

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.13
(1262 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
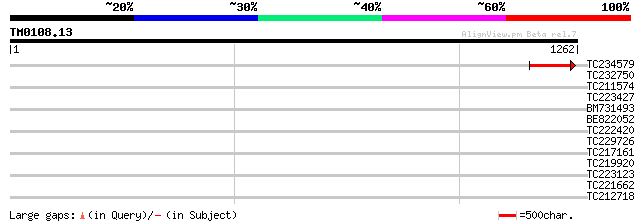
Sequences producing significant alignments: (bits) Value
TC234579 141 2e-33
TC232750 42 0.002
TC211574 40 0.007
TC223427 34 0.47
BM731493 33 1.0
BE822052 similar to GP|4557063|gb|A expressed protein {Arabidops... 32 1.8
TC222420 32 1.8
TC229726 similar to GB|AAP37709.1|30725374|BT008350 At5g06570 {A... 32 2.3
TC217161 32 2.3
TC219920 31 3.0
TC223123 similar to UP|Q9VY72 (Q9VY72) CG32611-PB, partial (3%) 30 6.7
TC221662 weakly similar to UP|Q6TDU2 (Q6TDU2) Coronatine-insensi... 30 8.8
TC212718 similar to UP|Q91CY9 (Q91CY9) ORF1, partial (4%) 30 8.8
>TC234579
Length = 506
Score = 141 bits (355), Expect = 2e-33
Identities = 71/102 (69%), Positives = 85/102 (82%)
Frame = +3
Query: 1158 AERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDSVNDNPGFDRDQVKEACRKAYA 1217
AERAFE+S+NQLLDAEEVA +L+KELSHLR+ML DSVN+NP F QVKEACRKA A
Sbjct: 12 AERAFEFSKNQLLDAEEVAHNLMKELSHLRNMLESSADSVNNNPVFGGSQVKEACRKACA 191
Query: 1218 AEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKIIK 1259
AE+LAK RLS M DDL+IHCRIT+LQ P V FA HV++++I+
Sbjct: 192 AEELAKNRLSQMHDDLNIHCRITSLQPPTVTFAVHVEKEVIQ 317
>TC232750
Length = 746
Score = 42.0 bits (97), Expect = 0.002
Identities = 30/108 (27%), Positives = 55/108 (50%), Gaps = 3/108 (2%)
Frame = +1
Query: 1147 TCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDS---VNDNPGF 1203
T + C S + ++ A +++ Q+ DAE +A L KEL ++D++ + S +N +
Sbjct: 160 TRIGCNSVQSCSKSAIAFTQQQMHDAECLAMRLTKELKSMKDIVDDMLRSEFCLNTSLRH 339
Query: 1204 DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFAD 1251
++ + A + A AE+ K L+ M D S C+I ++FAD
Sbjct: 340 KVNEARMAVKNATRAEEATKRCLAFMSRDCSRFCKI-------MKFAD 462
>TC211574
Length = 1008
Score = 40.0 bits (92), Expect = 0.007
Identities = 24/57 (42%), Positives = 30/57 (52%), Gaps = 3/57 (5%)
Frame = -1
Query: 26 PSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQ---GGLYSGLGGSSGNDEKREPP 79
P ++P RSR L+ R SG R P GPR+ G L SG GGS G+ +R P
Sbjct: 192 PGTAPRDCTPRSRRSLYPQRWFSGVRGPGGGPRRTGPGRLQSGRGGSCGSLRRRGGP 22
>TC223427
Length = 701
Score = 33.9 bits (76), Expect = 0.47
Identities = 19/48 (39%), Positives = 26/48 (53%), Gaps = 2/48 (4%)
Frame = +2
Query: 903 SKITPVLNRCPRI--KSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRH 948
S I ++NRCPRI SL+ PT++ + L + KK C NRH
Sbjct: 419 SDIVHLVNRCPRIINSSLEKNVIPTFELVKRFLQSDKKTIDCVFANRH 562
>BM731493
Length = 427
Score = 32.7 bits (73), Expect = 1.0
Identities = 30/106 (28%), Positives = 45/106 (42%)
Frame = +1
Query: 798 RKYVSRYPSEGQSSSQLDTDMQDLEENERSNGAIRKSDDAVVLNHVGVSNEVPRPPPENI 857
R +RY E D + D ++ E NG DD N+V PR P +
Sbjct: 46 RPMPTRY-QEPDDDDDDDNEPDDFDDGEAENGY---DDD----NNVAY----PRIPKKRK 189
Query: 858 ISKTQDMAAHGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENASES 903
+ AA GG+ E + + +YGS+G+ G + E N SE+
Sbjct: 190 VFSAAAAAAAGGSYEFAPRVKFSYGSRGSGSGSGSGSGEEWNESET 327
>BE822052 similar to GP|4557063|gb|A expressed protein {Arabidopsis
thaliana}, partial (28%)
Length = 668
Score = 32.0 bits (71), Expect = 1.8
Identities = 26/94 (27%), Positives = 40/94 (41%), Gaps = 3/94 (3%)
Frame = -1
Query: 206 ISSQARDFDRANTDSSDH---KQFEEGNDGDCSGKINDFDEEIVVTTPPDAELLGDSKVD 262
++ A D D + DH Q +E D D SG + +EE P+A G S D
Sbjct: 539 LNKDASDSDEDEDEEDDHDTNDQNDEEGDEDFSG---EGEEEADPEDDPEANGAGGSNDD 369
Query: 263 GDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSK 296
D + ++D D G+ E+ EG +D +
Sbjct: 368 DDSDEDDDD--DNDGDDDGEDEDEKEGEDEEDEE 273
>TC222420
Length = 582
Score = 32.0 bits (71), Expect = 1.8
Identities = 45/173 (26%), Positives = 65/173 (37%), Gaps = 16/173 (9%)
Frame = +2
Query: 967 SDLRATSGGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKV 1026
SD SG D+ DS+ +N+ +++ R +S Q+S SK
Sbjct: 101 SDKEDDSGVDNADSLEQIKTKSNAEHMDKRQ-------------RKKSNSKQTSTSKSIS 241
Query: 1027 MQMCDEQVLLNGLCKPESSSGT----PISV----RRIDSPSTPLRPIVNEVMAREEATFA 1078
+ +QVL + P S SGT ++V + + ST N + +
Sbjct: 242 SMLTKDQVLDSKKPSPSSGSGTHGKPSVAVTKKSKSTSTKSTNKSSKTNLELVEDSVRLE 421
Query: 1079 LSESLINSKVK-----GNCSFSISPDDKEKLSEAHGCCLSLS---QLELKEKL 1123
LS+S NS K CS DD + H LS QL KEKL
Sbjct: 422 LSDSRFNSGGKDAAQHSTCSPGEKDDDNLEAINGHTHAEDLSSSQQLSSKEKL 580
>TC229726 similar to GB|AAP37709.1|30725374|BT008350 At5g06570 {Arabidopsis
thaliana;} , partial (36%)
Length = 674
Score = 31.6 bits (70), Expect = 2.3
Identities = 16/54 (29%), Positives = 23/54 (41%)
Frame = -2
Query: 31 PGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLGGSSGNDEKREPPEEDDG 84
PG SR R R +RP S R G++ +R PP+++DG
Sbjct: 544 PGGAQCSRQTSHPRRRRGQSQRPHSTTRPPSSSGPASGATSPPPRRRPPQQEDG 383
>TC217161
Length = 1526
Score = 31.6 bits (70), Expect = 2.3
Identities = 41/203 (20%), Positives = 92/203 (45%), Gaps = 24/203 (11%)
Frame = +1
Query: 721 VDFHSTKIVANDGHDVDIKHMQNSISSLQERGPSPKDQSMLYVNVGVSNQTLEHHSSEV- 779
+D H ++ D HDV ++ QNS+ SL++ V ++ +++SSE+
Sbjct: 472 IDNHMLELCETDQHDVTEENKQNSV-SLED------------VITCHESERCQNYSSELE 612
Query: 780 --KGFSVNNGEV----KQLLNME-KRKYVSRYPSEGQSSSQLDTDMQDLE---ENERSNG 829
K V+ E+ K+ + + KRK + R S+ Q + L +QDL+ E ++ +
Sbjct: 613 TEKELGVDKLELWKTRKETTSEDSKRKILERLASDSQRLAILKMTLQDLKKKPETQKKSS 792
Query: 830 AIRKSD-----------DAVVLNHVGVSNEVPRPPPENIISKTQDMAAHGGNKEG--SVQ 876
+ + + + V+ +G+ +++ + E S + D + K+G + +
Sbjct: 793 KVNEIEYETVKRHIEDVEEAVMKQIGIYDQLAK-DTEECTSSSSDTSTMQLEKQGGQTQR 969
Query: 877 KGIAYGSKGNNGRKGCYAYEIEN 899
K + ++ + + G +E++N
Sbjct: 970 KKLTEQARRGSEQIGRLQFEVQN 1038
>TC219920
Length = 941
Score = 31.2 bits (69), Expect = 3.0
Identities = 20/84 (23%), Positives = 34/84 (39%)
Frame = +3
Query: 638 HGTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVEEPI 697
+G ++ + F+SA S G +P + S+ P +P V S P + +
Sbjct: 174 NGVASASVTPAMSFSSAMYGSGGTIPYMVDSRGAPVVPQVGGSSSTVLSSYSQPPIFMNM 353
Query: 698 EKAVVGSKDECSREPNFDLCSAIV 721
+G PNFDL S+ +
Sbjct: 354 TGTQLGLNGFGPSRPNFDLNSSFM 425
>TC223123 similar to UP|Q9VY72 (Q9VY72) CG32611-PB, partial (3%)
Length = 680
Score = 30.0 bits (66), Expect = 6.7
Identities = 16/49 (32%), Positives = 22/49 (44%)
Frame = +1
Query: 24 SNPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLGGSSGN 72
+ P PP R R L R+ + G GG++ G+GGS GN
Sbjct: 295 AQPPPPPPPLRHRDRPLLLGARSPTRASGSSGGGGGGGIWMGVGGSEGN 441
>TC221662 weakly similar to UP|Q6TDU2 (Q6TDU2) Coronatine-insensitive 1,
partial (44%)
Length = 993
Score = 29.6 bits (65), Expect = 8.8
Identities = 17/58 (29%), Positives = 29/58 (49%)
Frame = +2
Query: 1204 DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKIIKGR 1261
DRD V + CR+ Y + L ++ +++ C T R R RF H++ +KG+
Sbjct: 113 DRDAVSQVCRRLYELDSLTRKHVTIAL------CYTTTPDRLRRRF-PHLESLNLKGK 265
>TC212718 similar to UP|Q91CY9 (Q91CY9) ORF1, partial (4%)
Length = 428
Score = 29.6 bits (65), Expect = 8.8
Identities = 21/72 (29%), Positives = 32/72 (44%)
Frame = -3
Query: 8 RESASAMEETLISRKRSNPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLG 67
R S+ +++S RSN SSPP + R +L R R +RR G + +
Sbjct: 339 RSLPSSPSPSILSAARSNSRSSPPRTPSSPRRRLRHWRRRRRRRRRRKGS*RRSPTALAS 160
Query: 68 GSSGNDEKREPP 79
+SG+ E P
Sbjct: 159 SASGHPEATPSP 124
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,254,406
Number of Sequences: 63676
Number of extensions: 712176
Number of successful extensions: 3025
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2979
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3021
length of query: 1262
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1154
effective length of database: 5,762,624
effective search space: 6650068096
effective search space used: 6650068096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0108.13


