

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.12
(329 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
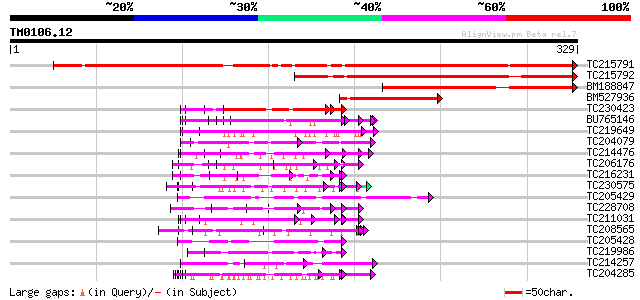
Score E
Sequences producing significant alignments: (bits) Value
TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to p... 479 e-136
TC215792 243 7e-65
BM188847 similar to GP|18700169|gb| AT3g07560/F21O3_27 {Arabidop... 170 7e-43
BM527936 similar to GP|18700169|gb| AT3g07560/F21O3_27 {Arabidop... 108 3e-24
TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fra... 83 1e-16
BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvo... 81 6e-16
TC219649 weakly similar to UP|Q95JD1 (Q95JD1) Basic proline-rich... 81 6e-16
TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding ... 77 1e-14
TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 77 1e-14
TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein... 72 4e-13
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 70 2e-12
TC230575 similar to GB|AAM97131.1|22531255|AY136466 ribonucleopr... 69 4e-12
TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich prote... 68 5e-12
TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wal... 67 1e-11
TC211031 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 67 1e-11
TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleopr... 66 2e-11
TC205428 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 66 2e-11
TC219986 weakly similar to UP|Q41187 (Q41187) Glycine-rich prote... 65 3e-11
TC214257 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 65 4e-11
TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 65 5e-11
>TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to peroxisomal
membrane protein pex13 {Neurospora crassa;} , partial
(9%)
Length = 1440
Score = 479 bits (1233), Expect = e-136
Identities = 244/305 (80%), Positives = 259/305 (84%), Gaps = 1/305 (0%)
Frame = +3
Query: 26 SNPPPKPWQQGGSSSGPTAPFKPPSAGSTSDVVEASGTARPGEIVSSSDRSAAVNRNTLA 85
++PPPKPW+Q GSSSGP APFKPPSAG+TSDVVEASGTA+PGEIVSSSDRSAAVNRNTL
Sbjct: 141 NSPPPKPWEQAGSSSGP-APFKPPSAGNTSDVVEASGTAKPGEIVSSSDRSAAVNRNTLG 317
Query: 86 RPVPTRPWEQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGM 145
RPVPTRPWEQN GGYGSTMNYNSGYGSG+YGSSY GGLGGG+ G YGGGM
Sbjct: 318 RPVPTRPWEQNYGNSTYGGYGSTMNYNSGYGSGMYGSSY-----GGLGGGMYGSSYGGGM 482
Query: 146 YGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
YG+SMY RGGY GG+YGSS GMYGGGMY GLGGPMGGY GMG GPY G+QDPNNPYG
Sbjct: 483 YGNSMY-RGGY-GGMYGSS-GMYGGGMYNSGLGGPMGGY--GMGGGPY-GEQDPNNPYGA 644
Query: 206 PPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLFDRSGLLYGELARF 265
PPSPPGFWIS +RVMQGVVNFFGRISILIDQNTQAFH+FM+ALLQLFDRSG+LYGELARF
Sbjct: 645 PPSPPGFWISALRVMQGVVNFFGRISILIDQNTQAFHLFMTALLQLFDRSGMLYGELARF 824
Query: 266 VLRLLGIKTKPNKVNPQGPNGHMLGPNGHPLPGPHNSSANVNAIEGAKPA-SGGWDNVWG 324
VLRLLGIKTK KV+P GPNG L P P PGPHN S NVN IE K A SG WDNVWG
Sbjct: 825 VLRLLGIKTKSKKVHPPGPNGQPL-PGPGPGPGPHNPSGNVNYIEAPKAAPSGSWDNVWG 1001
Query: 325 NDANQ 329
NDANQ
Sbjct: 1002NDANQ 1016
>TC215792
Length = 801
Score = 243 bits (621), Expect = 7e-65
Identities = 127/167 (76%), Positives = 136/167 (81%), Gaps = 3/167 (1%)
Frame = +2
Query: 166 GMYGGGMYGGGL-GGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGFWISVMRVMQGVV 224
GMYGGGMY G P+GGYG MG GPYG DQDPN+P+G PPSPPGFWISV+RVMQGVV
Sbjct: 32 GMYGGGMYNXVP*GVPIGGYG--MGGGPYG-DQDPNDPFGAPPSPPGFWISVLRVMQGVV 202
Query: 225 NFFGRISILIDQNTQAFHMFMSALLQLFDRSGLLYGELARFVLRLLGIKTKPNKVNPQGP 284
NFFGRISILIDQNTQAFH+FM+ALLQLFDRSG+LYGELARFVLRLLGI+TK KV P GP
Sbjct: 203 NFFGRISILIDQNTQAFHLFMTALLQLFDRSGMLYGELARFVLRLLGIRTKSKKVYPPGP 382
Query: 285 NGH-MLGPNGHPLPGPHNSSANVNAIEGAKPA-SGGWDNVWGNDANQ 329
NG +LG PGPHN S NVN IE K A SG WDNVWGNDANQ
Sbjct: 383 NGQPLLG------PGPHNPSGNVNYIEAPKAAPSGSWDNVWGNDANQ 505
>BM188847 similar to GP|18700169|gb| AT3g07560/F21O3_27 {Arabidopsis
thaliana}, partial (24%)
Length = 416
Score = 170 bits (431), Expect = 7e-43
Identities = 87/114 (76%), Positives = 94/114 (82%), Gaps = 1/114 (0%)
Frame = +2
Query: 217 MRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLFDRSGLLYGELARFVLRLLGIKTKP 276
MRVMQGVVNFFGRIS+LIDQNTQAFHMFM+ALLQLFDRSG+LYGELARFVLRLLGI+TKP
Sbjct: 5 MRVMQGVVNFFGRISMLIDQNTQAFHMFMTALLQLFDRSGVLYGELARFVLRLLGIRTKP 184
Query: 277 NKVNPQGPNGHMLGPNGHPLPGPHNSSANVNAIEGAKPA-SGGWDNVWGNDANQ 329
K+ P GP+G PLPG HNSS N N IEG KPA SG WDNVW NDA +
Sbjct: 185 KKIIPP-------GPDGLPLPGQHNSSVNQNFIEGTKPAPSGSWDNVWENDAKK 325
>BM527936 similar to GP|18700169|gb| AT3g07560/F21O3_27 {Arabidopsis
thaliana}, partial (20%)
Length = 355
Score = 108 bits (270), Expect = 3e-24
Identities = 51/60 (85%), Positives = 55/60 (91%)
Frame = +1
Query: 192 PYGGDQDPNNPYGGPPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQL 251
PYG + DPNNPYG P SPPGFWIS MRVMQGVVNFFGRIS+LIDQNTQAFHMFM+ALLQ+
Sbjct: 1 PYGAE-DPNNPYGVPSSPPGFWISFMRVMQGVVNFFGRISMLIDQNTQAFHMFMTALLQV 177
>TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fragment),
partial (40%)
Length = 522
Score = 83.2 bits (204), Expect = 1e-16
Identities = 43/87 (49%), Positives = 51/87 (58%)
Frame = +2
Query: 103 GGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYG 162
GG G + G G G +G +GGG+GGG+GGG GGG GG +G GG GGG G
Sbjct: 8 GGVGGGIGGGGGAGGG-FGGGHGGGVGGGIGGGAGGGGGAGGGFGGGA--GGGAGGGFGG 178
Query: 163 SSGGMYGGGMYGGGLGGPMGGYGGGMG 189
+GG GGG GG GG GG+GGG G
Sbjct: 179 GAGGGAGGGFGGGAGGGAGGGFGGGGG 259
Score = 78.6 bits (192), Expect = 4e-15
Identities = 44/88 (50%), Positives = 52/88 (59%), Gaps = 1/88 (1%)
Frame = +2
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLG-GGMYGGGMYGSSMYNRGGYGG 158
G GG G G+G G +G GGG+GGG GGG G GG +GGG G + GG+GG
Sbjct: 11 GVGGGIGGGGGAGGGFGGG-HGGGVGGGIGGGAGGGGGAGGGFGGGAGGGA---GGGFGG 178
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGG 186
G G +GG +GGG GGG GG GG GG
Sbjct: 179 GAGGGAGGGFGGGA-GGGAGGGFGGGGG 259
Score = 77.8 bits (190), Expect = 6e-15
Identities = 43/83 (51%), Positives = 48/83 (57%), Gaps = 1/83 (1%)
Frame = +2
Query: 114 GYGSGLYGSS-YGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGM 172
G G G+ G GGG GGG GGG+GGG+ GG G GG GGG G +GG GGG
Sbjct: 11 GVGGGIGGGGGAGGGFGGGHGGGVGGGIGGGAGGG------GGAGGGFGGGAGGGAGGGF 172
Query: 173 YGGGLGGPMGGYGGGMGVGPYGG 195
GG GG GG+GGG G G GG
Sbjct: 173 GGGAGGGAGGGFGGGAGGGAGGG 241
Score = 68.2 bits (165), Expect = 5e-12
Identities = 40/72 (55%), Positives = 44/72 (60%), Gaps = 1/72 (1%)
Frame = +2
Query: 125 GGGLGGGLGGGLG-GGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGG 183
GGG+GGG+GGG G GG +GGG G GG GGG G GG GGG GG GG GG
Sbjct: 5 GGGVGGGIGGGGGAGGGFGGGHGGGV---GGGIGGGAGG--GGGAGGGFGGGAGGGAGGG 169
Query: 184 YGGGMGVGPYGG 195
+GGG G G GG
Sbjct: 170 FGGGAGGGAGGG 205
Score = 35.4 bits (80), Expect = 0.036
Identities = 18/41 (43%), Positives = 21/41 (50%)
Frame = +2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGG 139
+GG G G+ + G G G G GG GG GGG GGG
Sbjct: 134 FGGGAGGGAGGGFGGGAGGGAGGGFGGGAGGGA-GGGFGGG 253
Score = 34.7 bits (78), Expect = 0.061
Identities = 22/50 (44%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +2
Query: 157 GGGLYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGGDQDPNNPYGG 205
GGG+ G GG GGG GG GG+GGG+G G GG GG
Sbjct: 5 GGGVGGGIGG-------GGGAGGGFGGGHGGGVGGGIGGGAGGGGGAGGG 133
>BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (25%)
Length = 407
Score = 81.3 bits (199), Expect = 6e-16
Identities = 50/106 (47%), Positives = 52/106 (48%)
Frame = +3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GGG GGG GGG GG + G GG GGG
Sbjct: 84 GGEGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGGGFLGGREXXGGGGEGGG 263
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GG GGG GGG G GG GGG G G GG + GG
Sbjct: 264 GGGGGGGGGGGGXGGGGGGRGEGGGGGGGGGGGGGGGGEGGGRRGG 401
Score = 79.7 bits (195), Expect = 2e-15
Identities = 51/111 (45%), Positives = 53/111 (46%), Gaps = 5/111 (4%)
Frame = +1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GGG GGG GGG GGG G GG GGG
Sbjct: 34 GGGGGGGGGGGGGGGGGGGRGGEGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGGGGGGG 213
Query: 160 LY-----GSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
++ GG GGG GGG GG GG GGG G G GG GG
Sbjct: 214 VF*GGERXREGGERGGGGGGGGGGGGGGGXGGGGGGGGRGGGGGGGGGGGG 366
Score = 79.0 bits (193), Expect = 3e-15
Identities = 52/108 (48%), Positives = 55/108 (50%), Gaps = 2/108 (1%)
Frame = +1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMY-GSSMYNRGG-YG 157
GG GG G G G G G GGG GGG GGG GGG GGG++ G GG G
Sbjct: 85 GGRGGEGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGGGG--GGGVF*GGERXREGGERG 258
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG G GG GGG GGG GG GG GGG G G GG+ GG
Sbjct: 259 GGGGGGGGGGGGGGXGGGGGGGGRGGGGGGGGGGGGGGEGREEGGGGG 402
Score = 76.6 bits (187), Expect = 1e-14
Identities = 49/106 (46%), Positives = 50/106 (46%)
Frame = +3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GG GGG GGG GGG G + R GGG
Sbjct: 69 GGGGGGGEGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGGGFLGGREXXGGG 248
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GG GGG GGG GG GG G G G G GG GG
Sbjct: 249 GEGGGGGGGGGGGGGGGXGGGGGGRGEGGGGGGGGGGGGGGGGEGG 386
Score = 76.6 bits (187), Expect = 1e-14
Identities = 49/106 (46%), Positives = 53/106 (49%)
Frame = +1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GGG GGG GGG++ GG RGG GGG
Sbjct: 94 GGEGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGGGGGGGVF*GGERXREGGERGGGGGG 273
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GG GG GGG GG GG GGG G G G ++ GG
Sbjct: 274 --GGGGGGGGGXGGGGGGGGRGGGGGGGGGGGGGGEGREEGGGGGG 405
Score = 76.3 bits (186), Expect = 2e-14
Identities = 51/114 (44%), Positives = 52/114 (44%), Gaps = 8/114 (7%)
Frame = +3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GGG GGG GGG GGG G GG GGG
Sbjct: 33 GGGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGGGGGGGGGRGGGGGGGGGGGGGGGGGG 212
Query: 160 LY--------GSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
+ G G GGG GGG GG GG GGG G G GG GG
Sbjct: 213 GFLGGREXXGGGGEGGGGGGGGGGGGGGGXGGGGGGRGEGGGGGGGGGGGGGGG 374
Score = 74.3 bits (181), Expect = 7e-14
Identities = 51/113 (45%), Positives = 52/113 (45%), Gaps = 8/113 (7%)
Frame = +3
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGL 160
G GG G G G G G GGG GGG GGG GGG GGG RGG GGG
Sbjct: 21 GGGGGGGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGGGGGGGG-------GRGGGGGGG 179
Query: 161 YGSSGGMYGGGMY--------GGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GG GGG + GGG GG GG GGG G G GG GG
Sbjct: 180 GGGGGGGGGGGGFLGGREXXGGGGEGGGGGGGGGGGGGGGXGGGGGGRGEGGG 338
Score = 73.2 bits (178), Expect = 2e-13
Identities = 52/117 (44%), Positives = 54/117 (45%), Gaps = 11/117 (9%)
Frame = +2
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G G G G GGG GGG GGG GGG GGG G GG GGG
Sbjct: 23 GGGGGGGGGGGGGGGGGGG--GGGGGGGRGGGGGGGGGGGGGGGGGKG------GGGGGG 178
Query: 160 LYGSSGGMYGGGMY-----------GGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GG GGG + GGG GG GG GGG G G GG+ GG
Sbjct: 179 GGGGGGGGGGGGFFRGERXXGRGGRGGGGGGGGGGGGGGGGGGGGGGEGGGGGGGGG 349
Score = 71.2 bits (173), Expect = 6e-13
Identities = 45/80 (56%), Positives = 45/80 (56%)
Frame = +3
Query: 116 GSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGG 175
G GL G GGG GGG GGG GGG GGG G GG GGG G GG GGG GG
Sbjct: 6 GRGLPGGGGGGGGGGGGGGGGGGG--GGGGEGGG----GGGGGGGGGGGGGGGGGGRGGG 167
Query: 176 GLGGPMGGYGGGMGVGPYGG 195
G GG GG GGG G G GG
Sbjct: 168 GGGGGGGGGGGGGGGGFLGG 227
Score = 67.4 bits (163), Expect = 8e-12
Identities = 40/77 (51%), Positives = 44/77 (56%)
Frame = +2
Query: 121 GSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGP 180
G + G GGG GGG GGG GGG G RGG GGG G GG GGG GGG GG
Sbjct: 5 GERFARGGGGGGGGGGGGGGGGGGGGGGGGGGRGGGGGG-GGGGGGGGGGGKGGGGGGGG 181
Query: 181 MGGYGGGMGVGPYGGDQ 197
GG GGG G G + G++
Sbjct: 182 GGGGGGGGGGGFFRGER 232
Score = 67.4 bits (163), Expect = 8e-12
Identities = 49/111 (44%), Positives = 49/111 (44%), Gaps = 8/111 (7%)
Frame = +2
Query: 103 GGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYG 162
GG G G G G GGG GGG GGG GGG GGG G GG GGG G
Sbjct: 2 GGERFARGGGGGGGGGGGGGGGGGGGGGGGGGGRGGG--GGGGGGGGGGGGGGKGGGGGG 175
Query: 163 SSGGMYGGGMYGG--------GLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG GGG GG G GG GG GGG G G GG GG
Sbjct: 176 GGGGGGGGGGGGGFFRGERXXGRGGRGGGGGGGGGGGGGGGGGGGGGGEGG 328
Score = 53.9 bits (128), Expect = 1e-07
Identities = 40/84 (47%), Positives = 40/84 (47%)
Frame = +3
Query: 129 GGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGM 188
G GL GG GGG GGG GG GGG G GG GGG GGG GG GG GGG
Sbjct: 6 GRGLPGGGGGGGGGGG---------GGGGGG--GGGGGGEGGG--GGGGGGGGGGGGGGG 146
Query: 189 GVGPYGGDQDPNNPYGGPPSPPGF 212
G G GG GG GF
Sbjct: 147 GGGRGGGGGGGGGGGGGGGGGGGF 218
Score = 51.2 bits (121), Expect = 6e-07
Identities = 39/89 (43%), Positives = 40/89 (44%)
Frame = +2
Query: 125 GGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGY 184
GGG GGG GGG GGG GGG GGG GGG GGG GG GG
Sbjct: 23 GGGGGGGGGGGGGGGGGGGG------------GGG---------GGGRGGGGGGG--GGG 133
Query: 185 GGGMGVGPYGGDQDPNNPYGGPPSPPGFW 213
GGG G G GG GG GF+
Sbjct: 134 GGGGGGGKGGGGGGGGGGGGGGGGGGGFF 220
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/62 (43%), Positives = 28/62 (44%)
Frame = +2
Query: 94 EQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNR 153
E+ GG GG G G G G G GG GGG GGG GGG GGG G
Sbjct: 227 ERXXGRGGRGGGGGGGGGGGGGGGGGGGGGGEGGGGGGGGGGGGGG--GGGRGGRREEGG 400
Query: 154 GG 155
GG
Sbjct: 401 GG 406
>TC219649 weakly similar to UP|Q95JD1 (Q95JD1) Basic proline-rich protein,
partial (37%)
Length = 750
Score = 81.3 bits (199), Expect = 6e-16
Identities = 54/116 (46%), Positives = 60/116 (51%), Gaps = 8/116 (6%)
Frame = -1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMY-NRGGYGG 158
GGAG G +G GSG YG GGG+G G GG GG GGG GS + GG GG
Sbjct: 438 GGAGPGGGEPGSGAGPGSGGYGGGPGGGVGPGGGGPGGGAGRGGGGPGSGAGPSGGGPGG 259
Query: 159 GLYGSS--GGMYGGGMYGGGLGGPMGGYGGG-----MGVGPYGGDQDPNNPYGGPP 207
G YG GG GG YGGG GG G GGG G GPYGG ++ +G P
Sbjct: 258 GSYGGEPGGGGPGGDGYGGGPGGGAGPDGGGPKDGGFGGGPYGGYGPRSHGFGVKP 91
Score = 76.3 bits (186), Expect = 2e-14
Identities = 51/111 (45%), Positives = 54/111 (47%), Gaps = 3/111 (2%)
Frame = -1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG G G G G+G G SYGGG GG G G GGG GS+ GGYGGG
Sbjct: 717 GGGGPVGGGYGGGPGGGAGSGGGSYGGGPGGDAGPG------GGGPGGSAGPGSGGYGGG 556
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMG--VGPYGGDQDPN-NPYGGPP 207
G +G GGG G G G GGYGGG G GP GG P GG P
Sbjct: 555 PGGDTGP--GGGEPGSGAGPGSGGYGGGPGGDAGPGGGGPGGGAGPGGGEP 409
Score = 72.8 bits (177), Expect = 2e-13
Identities = 56/128 (43%), Positives = 60/128 (46%), Gaps = 21/128 (16%)
Frame = -1
Query: 101 GAGGYGSTMNYNSGYGSGLYGSS-------YGGGLGGGLG---GGLGGGMY-GGGMYGSS 149
G+GGYG ++G G G GS YGGG GG G GG GGG GGG GS
Sbjct: 579 GSGGYGGGPGGDTGPGGGEPGSGAGPGSGGYGGGPGGDAGPGGGGPGGGAGPGGGEPGSG 400
Query: 150 MY-NRGGYGGGLYGSSGGMYGG-----GMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNP- 202
GGYGGG G G GG G GGG G G GGG G G YGG+ P
Sbjct: 399 AGPGSGGYGGGPGGGVGPGGGGPGGGAGRGGGGPGSGAGPSGGGPGGGSYGGEPGGGGPG 220
Query: 203 ---YGGPP 207
YGG P
Sbjct: 219 GDGYGGGP 196
Score = 72.0 bits (175), Expect = 3e-13
Identities = 60/140 (42%), Positives = 66/140 (46%), Gaps = 26/140 (18%)
Frame = -1
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYGGGLG--------------GGLGGGLGGGM--YGGG 144
G+GGYG ++G G G GGG G GG GGG GGG+ GGG
Sbjct: 501 GSGGYGGGPGGDAGPGGG----GPGGGAGPGGGEPGSGAGPGSGGYGGGPGGGVGPGGGG 334
Query: 145 MYGSSMYNRGGYGGGLYGSSGGMYGGGMYGG--GLGGPMG-GYGGGM--GVGPYGGDQDP 199
G + GG G G G SGG GGG YGG G GGP G GYGGG G GP GG
Sbjct: 333 PGGGAGRGGGGPGSGA-GPSGGGPGGGSYGGEPGGGGPGGDGYGGGPGGGAGPDGGGPKD 157
Query: 200 ----NNPYGG-PPSPPGFWI 214
PYGG P GF +
Sbjct: 156 GGFGGGPYGGYGPRSHGFGV 97
Score = 71.6 bits (174), Expect = 4e-13
Identities = 49/105 (46%), Positives = 52/105 (48%), Gaps = 8/105 (7%)
Frame = -1
Query: 111 YNSGYGSGLYGSSYGGGLGGGLGGGLGGGM-YGGGMYGSSMYNRGGYGGGLYGSSGGMYG 169
Y G G G+ G GG +GGG GGG GGG GGG YG G GGG G S G G
Sbjct: 750 YGGGPGGGV-GPGGGGPVGGGYGGGPGGGAGSGGGSYGGGPGGDAGPGGGGPGGSAGP-G 577
Query: 170 GGMYGGGLGGPMG------GYGGGMGVGPYGGDQDPN-NPYGGPP 207
G YGGG GG G G G G G G YGG + P GG P
Sbjct: 576 SGGYGGGPGGDTGPGGGEPGSGAGPGSGGYGGGPGGDAGPGGGGP 442
Score = 71.2 bits (173), Expect = 6e-13
Identities = 50/117 (42%), Positives = 55/117 (46%), Gaps = 9/117 (7%)
Frame = -1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG G GS + GYG G G + GG G G G G G YGGG G + GG GGG
Sbjct: 612 GGGGPGGSAGPGSGGYGGGPGGDTGPGGGEPGSGAGPGSGGYGGGPGGDAGPGGGGPGGG 433
Query: 160 LYGSSGGMYGGGM------YGGGLGGPMGGYGGGMGVGPYGGDQDPNN---PYGGPP 207
G GG G G YGGG GG +G GGG G G G P + P GG P
Sbjct: 432 A-GPGGGEPGSGAGPGSGGYGGGPGGGVGPGGGGPGGGAGRGGGGPGSGAGPSGGGP 265
Score = 67.8 bits (164), Expect = 6e-12
Identities = 50/113 (44%), Positives = 53/113 (46%), Gaps = 6/113 (5%)
Frame = -1
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYG---GGLGGGLGG--GLGGGMYGGGMYGSSMYNRGG 155
G G YG ++G G G G S G GG GGG GG G GGG G G S GG
Sbjct: 657 GGGSYGGGPGGDAGPGGGGPGGSAGPGSGGYGGGPGGDTGPGGGEPGSGAGPGS----GG 490
Query: 156 YGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPN-NPYGGPP 207
YGGG G +G GG G G GG G G G G G YGG P GG P
Sbjct: 489 YGGGPGGDAGPGGGGPGGGAGPGGGEPGSGAGPGSGGYGGGPGGGVGPGGGGP 331
Score = 38.9 bits (89), Expect = 0.003
Identities = 25/63 (39%), Positives = 28/63 (43%), Gaps = 3/63 (4%)
Frame = -1
Query: 98 TYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGL---GGGLGGGMYGGGMYGSSMYNRG 154
+YGG G G G G YGGG GGG GGG G +GGG YG
Sbjct: 255 SYGGEPG-----------GGGPGGDGYGGGPGGGAGPDGGGPKDGGFGGGPYGGYGPRSH 109
Query: 155 GYG 157
G+G
Sbjct: 108 GFG 100
Score = 28.9 bits (63), Expect = 3.3
Identities = 13/25 (52%), Positives = 13/25 (52%)
Frame = +2
Query: 25 PSNPPPKPWQQGGSSSGPTAPFKPP 49
P PPPKP G SGP P PP
Sbjct: 131 P*GPPPKPPSLGPPPSGPAPPPGPP 205
>TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein,
complete
Length = 748
Score = 77.0 bits (188), Expect = 1e-14
Identities = 51/107 (47%), Positives = 53/107 (48%)
Frame = +1
Query: 106 GSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSG 165
G + N G G GGG GGG GGG GGG GGG G YNR G GGG YG
Sbjct: 292 GRNITVNEAQSRG--GGGGGGGGGGGYGGGRGGGYGGGGRGGGGGYNRSGGGGG-YG--- 453
Query: 166 GMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGF 212
GGG YGGG G GGYGGG G YGG D GG G+
Sbjct: 454 ---GGGGYGGGGG---GGYGGGRDRG-YGGGGDRGYSRGGDGGDGGW 573
Score = 50.1 bits (118), Expect = 1e-06
Identities = 38/76 (50%), Positives = 39/76 (51%), Gaps = 5/76 (6%)
Frame = +1
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYG----GGLGG-GLGGGLGGGMYGGGMYGSSMYNRG 154
GG GGYG GYG G G G GG GG G GGG GGG GGG YG +RG
Sbjct: 349 GGGGGYGG--GRGGGYGGGGRGGGGGYNRSGGGGGYGGGGGYGGG--GGGGYGGGR-DRG 513
Query: 155 GYGGGLYGSSGGMYGG 170
GGG G S G GG
Sbjct: 514 YGGGGDRGYSRGGDGG 561
>TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (14%)
Length = 642
Score = 76.6 bits (187), Expect = 1e-14
Identities = 56/114 (49%), Positives = 56/114 (49%), Gaps = 8/114 (7%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLG--GGLGGGMYGGGMYGSSMYNRGGYG 157
G AGG GS GYG G GS GGG GG G GG GG GG GS GG G
Sbjct: 625 GSAGGGGS--KGGGGYGGG--GSQGGGGSAGGGGS*GGSGGSAGGGEGSGSQGGGSGGGG 458
Query: 158 --GGLYGSSGGMYG----GGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG GS GG YG GG GGG G GG GGG G G YGG D GG
Sbjct: 457 SQGGGSGSGGGGYGGGGSGGSEGGGSSGGGGGGGGGGGGGGYGGGGDKGGGIGG 296
Score = 74.3 bits (181), Expect = 7e-14
Identities = 51/113 (45%), Positives = 55/113 (48%), Gaps = 7/113 (6%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGG----GLGGGLGGGMYGGGMYGSSM---YN 152
G AGG GS G G S GGG GG G G G GGG YGGG G S +
Sbjct: 556 GSAGGGGS*GGSGGSAGGGEGSGSQGGGSGGGGSQGGGSGSGGGGYGGGGSGGSEGGGSS 377
Query: 153 RGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG GGG G GG GGG GGG+GG G Y G G GD+D + GG
Sbjct: 376 GGGGGGGGGGGGGGYGGGGDKGGGIGGDKGDYHHGKG---GKGDKDGGDKGGG 227
Score = 73.2 bits (178), Expect = 2e-13
Identities = 57/118 (48%), Positives = 60/118 (50%), Gaps = 6/118 (5%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGG--MYGSSMYNRGGYG 157
GG GG GS S G G G S GG GGG G G GG GGG G S GGYG
Sbjct: 592 GGYGGGGSQGGGGSAGGGGS*GGS-GGSAGGGEGSGSQGGGSGGGGSQGGGSGSGGGGYG 416
Query: 158 GGLYGSS--GGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGGDQ-DPNNPYGGPPSPPG 211
GG G S GG GGG GGG GG GGYGGG G GGD+ D ++ GG G
Sbjct: 415 GGGSGGSEGGGSSGGGGGGGG-GGGGGGYGGGGDKGGGIGGDKGDYHHGKGGKGDKDG 245
Score = 68.2 bits (165), Expect = 5e-12
Identities = 46/101 (45%), Positives = 48/101 (46%), Gaps = 3/101 (2%)
Frame = -3
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGL--GGGLGGGMYGGGM-YGSSMYNRGG 155
YGG G G + G G G S GGG G G GG GGG GGG G Y GG
Sbjct: 586 YGGGGSQGGGGSAGGGGS*GGSGGSAGGGEGSGSQGGGSGGGGSQGGGSGSGGGGYGGGG 407
Query: 156 YGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGD 196
GG G S G GGG GGG GG GG G G+G GD
Sbjct: 406 SGGSEGGGSSGGGGGGGGGGGGGGYGGGGDKGGGIGGDKGD 284
Score = 67.0 bits (162), Expect = 1e-11
Identities = 42/92 (45%), Positives = 44/92 (47%)
Frame = -3
Query: 114 GYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMY 173
G +G GS GGG GGG G GG GGG G S + GG G G GG GGG
Sbjct: 628 GGSAGGGGSKGGGGYGGGGSQGGGGSAGGGGS*GGSGGSAGG-GEGSGSQGGGSGGGGSQ 452
Query: 174 GGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GGG G GGYGGG G GG GG
Sbjct: 451 GGGSGSGGGGYGGGGSGGSEGGGSSGGGGGGG 356
Score = 62.8 bits (151), Expect = 2e-10
Identities = 43/92 (46%), Positives = 48/92 (51%), Gaps = 5/92 (5%)
Frame = -3
Query: 100 GGAGGYGS----TMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGG 155
GG+GG GS + + GYG G G S GGG GG GGG GGG GGG YG G
Sbjct: 478 GGSGGGGSQGGGSGSGGGGYGGGGSGGSEGGGSSGGGGGGGGGG--GGGGYGGG----GD 317
Query: 156 YGGGLYGSSGGM-YGGGMYGGGLGGPMGGYGG 186
GGG+ G G +G G G GG GG GG
Sbjct: 316 KGGGIGGDKGDYHHGKGGKGDKDGGDKGGGGG 221
Score = 62.0 bits (149), Expect = 4e-10
Identities = 40/86 (46%), Positives = 42/86 (48%), Gaps = 3/86 (3%)
Frame = -3
Query: 123 SYGGGLGGGLGGGLGGGMYGGG--MYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGL-GG 179
S GGG G GG GGG YGGG G GG GG GS+GG G G GGG GG
Sbjct: 640 SEGGGGSAGGGGSKGGGGYGGGGSQGGGGSAGGGGS*GGSGGSAGGGEGSGSQGGGSGGG 461
Query: 180 PMGGYGGGMGVGPYGGDQDPNNPYGG 205
G G G G G YGG + GG
Sbjct: 460 GSQGGGSGSGGGGYGGGGSGGSEGGG 383
>TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein-1, partial
(77%)
Length = 883
Score = 71.6 bits (174), Expect = 4e-13
Identities = 54/113 (47%), Positives = 56/113 (48%), Gaps = 16/113 (14%)
Frame = +3
Query: 95 QNTTYGGAGG--YGSTMNYNSGYGSGLYGSS--YGGGLGGGLGGGLGGGMYGGGMYGSSM 150
Q T GG GG YG G G G YG YGGG G G GGG GGG GGG G +
Sbjct: 276 QGTRRGGDGGRSYGG----GRGGGGGGYGGGGGYGGGGGYGGGGGYGGGGRGGG--GGAC 437
Query: 151 YN-----------RGGYGGGLYGSSGGMYGGGMYGGGLGGP-MGGYGGGMGVG 191
YN GG GG YG GG GG YGGG GG +GG GGG G G
Sbjct: 438 YNCGESGHLARDCSGGGGGDRYGGGGG---GGRYGGGGGGRYVGGGGGGGGGG 587
Score = 64.3 bits (155), Expect = 7e-11
Identities = 46/90 (51%), Positives = 46/90 (51%), Gaps = 15/90 (16%)
Frame = +3
Query: 121 GSSYGGGLGGGLGGGLGGGMYGGG--MYGSSMYNRGGYGGG-----LYGSSG-------G 166
G SYGGG GGG GG GGG YGGG G Y GG GGG G SG G
Sbjct: 303 GRSYGGGRGGGGGGYGGGGGYGGGGGYGGGGGYGGGGRGGGGGACYNCGESGHLARDCSG 482
Query: 167 MYGGGMY-GGGLGGPMGGYGGGMGVGPYGG 195
GG Y GGG GG GG GGG VG GG
Sbjct: 483 GGGGDRYGGGGGGGRYGGGGGGRYVGGGGG 572
Score = 60.5 bits (145), Expect = 1e-09
Identities = 43/98 (43%), Positives = 46/98 (46%), Gaps = 18/98 (18%)
Frame = +3
Query: 116 GSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGG 175
G+ + G+ GG G GGG GG GGG YG GGYGGG GG YGGG GG
Sbjct: 264 GASVQGTRRGGDGGRSYGGGRGG---GGGGYGGG----GGYGGGGGYGGGGGYGGGGRGG 422
Query: 176 GLG----------------GPMGG--YGGGMGVGPYGG 195
G G G GG YGGG G G YGG
Sbjct: 423 GGGACYNCGESGHLARDCSGGGGGDRYGGGGGGGRYGG 536
Score = 53.5 bits (127), Expect = 1e-07
Identities = 39/97 (40%), Positives = 39/97 (40%), Gaps = 16/97 (16%)
Frame = +3
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGL--------------GGGLGGGLGGGMYGGG 144
YGG GGYG Y G G G G GGG GG GG GG GGG
Sbjct: 345 YGGGGGYGGGGGYGGGGGYGGGGRGGGGGACYNCGESGHLARDCSGGGGGDRYGGGGGGG 524
Query: 145 MYGSSMYNRGGYGGGLY--GSSGGMYGGGMYGGGLGG 179
Y GG GGG Y G GG GG Y G G
Sbjct: 525 RY-------GGGGGGRYVGGGGGGGGGGSCYSCGESG 614
Score = 45.1 bits (105), Expect = 4e-05
Identities = 25/52 (48%), Positives = 27/52 (51%)
Frame = +3
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G G + G+ G GG YGGG GG GGYGGG G G GG YGG
Sbjct: 255 GPDGASVQGTRRGGDGGRSYGGGRGGGGGGYGGGGGYGG-GGGYGGGGGYGG 407
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 69.7 bits (169), Expect = 2e-12
Identities = 49/106 (46%), Positives = 49/106 (46%), Gaps = 11/106 (10%)
Frame = +3
Query: 95 QNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGG--GLGGGLGGGMYGGGMYGSSMYN 152
Q T GG GS Y G G G YG GG GG G GGG GGG YGGG
Sbjct: 45 QGTRGGGGXXRGS---YGXGGGRGGYGGGGRGGRGGYGGGGGGYGGGGYGGG-------- 191
Query: 153 RGGYGGGLYGSSGGMYGGG---------MYGGGLGGPMGGYGGGMG 189
GGYGGG G GG Y G GGG GG GG GGG G
Sbjct: 192 -GGYGGGGGGGGGGCYKCGETGHIARDCSQGGGGGGRYGGGGGGGG 326
Score = 62.0 bits (149), Expect = 4e-10
Identities = 41/79 (51%), Positives = 43/79 (53%), Gaps = 1/79 (1%)
Frame = +3
Query: 117 SGLYGSSYGGGLG-GGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGG 175
S + G+ GGG G G G G G YGGG G RGGYGGG GG YGGG YGG
Sbjct: 36 SPVQGTRGGGGXXRGSYGXGGGRGGYGGGGRGG----RGGYGGG-----GGGYGGGGYGG 188
Query: 176 GLGGPMGGYGGGMGVGPYG 194
G GGYGGG G G G
Sbjct: 189 G-----GGYGGGGGGGGGG 230
Score = 57.8 bits (138), Expect = 7e-09
Identities = 36/63 (57%), Positives = 36/63 (57%)
Frame = +3
Query: 133 GGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGP 192
GGG G YG G RGGYGGG G GG YGGG GGG GG GGYGGG G G
Sbjct: 60 GGGXXRGSYGXGG------GRGGYGGGGRGGRGG-YGGG--GGGYGG--GGYGGGGGYGG 206
Query: 193 YGG 195
GG
Sbjct: 207 GGG 215
Score = 48.5 bits (114), Expect = 4e-06
Identities = 37/87 (42%), Positives = 37/87 (42%), Gaps = 21/87 (24%)
Frame = +3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSS--YGGGLGGGLGG-----------------GLGGGM 140
GG GGYG GYG G YG YGGG GGG GG G GGG
Sbjct: 129 GGRGGYGGG---GGGYGGGGYGGGGGYGGGGGGGGGGCYKCGETGHIARDCSQGGGGGGR 299
Query: 141 YGGGMYGSSMYNRGGYGGGLY--GSSG 165
YGGG GG GG Y G SG
Sbjct: 300 YGGG---------GGGGGSCYNCGESG 353
Score = 35.8 bits (81), Expect = 0.027
Identities = 26/61 (42%), Positives = 26/61 (42%), Gaps = 12/61 (19%)
Frame = +3
Query: 99 YGGAGGYGSTMNYNSGYGSGLY----------GSSYGGGLGG--GLGGGLGGGMYGGGMY 146
YGG GGYG G G G Y S GGG GG G GGG GG Y G
Sbjct: 180 YGGGGGYGGG---GGGGGGGCYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGES 350
Query: 147 G 147
G
Sbjct: 351 G 353
Score = 33.1 bits (74), Expect = 0.18
Identities = 26/60 (43%), Positives = 27/60 (44%), Gaps = 7/60 (11%)
Frame = +3
Query: 153 RGGYGGGLYGSSGGMYGGGMYGGGLG-------GPMGGYGGGMGVGPYGGDQDPNNPYGG 205
+G GGG G G YG G GG G G GGYGGG G G GG YGG
Sbjct: 45 QGTRGGG--GXXRGSYGXG---GGRGGYGGGGRGGRGGYGGG-GGGYGGGGYGGGGGYGG 206
>TC230575 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprotein-like
{Arabidopsis thaliana;} , partial (21%)
Length = 788
Score = 68.6 bits (166), Expect = 4e-12
Identities = 47/112 (41%), Positives = 54/112 (47%), Gaps = 14/112 (12%)
Frame = +2
Query: 98 TYGGAGGYGSTMNYNSGYGSGLY---GSSYGG--GLGGGLGGGLGGGMYG-GGMYGSSM- 150
++G GGY S Y G GSG Y S +GG G G +G G G G YG S
Sbjct: 107 SFGMGGGYRSGGAYGGGRGSGAYXXYASEFGGYGGYAGAMGPYRGDPSLGYTGRYGGSFS 286
Query: 151 --YNRGGYGG-----GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
Y+ GGYGG G YG+ GG GG G G G GY G +G G YGG
Sbjct: 287 RGYDLGGYGGPSENYGAYGAGGGSSGGAGAGAGAGAYQSGYDGNLG-GGYGG 439
Score = 55.8 bits (133), Expect = 3e-08
Identities = 41/109 (37%), Positives = 44/109 (39%), Gaps = 13/109 (11%)
Frame = +2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLG-----GGLGG--------GLGGGMYGGGM 145
+GG GGY M G S Y YGG GG GG G GGG GG
Sbjct: 194 FGGYGGYAGAMGPYRGDPSLGYTGRYGGSFSRGYDLGGYGGPSENYGAYGAGGGSSGGAG 373
Query: 146 YGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYG 194
G+ G Y G G+ GG YGG G G GGY G PYG
Sbjct: 374 AGAGA---GAYQSGYDGNLGGGYGGASGGPFYGSTRGGYSAGR-YHPYG 508
Score = 55.5 bits (132), Expect = 3e-08
Identities = 48/138 (34%), Positives = 52/138 (36%), Gaps = 34/138 (24%)
Frame = +2
Query: 107 STMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGM-------YGSSMYNRGGYGGG 159
S +Y+SG G G +GG G G GG GG YGGG Y S GGY G
Sbjct: 50 SRSSYSSG-GXGDAXXGFGGSFGMG-GGYRSGGAYGGGRGSGAYXXYASEFGGYGGYAGA 223
Query: 160 LYGSSG-------GMYGG------------------GMYGGGLGGPMGGYGGGMGVGPYG 194
+ G G YGG G YG G GG GG G G G G Y
Sbjct: 224 MGPYRGDPSLGYTGRYGGSFSRGYDLGGYGGPSENYGAYGAG-GGSSGGAGAGAGAGAYQ 400
Query: 195 GDQDPN--NPYGGPPSPP 210
D N YGG P
Sbjct: 401 SGYDGNLGGGYGGASGGP 454
Score = 53.9 bits (128), Expect = 1e-07
Identities = 41/113 (36%), Positives = 47/113 (41%), Gaps = 15/113 (13%)
Frame = +2
Query: 107 STMNYNSGYGSGLYGSSYGGGLGGGLGG--GLGGGMYGGGMYGSSMYNRGGYGGGLYGSS 164
S+ YN S Y S G G GG G+GGG GG YG GG G G Y
Sbjct: 29 SSKRYNDSRSS--YSSGGXGDAXXGFGGSFGMGGGYRSGGAYG------GGRGSGAYXXY 184
Query: 165 GGMYGG-GMYGGGLG--------GPMGGYGG----GMGVGPYGGDQDPNNPYG 204
+GG G Y G +G G G YGG G +G YGG + YG
Sbjct: 185 ASEFGGYGGYAGAMGPYRGDPSLGYTGRYGGSFSRGYDLGGYGGPSENYGAYG 343
Score = 53.9 bits (128), Expect = 1e-07
Identities = 38/98 (38%), Positives = 47/98 (47%), Gaps = 4/98 (4%)
Frame = +2
Query: 92 PWEQNTTYGGAGGYGSTMNYNSGYGSGLYGS---SYGG-GLGGGLGGGLGGGMYGGGMYG 147
P+ + + G G YG + ++ GY G YG +YG G GGG GG G G G G Y
Sbjct: 230 PYRGDPSLGYTGRYGGS--FSRGYDLGGYGGPSENYGAYGAGGGSSGGAGAGA-GAGAYQ 400
Query: 148 SSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG 185
S G GGG G+SGG + G GG G YG
Sbjct: 401 SGY--DGNLGGGYGGASGGPFYGSTRGGYSAGRYHPYG 508
>TC205429 weakly similar to UP|O22600 (O22600) Glycine-rich protein, partial
(49%)
Length = 611
Score = 68.2 bits (165), Expect = 5e-12
Identities = 56/150 (37%), Positives = 77/150 (51%), Gaps = 1/150 (0%)
Frame = -2
Query: 98 TYGGAGGYGSTMNYNSGYGS-GLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGY 156
+YGG G Y SG GS GL G G+GGG GGGLGGG+ GG+ GG
Sbjct: 523 SYGGVGAY-------SGIGSNGLPLGGVGAGIGGGFGGGLGGGL--GGL--------GGT 395
Query: 157 GGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGFWISV 216
GG+ G G GGG+ GGG+G G GGG GV P+ + + +G P F+ S
Sbjct: 394 TGGIGGFGG--TGGGVGGGGVG---TGVGGGSGVLPFP*NDEHACMHG--CVAPSFFTSF 236
Query: 217 MRVMQGVVNFFGRISILIDQNTQAFHMFMS 246
V + +V F +S++ + +F +F+S
Sbjct: 235 FFVREIIVV*FVHVSVV---SRSSFLVFLS 155
>TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (15%)
Length = 494
Score = 67.0 bits (162), Expect = 1e-11
Identities = 39/73 (53%), Positives = 42/73 (57%), Gaps = 1/73 (1%)
Frame = +3
Query: 118 GLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYG-GGMYGGG 176
G +G YGG GG GGG GGG GG +G GGYGGG GG YG GG +GGG
Sbjct: 15 GEHGXGYGGA--GGSGGGSGGGYGDGGAHG------GGYGGGAGAGGGGGYGAGGAHGGG 170
Query: 177 LGGPMGGYGGGMG 189
GG GG GGG G
Sbjct: 171 FGGG-GGSGGGHG 206
Score = 61.2 bits (147), Expect = 6e-10
Identities = 39/77 (50%), Positives = 43/77 (55%)
Frame = +3
Query: 94 EQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNR 153
E YGGAGG G G G G YG GG GGG GGG G G GGG YG+ +
Sbjct: 18 EHGXGYGGAGGSGG------GSGGG-YGD--GGAHGGGYGGGAGAG--GGGGYGAGGAHG 164
Query: 154 GGYGGGLYGSSGGMYGG 170
GG+GGG G SGG +GG
Sbjct: 165 GGFGGG--GGSGGGHGG 209
Score = 48.1 bits (113), Expect = 5e-06
Identities = 33/70 (47%), Positives = 38/70 (54%), Gaps = 2/70 (2%)
Frame = +3
Query: 138 GGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLG-GPMGGYG-GGMGVGPYGG 195
GG +G G G+ GG GG YG GG +GGG YGGG G G GGYG GG G +GG
Sbjct: 12 GGEHGXGYGGAG--GSGGGSGGGYG-DGGAHGGG-YGGGAGAGGGGGYGAGGAHGGGFGG 179
Query: 196 DQDPNNPYGG 205
+GG
Sbjct: 180 GGGSGGGHGG 209
Score = 41.6 bits (96), Expect = 5e-04
Identities = 24/47 (51%), Positives = 26/47 (55%), Gaps = 2/47 (4%)
Frame = +3
Query: 151 YNRGGYGGGLYGSSGGMYG--GGMYGGGLGGPMGGYGGGMGVGPYGG 195
+ GG G YG +GG G GG YG G G GGYGGG G G GG
Sbjct: 3 HEAGGEHGXGYGGAGGSGGGSGGGYGDG-GAHGGGYGGGAGAGGGGG 140
>TC211031 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (11%)
Length = 678
Score = 67.0 bits (162), Expect = 1e-11
Identities = 46/104 (44%), Positives = 50/104 (47%), Gaps = 8/104 (7%)
Frame = -3
Query: 100 GGAGGYGSTMNYN-----SGYGSGLYGSSYGGGLGGGLGGGLGGGM---YGGGMYGSSMY 151
GG+GG G N SG G G +G GG GGG GG GGG+ GGG G
Sbjct: 442 GGSGGGGGASGGNGGGGASGGGDGRFGGEGDGGKGGGGDGGKGGGVDSGIGGGKGG---- 275
Query: 152 NRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GG GG G GG GGG G G GG G GGG +G GG
Sbjct: 274 --GGDGGNGGGGDGGNGGGGDGGNGGGGASGIGGGGHSLGGGGG 149
Score = 60.8 bits (146), Expect = 8e-10
Identities = 43/97 (44%), Positives = 47/97 (48%), Gaps = 8/97 (8%)
Frame = -3
Query: 103 GGYGSTMNYNSGYGSGLYGSSYGGGL-GGGLG-------GGLGGGMYGGGMYGSSMYNRG 154
G +G N SG G G G + GGG GGG G GG GGG GG G G
Sbjct: 466 GPFGGIGNGGSGGGGGASGGNGGGGASGGGDGRFGGEGDGGKGGGGDGGKGGGVDSGIGG 287
Query: 155 GYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVG 191
G GGG G +GG GG GGG GG GG G+G G
Sbjct: 286 GKGGGGDGGNGGGGDGGNGGGGDGGNGGGGASGIGGG 176
Score = 60.5 bits (145), Expect = 1e-09
Identities = 44/97 (45%), Positives = 47/97 (48%), Gaps = 1/97 (1%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGG-GMYGSSMYNRGGYGG 158
G GG G N G G G G + GG GGG GG G G +GG G G GG GG
Sbjct: 466 GPFGGIG-----NGGSGGG--GGASGGNGGGGASGG-GDGRFGGEGDGGKGGGGDGGKGG 311
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G+ GG GGG GG GG GG GGG G GG
Sbjct: 310 GVDSGIGGGKGGGGDGGNGGGGDGGNGGGGDGGNGGG 200
Score = 59.3 bits (142), Expect = 2e-09
Identities = 43/101 (42%), Positives = 47/101 (45%), Gaps = 4/101 (3%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
G G G Y+ G G+ GGG GG GG GGG GGG GG GGG
Sbjct: 508 GRFGSRGGGTRYSPGPFGGIGNGGSGGG-GGASGGNGGGGASGGGDGRFGGEGDGGKGGG 332
Query: 160 LYG-SSGGM---YGGGMYGGGLGGPMGGYGGGMGVGPYGGD 196
G GG+ GGG GGG GG GG GG G G GG+
Sbjct: 331 GDGGKGGGVDSGIGGGKGGGGDGGNGGGGDGGNGGGGDGGN 209
Score = 56.6 bits (135), Expect = 1e-08
Identities = 38/95 (40%), Positives = 44/95 (46%)
Frame = -3
Query: 111 YNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGG 170
+ S G Y GG+G G GG GG G G G+S G +GG G GG G
Sbjct: 502 FGSRGGGTRYSPGPFGGIGNGGSGGGGGASGGNGGGGASGGGDGRFGGEGDGGKGGGGDG 323
Query: 171 GMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G GGG+ +GG GG G G GG D N GG
Sbjct: 322 GK-GGGVDSGIGGGKGGGGDGGNGGGGDGGNGGGG 221
Score = 47.0 bits (110), Expect = 1e-05
Identities = 30/70 (42%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Frame = -3
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYG-G 158
GG G G +SG G G G GG GGG GG GGG G G G+S GG+ G
Sbjct: 340 GGGGDGGKGGGVDSGIGGGKGGGGDGGNGGGGDGGNGGGGDGGNGGGGASGIGGGGHSLG 161
Query: 159 GLYGSSGGMY 168
G GSS ++
Sbjct: 160 GGGGSSHSLH 131
Score = 47.0 bits (110), Expect = 1e-05
Identities = 34/80 (42%), Positives = 38/80 (47%)
Frame = -3
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G + G G G+ GG GGG GG GGG G G G GG GG
Sbjct: 367 FGGEGDGGKGGGGDGGKGGGVDSGIGGGKGGGGDGGNGGGGDGGNGGGGD-----GGNGG 203
Query: 159 GLYGSSGGMYGGGMYGGGLG 178
G G+SG GG GGG G
Sbjct: 202 G--GASGIGGGGHSLGGGGG 149
Score = 34.3 bits (77), Expect = 0.079
Identities = 27/64 (42%), Positives = 30/64 (46%), Gaps = 6/64 (9%)
Frame = -3
Query: 148 SSMYNRGGYGGGLYGSSGGMYGGGMYG--GGLGGPMGGYGG----GMGVGPYGGDQDPNN 201
SS R G GG S G +GG G GG GG GG GG G G G +GG+ D
Sbjct: 520 SSRSGRFGSRGGGTRYSPGPFGGIGNGGSGGGGGASGGNGGGGASGGGDGRFGGEGDGGK 341
Query: 202 PYGG 205
GG
Sbjct: 340 GGGG 329
>TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprotein-like
{Arabidopsis thaliana;} , partial (34%)
Length = 936
Score = 66.2 bits (160), Expect = 2e-11
Identities = 46/105 (43%), Positives = 51/105 (47%), Gaps = 6/105 (5%)
Frame = +2
Query: 107 STMNYN---SGYGSGLYGSSYGGGLGG-GLGGGLGGGMYGG--GMYGSSMYNRGGYGGGL 160
S+ YN S YG G YG +Y G G G+GG GG YGG YG GGYGG
Sbjct: 179 SSKRYNDSRSSYGGG-YGDAYDGFGGNFGMGGYRSGGAYGGRGSAYGGFGSEFGGYGG-- 349
Query: 161 YGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
Y + G Y G G G GGYG G +G YGG D YGG
Sbjct: 350 YAGAMGPYRGDPSLGYAGRYGGGYGRGYDLGGYGGPSDGYGAYGG 484
Score = 65.5 bits (158), Expect = 3e-11
Identities = 44/107 (41%), Positives = 47/107 (43%), Gaps = 1/107 (0%)
Frame = +2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLG-GGLGGGMYGGGMYGSSMYNRGGYG 157
+GG GGY M G S Y YGGG G G GG GG G G YG G G
Sbjct: 332 FGGYGGYAGAMGPYRGDPSLGYAGRYGGGYGRGYDLGGYGGPSDGYGAYG-------GVG 490
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYG 204
GG GS+ G GGG GGP GG G G G +PYG
Sbjct: 491 GGSSGSAYGSSYDASLGGGYGGPAGGSFYGTRGGYAGAGTGRYHPYG 631
Score = 62.0 bits (149), Expect = 4e-10
Identities = 51/132 (38%), Positives = 61/132 (45%), Gaps = 12/132 (9%)
Frame = +2
Query: 87 PVPTRPWEQNTTYGGAGGYGSTMN-YNSGYGSGLY--GSSYGGGLGGGLGGGLGGGMYGG 143
P R + ++YGG GYG + + +G G Y G +YGG G GG G G
Sbjct: 176 PSSKRYNDSRSSYGG--GYGDAYDGFGGNFGMGGYRSGGAYGG--RGSAYGGFGSEFGGY 343
Query: 144 GMYGSSM-YNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG--GGMGVG----PYGGD 196
G Y +M RG G G GG YG G GG GGP GYG GG+G G YG
Sbjct: 344 GGYAGAMGPYRGDPSLGYAGRYGGGYGRGYDLGGYGGPSDGYGAYGGVGGGSSGSAYGSS 523
Query: 197 QDPN--NPYGGP 206
D + YGGP
Sbjct: 524 YDASLGGGYGGP 559
Score = 45.1 bits (105), Expect = 4e-05
Identities = 31/69 (44%), Positives = 32/69 (45%), Gaps = 8/69 (11%)
Frame = +2
Query: 148 SSMYN--RGGYGGGL---YGSSGGMYGGGMY--GGGLGGPMGGYGG-GMGVGPYGGDQDP 199
S YN R YGGG Y GG +G G Y GG GG YGG G G YGG
Sbjct: 182 SKRYNDSRSSYGGGYGDAYDGFGGNFGMGGYRSGGAYGGRGSAYGGFGSEFGGYGGYAGA 361
Query: 200 NNPYGGPPS 208
PY G PS
Sbjct: 362 MGPYRGDPS 388
>TC205428 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (5%)
Length = 720
Score = 65.9 bits (159), Expect = 2e-11
Identities = 42/98 (42%), Positives = 49/98 (49%)
Frame = +1
Query: 98 TYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYG 157
+YGG GGY G G G +GG G G+GGG GGG GG G
Sbjct: 172 SYGGVGGYS---------GIGNNGLPFGGA-GAGIGGGFGGG--------------GGLG 279
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GGL G +GG+ G G G G+GG G GGG+G G GG
Sbjct: 280 GGLGGPTGGIGGFGGTGSGVGG---GIGGGVGTGVGGG 384
Score = 36.2 bits (82), Expect = 0.021
Identities = 25/58 (43%), Positives = 31/58 (53%)
Frame = +1
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPG 211
GGY G G++G +GG G G+GG GG GGG+G G GG +GG S G
Sbjct: 187 GGYSG--IGNNGLPFGGA--GAGIGGGFGG-GGGLG-GGLGGPTGGIGGFGGTGSGVG 342
>TC219986 weakly similar to UP|Q41187 (Q41187) Glycine-rich protein
(Fragment), partial (47%)
Length = 584
Score = 65.5 bits (158), Expect = 3e-11
Identities = 39/83 (46%), Positives = 46/83 (54%), Gaps = 2/83 (2%)
Frame = +2
Query: 104 GYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGS 163
GYG +SG G G YGGG G G GGLGGG GG G +Y G GGG+Y
Sbjct: 71 GYGGGAGGDSGLGGG-----YGGGAGSGGVGGLGGGSGLGGGIGGGIYKGIGIGGGIYKG 235
Query: 164 SG-GMYGGGMYG-GGLGGPMGGY 184
G G GGG+ GG+GG +GG+
Sbjct: 236 IGFGGVGGGVGAIGGIGGVIGGF 304
Score = 59.7 bits (143), Expect = 2e-09
Identities = 42/82 (51%), Positives = 46/82 (55%)
Frame = +2
Query: 114 GYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMY 173
GYG G G S GLGGG GGG G G GG GS + GG GGG+Y G GGG+Y
Sbjct: 71 GYGGGAGGDS---GLGGGYGGGAGSGGVGGLGGGSGL--GGGIGGGIYKGIG--IGGGIY 229
Query: 174 GGGLGGPMGGYGGGMGVGPYGG 195
G +G GG GGG VG GG
Sbjct: 230 KG-IG--FGGVGGG--VGAIGG 280
Score = 33.5 bits (75), Expect(2) = 0.001
Identities = 19/39 (48%), Positives = 19/39 (48%), Gaps = 1/39 (2%)
Frame = +2
Query: 168 YGGGMYG-GGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
YGGG G GLGG GG G GVG GG GG
Sbjct: 74 YGGGAGGDSGLGGGYGGGAGSGGVGGLGGGSGLGGGIGG 190
Score = 26.2 bits (56), Expect(2) = 0.001
Identities = 14/25 (56%), Positives = 17/25 (68%), Gaps = 4/25 (16%)
Frame = +1
Query: 121 GSSYGG--GLG--GGLGGGLGGGMY 141
G S GG G+G GG GGGLGG ++
Sbjct: 4 GGSIGGVGGIGDVGGAGGGLGGWLW 78
>TC214257 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (7%)
Length = 607
Score = 65.1 bits (157), Expect = 4e-11
Identities = 40/80 (50%), Positives = 41/80 (51%), Gaps = 3/80 (3%)
Frame = +2
Query: 137 GGGMYGGGMY---GSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPY 193
GGG YGGG G YNR G GGG YG GG GGG +GGG GGYGGG Y
Sbjct: 8 GGGGYGGGGRREGGGGGYNRNGGGGG-YGGGGGYGGGGGHGGG-----GGYGGGGRDRGY 169
Query: 194 GGDQDPNNPYGGPPSPPGFW 213
GGD GG S G W
Sbjct: 170 GGDGGSRYSRGGGASDGGSW 229
Score = 53.1 bits (126), Expect = 2e-07
Identities = 34/76 (44%), Positives = 36/76 (46%), Gaps = 2/76 (2%)
Frame = +2
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNR--GGYG 157
GG GG G GY G YGGG GG GG GGG GGG YG +R GG G
Sbjct: 14 GGYGGGGRREGGGGGYNRNGGGGGYGGG--GGYGG--GGGHGGGGGYGGGGRDRGYGGDG 181
Query: 158 GGLYGSSGGMYGGGMY 173
G Y GG GG +
Sbjct: 182 GSRYSRGGGASDGGSW 229
Score = 40.0 bits (92), Expect = 0.001
Identities = 25/57 (43%), Positives = 26/57 (44%)
Frame = +2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGG 155
YGG GGYG GGG GGG G G GG G G G S Y+RGG
Sbjct: 86 YGGGGGYGG-----------------GGGHGGGGGYGGGGRDRGYGGDGGSRYSRGG 205
Score = 33.9 bits (76), Expect = 0.10
Identities = 20/48 (41%), Positives = 22/48 (45%)
Frame = +2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMY 146
YGG GG+G GYG G YGG GG GGG GG +
Sbjct: 104 YGGGGGHGG----GGGYGGGGRDRGYGG--DGGSRYSRGGGASDGGSW 229
>TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (32%)
Length = 1539
Score = 64.7 bits (156), Expect = 5e-11
Identities = 50/124 (40%), Positives = 60/124 (48%), Gaps = 25/124 (20%)
Frame = -2
Query: 97 TTYGGAGGY--------GSTMNYNSGYG--------------SGLYGSSYG--GGLGGGL 132
TT+GG GG+ GST G G G GS++G GG GG
Sbjct: 587 TTFGGLGGFFGGFGG*GGSTFGGFGGCGGLTFGGHGGFFGGFGG*GGSTFGGFGGCGGLT 408
Query: 133 GGGLGGGMYG-GGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVG 191
GGLGG G GG G + GG GG +G GG+ GG +GGGL G GG+GGG G
Sbjct: 407 FGGLGGFFGGCGG*GGLTFGGFGG*GGSTFGGHGGLTGG--FGGGLTG--GGFGGGTTFG 240
Query: 192 PYGG 195
+GG
Sbjct: 239 GHGG 228
Score = 63.9 bits (154), Expect = 9e-11
Identities = 43/103 (41%), Positives = 54/103 (51%), Gaps = 8/103 (7%)
Frame = -2
Query: 101 GAGGYGSTMNYN-SGY----GSGLYGSSYGGGLGGGLGGGLGG-GMYGGGMYGSSMYNRG 154
G GG G + SG+ G G +G GGG GG+GG GG G GGG +G G
Sbjct: 764 GTGGSGGNLGIGGSGFLKIGGHGFFGG-VGGGSFGGVGGSFGGFGGRGGGFFGGF----G 600
Query: 155 GYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG--GGMGVGPYGG 195
G GG +G GG +GG +GG G GG+G GG+ G +GG
Sbjct: 599 GSGGTTFGGLGGFFGG--FGG*GGSTFGGFGGCGGLTFGGHGG 477
Score = 63.2 bits (152), Expect = 2e-10
Identities = 49/138 (35%), Positives = 59/138 (42%), Gaps = 25/138 (18%)
Frame = -2
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYG------GGLGGGLGG--GLGGGMYGG--GMYGSS 149
G GG G + G G G +G G GGLGG GG G GG +GG G G +
Sbjct: 674 GSFGGVGGSFGGFGGRGGGFFGGFGGSGGTTFGGLGGFFGGFGG*GGSTFGGFGGCGGLT 495
Query: 150 MYNRGGYGGGLYGSSGGMYGG-----GMYGGGLGGPMGGYGG----------GMGVGPYG 194
GG+ GG G G +GG G+ GGLGG GG GG G G +G
Sbjct: 494 FGGHGGFFGGFGG*GGSTFGGFGGCGGLTFGGLGGFFGGCGG*GGLTFGGFGG*GGSTFG 315
Query: 195 GDQDPNNPYGGPPSPPGF 212
G +GG + GF
Sbjct: 314 GHGGLTGGFGGGLTGGGF 261
Score = 59.3 bits (142), Expect = 2e-09
Identities = 46/116 (39%), Positives = 58/116 (49%), Gaps = 17/116 (14%)
Frame = -2
Query: 97 TTYGGAGGYGS-TMNYNSGYGSGLYG---SSYGG--GLGGGLGGGLGG---------GMY 141
+T+GG GG G T + G+ G G S++GG G GG GGLGG G+
Sbjct: 533 STFGGFGGCGGLTFGGHGGFFGGFGG*GGSTFGGFGGCGGLTFGGLGGFFGGCGG*GGLT 354
Query: 142 GGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGG--LGGPMGGYGGGMGVGPYGG 195
GG G GG+GG G GG+ GGG +GGG GG GG+ G G G + G
Sbjct: 353 FGGFGG*GGSTFGGHGGLTGGFGGGLTGGG-FGGGTTFGGHGGGFSGTGGGGFFSG 189
Score = 58.5 bits (140), Expect = 4e-09
Identities = 44/112 (39%), Positives = 55/112 (48%), Gaps = 19/112 (16%)
Frame = -2
Query: 103 GGYGSTMNYNSGYGSGLYGS---SYGG--GLGGGLGGGLG----------GGMYG--GGM 145
GG+G + G G G +G S+GG G GGG GG G GG +G GG
Sbjct: 707 GGHG----FFGGVGGGSFGGVGGSFGGFGGRGGGFFGGFGGSGGTTFGGLGGFFGGFGG* 540
Query: 146 YGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG--GGMGVGPYGG 195
GS+ GG GG +G GG +GG +GG G GG+G GG+ G GG
Sbjct: 539 GGSTFGGFGGCGGLTFGGHGGFFGG--FGG*GGSTFGGFGGCGGLTFGGLGG 390
Score = 58.5 bits (140), Expect = 4e-09
Identities = 49/123 (39%), Positives = 59/123 (47%), Gaps = 25/123 (20%)
Frame = -2
Query: 98 TYGGAGGY----GSTMNYNSGYGSGLYGSSYGG--GLGGGLGGGL-GGGMYGGGMYGSSM 150
T+GG GG+ G G G GS++GG GL GG GGGL GGG GG +
Sbjct: 410 TFGGLGGFFGGCGG*GGLTFGGFGG*GGSTFGGHGGLTGGFGGGLTGGGFGGGTTF---- 243
Query: 151 YNRGGYGGGLYGSSGGMY----GGGM----------YGGGL----GGPMGGYGGGMGVGP 192
GG+GGG G+ GG + GGG+ GGGL GG G GGM +G
Sbjct: 242 ---GGHGGGFSGTGGGGFFSGTGGGV*TGTGGGV*TGGGGLWKISGGSGGNVIGGMVIGG 72
Query: 193 YGG 195
GG
Sbjct: 71 *GG 63
Score = 58.2 bits (139), Expect = 5e-09
Identities = 41/101 (40%), Positives = 47/101 (45%), Gaps = 16/101 (15%)
Frame = -2
Query: 98 TYGGAGG-----YGSTMNYNSGYGSGLYGSSYGGGL-----GGGLGGGLGGGMYGGGMYG 147
T+GG GG +G G+G GL G +GGG GGG G GGG + G G
Sbjct: 356 TFGGFGG*GGSTFGGHGGLTGGFGGGLTGGGFGGGTTFGGHGGGFSGTGGGGFFSGTGGG 177
Query: 148 SSMYNRGGY---GGGLY---GSSGGMYGGGMYGGGLGGPMG 182
GG GGGL+ G SGG GGM GG GG G
Sbjct: 176 V*TGTGGGV*TGGGGLWKISGGSGGNVIGGMVIGG*GGKRG 54
Score = 57.4 bits (137), Expect = 9e-09
Identities = 49/125 (39%), Positives = 56/125 (44%), Gaps = 25/125 (20%)
Frame = -2
Query: 96 NTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLG------GGLGG--GLGG--------- 138
N GG GG G + +G G GS GG G GG GG G+GG
Sbjct: 875 NLGSGGRGGSGF-L*IGTG*SGGNLGSGGRGGSGFL*IGTGGSGGNLGIGGSGFLKIGGH 699
Query: 139 ---GMYGGGMY---GSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG--GGMGV 190
G GGG + G S GG GGG +G GG GG GGLGG GG+G GG
Sbjct: 698 GFFGGVGGGSFGGVGGSFGGFGGRGGGFFGGFGG--SGGTTFGGLGGFFGGFGG*GGSTF 525
Query: 191 GPYGG 195
G +GG
Sbjct: 524 GGFGG 510
Score = 46.2 bits (108), Expect = 2e-05
Identities = 41/120 (34%), Positives = 50/120 (41%), Gaps = 24/120 (20%)
Frame = -2
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGG-----LGGGLGGGMYGGGMYGSSMY-- 151
+GG+ G G GSGL G + G G GG +G G GG G G G S +
Sbjct: 1052 FGGSLGRG---------GSGLSGGNLGSGGRGGSGFL*IGTG*SGGNLGSGGRGGSGFL* 900
Query: 152 -----NRGGYGGGLYGSSG------GMYGGGMYGGGLGGP------MGGYGGGMGVGPYG 194
+ G G G G SG G GG + GG GG GG GG +G+G G
Sbjct: 899 IGTG*SGGNLGSGGRGGSGFL*IGTG*SGGNLGSGGRGGSGFL*IGTGGSGGNLGIGGSG 720
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.139 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,039,019
Number of Sequences: 63676
Number of extensions: 347904
Number of successful extensions: 13912
Number of sequences better than 10.0: 888
Number of HSP's better than 10.0 without gapping: 4321
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7745
length of query: 329
length of database: 12,639,632
effective HSP length: 98
effective length of query: 231
effective length of database: 6,399,384
effective search space: 1478257704
effective search space used: 1478257704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0106.12


