR+AN VA+ LA+
Sbjct: 697 LARLDYVPRKANRVANYLAK 756
>AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa protein 3
(Hsc70.3). [Mouse-ear cress] {Arabidopsis thaliana},
partial (8%)
Length = 432
Score = 46.6 bits (109), Expect = 6e-06
Identities = 24/62 (38%), Positives = 34/62 (54%), Gaps = 5/62 (8%)
Frame = +1
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGC-----GGVLRDCDGNWVQGFCRRL*SSNPLLAELW 57
W PP G N DGSV++ GC GG++RD G ++ GF L +++ LAELW
Sbjct: 247 WRCPPNGWVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELW 426
Query: 58 AI 59
+
Sbjct: 427 GV 432
>BQ080437 homologue to GP|20197033|gb| expressed protein {Arabidopsis
thaliana}, partial (13%)
Length = 422
Score = 46.2 bits (108), Expect = 7e-06
Identities = 23/48 (47%), Positives = 29/48 (59%)
Frame = -2
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTVVASLSLP 169
E N VAD LA AH + G+H++ HP ECR LL +D V +SLP
Sbjct: 277 EMNSVADYLANEAHNYEVGIHIVDHPPAECRLLLAED----VGGISLP 146
>AW832520
Length = 423
Score = 45.1 bits (105), Expect = 2e-05
Identities = 25/57 (43%), Positives = 32/57 (55%), Gaps = 1/57 (1%)
Frame = -3
Query: 3 WLLPPVGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWA 58
W PP G +K N DG+V + + GGV+RD GN+V GF S +AELWA
Sbjct: 250 WSPPPEGLFKLNYDGAVNCNSLKVSAGGVVRDSHGNFVIGFTTTPDSCTINIAELWA 80
>TC223668
Length = 866
Score = 43.1 bits (100), Expect = 6e-05
Identities = 25/66 (37%), Positives = 36/66 (53%), Gaps = 3/66 (4%)
Frame = -3
Query: 1 LCWLLPP--VGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELW 57
L W PP + K NVDGS ++ GG++RD G W+ F R NPLL+EL+
Sbjct: 591 LTWTSPPPLLNEVKLNVDGSGHYEPHVMEVGGLIRDGVGRWLFRFARHCGLGNPLLSELF 412
Query: 58 AIRTAV 63
A+ ++
Sbjct: 411 ALEASL 394
>CA851573
Length = 614
Score = 42.7 bits (99), Expect = 8e-05
Identities = 36/130 (27%), Positives = 58/130 (43%), Gaps = 2/130 (1%)
Frame = +1
Query: 42 FCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIV 101
F +RL N +LAELW+++ + LA+ + E D+ EA + + + +
Sbjct: 64 FSKRLGYCNVILAELWSLKYGIKLAIDLNTQFSLFEMDAREAVHMIDXIKSVPSSLRPLA 243
Query: 102 RHIRRSI-PHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSM 160
IRR + +G + I REAN A+ A+ H N+ L+ + L+ D
Sbjct: 244 ADIRRVLWANGWQSSIVHINREANSAANWAAKFGHQRNWSSFLVDEIPFVLKALVANDVR 423
Query: 161 TVVAS-LSLP 169
AS L LP
Sbjct: 424 RAAASPLRLP 453
>TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transport exbB
protein, partial (8%)
Length = 825
Score = 41.6 bits (96), Expect = 2e-04
Identities = 20/56 (35%), Positives = 30/56 (52%)
Frame = +3
Query: 25 AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDS 80
A CGG+LRDC G W F +L +++ +AEL + LA +G + + DS
Sbjct: 351 AACGGLLRDCHGRWAASFV*KLGNTSAFVAEL*GAYLGLSLAWDKGDKNLEVNIDS 518
>BU579105 similar to GP|12328468|dbj P0416D03.16 {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 425
Score = 35.0 bits (79), Expect = 0.017
Identities = 17/32 (53%), Positives = 21/32 (65%), Gaps = 2/32 (6%)
Frame = +3
Query: 3 WLLPPVGSWKFNVDGS--VWQTREAGCGGVLR 32
W P +G K NVDGS +++ AGCGGVLR
Sbjct: 330 WKKPEIGWVKLNVDGSRDPYKSSSAGCGGVLR 425
>BQ253293
Length = 420
Score = 35.0 bits (79), Expect = 0.017
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = +2
Query: 1 LCWLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQG 41
+CW PP K N + +V + A CGGV+RD G + G
Sbjct: 128 ICWEPPPTVQLKLNCEAAVNRNGMASCGGVMRDPYGKVLWG 250
>CO980802
Length = 803
Score = 32.7 bits (73), Expect = 0.085
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Frame = -3
Query: 4 LLPPVGSWKFNVDGSVW-QTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTA 62
LL P+ +K NVDGS T + C G++ D G + F + N + A+LW ++
Sbjct: 423 LLGPLLFFKVNVDGSYKASTAKMACRGIILDSFGRVLSCFHSNIGVCNSIWAKLWGLKHG 244
Query: 63 V 63
+
Sbjct: 243 I 241
>TC233357 homologue to UP|Q8VWW2 (Q8VWW2) Two-component response
regulator-like protein, partial (8%)
Length = 811
Score = 30.4 bits (67), Expect = 0.42
Identities = 20/64 (31%), Positives = 28/64 (43%)
Frame = +1
Query: 72 SPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLA 131
S + +E+DS + P IV+ I I G+ + + REAN V DC A
Sbjct: 484 SYLYVESDSLINIQLLKLKYTTTHPAYGIVKMIPDFIDDGVHITWSRALREANQVVDCFA 663
Query: 132 QNAH 135
AH
Sbjct: 664 NYAH 675
>BI967772
Length = 719
Score = 27.3 bits (59), Expect(2) = 0.45
Identities = 17/52 (32%), Positives = 27/52 (51%)
Frame = +3
Query: 29 GVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDS 80
G+ RD G+ + F ++ L AE W I + +A + G +IIE+DS
Sbjct: 435 GLQRDHSGDVLFCFAAKVDYCFVLHAESWTIYIGLIIAWEEGFKNLIIESDS 590
Score = 21.6 bits (44), Expect(2) = 0.45
Identities = 8/19 (42%), Positives = 10/19 (52%)
Frame = +1
Query: 1 LCWLLPPVGSWKFNVDGSV 19
+CW P G K N + SV
Sbjct: 352 ICWNKAPTGVLKLNCNASV 408
>TC234500
Length = 866
Score = 30.0 bits (66), Expect = 0.55
Identities = 18/43 (41%), Positives = 26/43 (59%), Gaps = 5/43 (11%)
Frame = +3
Query: 42 FC-RRL*SSNP----LLAELWAIRTAVDLAVQRGCSPIIIETD 79
FC R L S+ P + LW++ TA+ LA ++G S + IETD
Sbjct: 396 FCVRSLDSAKPCSHWFIVVLWSVITALKLAKEKGVSQLWIETD 524
Score = 29.3 bits (64), Expect = 0.94
Identities = 16/34 (47%), Positives = 21/34 (61%), Gaps = 1/34 (2%)
Frame = +1
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMEC-RYL 154
EA VAD L++ AH FG+H+L P C +YL
Sbjct: 652 EATTVADWLSKYAHNIAFGLHVLDVPQHGCVKYL 753
>TC213868 Glycine max mRNA sequence
Length = 1272
Score = 29.6 bits (65), Expect = 0.72
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = -1
Query: 38 WVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDS 80
W++ + SS+PLL L + + + + RGCSP + ++DS
Sbjct: 627 WLESCTKPSISSSPLLLLLKSDSASAAVLMDRGCSPKLTQSDS 499
>TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/k19m22_160),
partial (36%)
Length = 827
Score = 28.9 bits (63), Expect = 1.2
Identities = 18/55 (32%), Positives = 25/55 (44%), Gaps = 4/55 (7%)
Frame = -1
Query: 119 IPREANMVADCLAQNAHVFNFGVHLL----HHPLMECRYLLYQDSMTVVASLSLP 169
I E+N L Q H+ +HLL HH L+ C YL ++ A+ S P
Sbjct: 500 ISSESNSTIKLLPQILHLQILHLHLLLQLLHHRLVRCHYLPHRRHPIAAAATSSP 336
>TC222348
Length = 995
Score = 28.9 bits (63), Expect = 1.2
Identities = 16/42 (38%), Positives = 24/42 (57%)
Frame = +2
Query: 121 REANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTV 162
++ N VAD LA AH + +LH P + C LL++D + V
Sbjct: 779 KKGNRVADWLANWAHHGHESPLILHSPSVGCVSLLWEDFIGV 904
>TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragment), partial
(9%)
Length = 1259
Score = 28.5 bits (62), Expect = 1.6
Identities = 21/63 (33%), Positives = 33/63 (52%), Gaps = 5/63 (7%)
Frame = +3
Query: 9 GSW--KFNVDGSVWQTRE-AGCGGVLRDCDGNWVQGFCRRL*S--SNPLLAELWAIRTAV 63
G W + NV G+ ++ + A CGG+ RD + +V GF +L SN E+W I +
Sbjct: 612 GHWLLRLNVSGAYDRSSDTAACGGIFRDNNDRFVLGFSVKLGECLSND-EGEIWGIYHGM 788
Query: 64 DLA 66
+A
Sbjct: 789 KIA 797
>TC209769
Length = 1204
Score = 27.7 bits (60), Expect = 2.7
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +2
Query: 132 QNAHVFNFGVHLLHHPL 148
Q++H+F G+HLLHH L
Sbjct: 704 QHSHLFKPGMHLLHHSL 754
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,380,379
Number of Sequences: 63676
Number of extensions: 167696
Number of successful extensions: 934
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 927
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 932
length of query: 169
length of database: 12,639,632
effective HSP length: 90
effective length of query: 79
effective length of database: 6,908,792
effective search space: 545794568
effective search space used: 545794568
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0108.3


