

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.15
(120 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
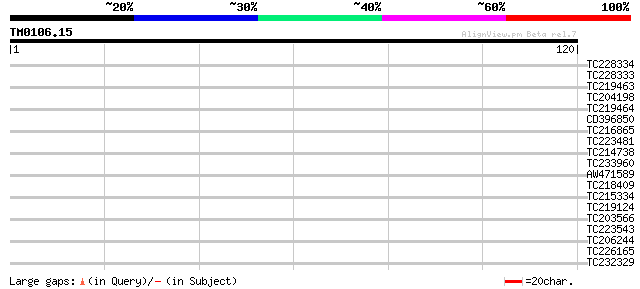
Score E
Sequences producing significant alignments: (bits) Value
TC228334 similar to UP|Q9SW80 (Q9SW80) Bel1-like homeodomain 2 (... 28 0.46
TC228333 homologue to UP|Q9M7S0 (Q9M7S0) Homeodomain protein, pa... 28 0.60
TC219463 homologue to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, p... 28 0.60
TC204198 weakly similar to UP|Q84VQ4 (Q84VQ4) DnaJ domain family... 27 1.0
TC219464 similar to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic pro... 27 1.0
CD396850 27 1.8
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 27 1.8
TC223481 similar to UP|Q9FFR3 (Q9FFR3) 6-phosphogluconate dehydr... 26 2.3
TC214738 weakly similar to UP|Q9FQ03 (Q9FQ03) XRN3, partial (17%) 26 3.0
TC233960 similar to UP|OPSD_SEPOF (O16005) Rhodopsin, partial (10%) 25 3.9
AW471589 25 3.9
TC218409 similar to UP|Q9LTW3 (Q9LTW3) DNA-3-methyladenine glyco... 25 3.9
TC215334 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (N... 25 3.9
TC219124 similar to UP|Q9LH71 (Q9LH71) Receptor-like serine/thre... 25 3.9
TC203566 similar to UP|Q9FFN2 (Q9FFN2) Similarity to limonene cy... 25 5.1
TC223543 25 5.1
TC206244 similar to PIR|B86314|B86314 F2H15.12 protein - Arabido... 25 6.7
TC226165 GAI1 [Glycine max] 25 6.7
TC232329 similar to PIR|F86340|F86340 protein F2D10.34 [imported... 24 8.7
>TC228334 similar to UP|Q9SW80 (Q9SW80) Bel1-like homeodomain 2
(AT4g36870/C7A10_490), partial (9%)
Length = 910
Score = 28.5 bits (62), Expect = 0.46
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = +1
Query: 90 YGTLNGKESLTLGCEHGGKFKPYKGKFDIR 119
+GT G SLTLG H G K F +R
Sbjct: 502 FGTTPGDVSLTLGLRHAGNMPSEKTPFSVR 591
>TC228333 homologue to UP|Q9M7S0 (Q9M7S0) Homeodomain protein, partial (14%)
Length = 1129
Score = 28.1 bits (61), Expect = 0.60
Identities = 14/30 (46%), Positives = 16/30 (52%)
Frame = +3
Query: 90 YGTLNGKESLTLGCEHGGKFKPYKGKFDIR 119
+GT G SLTLG H G P K F +R
Sbjct: 597 FGTTPGDVSLTLGLRHAGNM-PEKSPFSVR 683
>TC219463 homologue to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, partial (21%)
Length = 1111
Score = 28.1 bits (61), Expect = 0.60
Identities = 14/30 (46%), Positives = 16/30 (52%)
Frame = +2
Query: 90 YGTLNGKESLTLGCEHGGKFKPYKGKFDIR 119
+GT G SLTLG H G P K F +R
Sbjct: 644 FGTTTGDVSLTLGLRHAGNM-PEKTPFSVR 730
>TC204198 weakly similar to UP|Q84VQ4 (Q84VQ4) DnaJ domain family, partial
(50%)
Length = 875
Score = 27.3 bits (59), Expect = 1.0
Identities = 16/71 (22%), Positives = 35/71 (48%)
Frame = +2
Query: 10 DNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVF 69
D V +IKV +K + E D++++ +LKE ++++ ++ R+++
Sbjct: 359 DLVGGAYNIKVQELKSQNLEEDELEKKLELLKESYTILSSEEE-------------RRIY 499
Query: 70 DWAQLVGKNND 80
DW+ +N D
Sbjct: 500 DWSLARAENTD 532
>TC219464 similar to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic protein 13
(Fragment), partial (10%)
Length = 634
Score = 27.3 bits (59), Expect = 1.0
Identities = 14/30 (46%), Positives = 16/30 (52%)
Frame = +3
Query: 90 YGTLNGKESLTLGCEHGGKFKPYKGKFDIR 119
+GT G SLTLG H G P K F +R
Sbjct: 408 FGTTAGDVSLTLGLRHAGNM-PEKTPFSVR 494
>CD396850
Length = 638
Score = 26.6 bits (57), Expect = 1.8
Identities = 9/17 (52%), Positives = 14/17 (81%)
Frame = +2
Query: 59 VVMKFQNRQVFDWAQLV 75
++ FQ R+++DWAQLV
Sbjct: 368 IIGAFQRRKLYDWAQLV 418
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 26.6 bits (57), Expect = 1.8
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +1
Query: 12 VAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAID 51
+ G + R KE E + EW H+L+E D+M+ +D
Sbjct: 580 IINGRSNTIHRQKE---ESFSVLEWAHLLREKGDIMDLVD 690
>TC223481 similar to UP|Q9FFR3 (Q9FFR3) 6-phosphogluconate dehydrogenase
(AT5g41670/MBK23_20), partial (24%)
Length = 469
Score = 26.2 bits (56), Expect = 2.3
Identities = 16/57 (28%), Positives = 27/57 (47%)
Frame = -3
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVM 61
L+ VD+ G V+ V D + +WG L ED+D A++ N ++ V+
Sbjct: 419 LVAAVDDAVAG-------VEVVGEGGDGVVDWGACLDEDDDGSGALEGENEVARGVL 270
>TC214738 weakly similar to UP|Q9FQ03 (Q9FQ03) XRN3, partial (17%)
Length = 1587
Score = 25.8 bits (55), Expect = 3.0
Identities = 12/51 (23%), Positives = 25/51 (48%)
Frame = +3
Query: 36 WGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGKNNDFVVIMI 86
WG I+ +++ +N L H++MK +V Q + N+ ++ M+
Sbjct: 912 WGTIILASNQILSKEMTINTLDHIIMKEITIKVVGVVQGMDINHQEIITML 1064
>TC233960 similar to UP|OPSD_SEPOF (O16005) Rhodopsin, partial (10%)
Length = 862
Score = 25.4 bits (54), Expect = 3.9
Identities = 12/38 (31%), Positives = 24/38 (62%), Gaps = 3/38 (7%)
Frame = -3
Query: 14 EGDHIKVDRVKEVDNEVDDMKEWGHILKE---DEDLMN 48
EG+H++ D E DN+ ++E G++ ++ +ED+ N
Sbjct: 785 EGNHLEGDNRLEEDNQNHKLEEKGNLEEDNRLEEDIQN 672
>AW471589
Length = 405
Score = 25.4 bits (54), Expect = 3.9
Identities = 11/39 (28%), Positives = 20/39 (51%)
Frame = +2
Query: 7 EKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDED 45
+ +DN E + ++ D E D+ D + GH + DE+
Sbjct: 275 QSIDNGTEENEVQDDNDSETDSRSDTIDVEGHDEESDEE 391
>TC218409 similar to UP|Q9LTW3 (Q9LTW3) DNA-3-methyladenine glycosidase
I-like protein, partial (77%)
Length = 1263
Score = 25.4 bits (54), Expect = 3.9
Identities = 18/83 (21%), Positives = 33/83 (39%), Gaps = 3/83 (3%)
Frame = +2
Query: 38 HILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGKNNDFVVIMIRSDYGT---LN 94
+I DE+ + D L +++ + DW ++ K DF D T L
Sbjct: 320 YIAYHDEEWGVPVHDDKMLFELLVLSGAQVGSDWTSILKKRQDFRAAFSEFDVATLANLT 499
Query: 95 GKESLTLGCEHGGKFKPYKGKFD 117
K+ +++ E+G +G D
Sbjct: 500 DKQMVSISLEYGIDISQVRGVVD 568
>TC215334 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (NAD+) ,
partial (92%)
Length = 2249
Score = 25.4 bits (54), Expect = 3.9
Identities = 11/28 (39%), Positives = 14/28 (49%)
Frame = +2
Query: 9 VDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
+ +VAEGDH VDR + D W
Sbjct: 545 ISHVAEGDHEDVDRAVAAARKAFDRGPW 628
>TC219124 similar to UP|Q9LH71 (Q9LH71) Receptor-like serine/threonine
kinase, partial (15%)
Length = 871
Score = 25.4 bits (54), Expect = 3.9
Identities = 10/33 (30%), Positives = 17/33 (51%)
Frame = +3
Query: 19 KVDRVKEVDNEVDDMKEWGHILKEDEDLMNAID 51
K + + E + +W H+LKE +LM +D
Sbjct: 255 KSNTIHRSKQEALHLLDWAHLLKEKGNLMELVD 353
>TC203566 similar to UP|Q9FFN2 (Q9FFN2) Similarity to limonene cyclase,
partial (62%)
Length = 951
Score = 25.0 bits (53), Expect = 5.1
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +3
Query: 98 SLTLGCEHGGKFKPYKGKF 116
S TL C+H + +PYKG F
Sbjct: 168 SCTLQCQHL*RIRPYKGCF 224
>TC223543
Length = 549
Score = 25.0 bits (53), Expect = 5.1
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = -2
Query: 15 GDHIKVDRVKEVDNEVDDMKEWGHILKEDED 45
G HIKV +N VD ++ LK ++D
Sbjct: 425 GQHIKVGGTSSCNNRVDQSRKQTFQLKNEQD 333
>TC206244 similar to PIR|B86314|B86314 F2H15.12 protein - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (37%)
Length = 479
Score = 24.6 bits (52), Expect = 6.7
Identities = 10/28 (35%), Positives = 18/28 (63%), Gaps = 2/28 (7%)
Frame = -1
Query: 95 GKESLTLGCEHGGKFKPYKGK--FDIRV 120
G ++ GC + KF+ ++GK F+IR+
Sbjct: 347 GMDATNFGCSYNDKFRFFRGKEGFNIRL 264
>TC226165 GAI1 [Glycine max]
Length = 1313
Score = 24.6 bits (52), Expect = 6.7
Identities = 15/50 (30%), Positives = 24/50 (48%), Gaps = 1/50 (2%)
Frame = +3
Query: 32 DMKEWGHILKEDEDLM-NAIDDMNALSHVVMKFQNRQVFDWAQLVGKNND 80
DM E L+ E+ M N DD+ LS+ + + + +W Q + N D
Sbjct: 237 DMAEVAQKLERLEEAMGNVQDDLTDLSNDAVHYNPSDISNWLQTMLSNFD 386
>TC232329 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(32%)
Length = 650
Score = 24.3 bits (51), Expect = 8.7
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = -2
Query: 77 KNNDFVVIMIRSDYGTLNGKESLTLGC 103
+NN+ + I IRS G++ + L L C
Sbjct: 421 RNNNIITIRIRSAIGSIVAADGLGLRC 341
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.139 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,155,846
Number of Sequences: 63676
Number of extensions: 40495
Number of successful extensions: 190
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 190
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 190
length of query: 120
length of database: 12,639,632
effective HSP length: 96
effective length of query: 24
effective length of database: 6,526,736
effective search space: 156641664
effective search space used: 156641664
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0106.15


