

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.17
(488 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
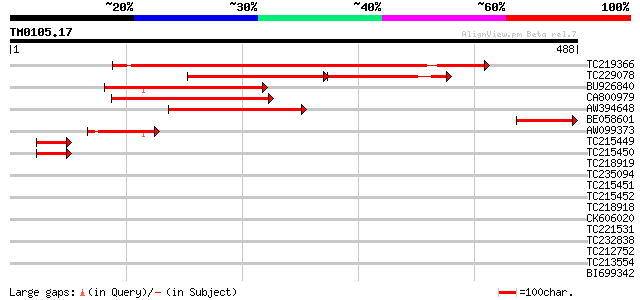
Score E
Sequences producing significant alignments: (bits) Value
TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating pro... 367 e-102
TC229078 similar to UP|O82458 (O82458) Rac GTPase activating pro... 181 5e-76
BU926840 similar to PIR|T01383|T01 GTPase-activating protein hom... 202 4e-52
CA800979 similar to GP|3695059|gb| rac GTPase activating protein... 191 5e-49
AW394648 similar to GP|3695059|gb| rac GTPase activating protein... 186 2e-47
BE058601 similar to GP|3695061|gb|A rac GTPase activating protei... 98 1e-20
AW099373 similar to GP|3695063|gb|A rac GTPase activating protei... 55 7e-08
TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%) 42 8e-04
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 42 8e-04
TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly pro... 41 0.001
TC235094 similar to UP|PRT3_SCYCA (P30258) Protamine Z3 (Scyllio... 40 0.002
TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembl... 40 0.002
TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%) 40 0.002
TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete 39 0.005
CK606020 39 0.007
TC221531 similar to UP|Q84N48 (Q84N48) CRS2-associated factor 2,... 38 0.012
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 37 0.015
TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific... 37 0.015
TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protei... 37 0.020
BI699342 37 0.026
>TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating protein,
partial (52%)
Length = 1397
Score = 367 bits (942), Expect = e-102
Identities = 192/326 (58%), Positives = 237/326 (71%), Gaps = 1/326 (0%)
Frame = +1
Query: 89 ILDILVAALKKSLVTCSVDREDVSSL-DISWPTEVRHVSHVTFDRFNGFLGLPTELQPEV 147
+L +L+ L+KS + R+++ S+ DI PT VRHV+HVTFDRFNGFLGLP E +PEV
Sbjct: 7 LLALLLTLLRKSF---QLPRKNIGSIMDIGSPTNVRHVAHVTFDRFNGFLGLPVEFEPEV 177
Query: 148 PQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNS 207
P++ PSASA VFGVS +SMQ SYD RGNSVPTILL+MQ LY +GGL+ EGIFRINADN
Sbjct: 178 PRRPPSASASVFGVSTESMQLSYDSRGNSVPTILLLMQRHLYVQGGLQVEGIFRINADNG 357
Query: 208 QEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLV 267
QEE R QLN G+VP G++VHCL+GLIKAWFRELPTG+LDSL+PEQVM C +E++C LV
Sbjct: 358 QEEHARDQLNLGVVPEGIDVHCLAGLIKAWFRELPTGILDSLSPEQVMQCQTEDECAELV 537
Query: 268 KLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVM 327
+ LP TEA+LLDWAINLMADVV+HE NKMNA NIAMVFAPNMTQM DP++AL++AVQVM
Sbjct: 538 RHLPHTEASLLDWAINLMADVVQHENVNKMNAHNIAMVFAPNMTQMADPISALMYAVQVM 717
Query: 328 NFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDTCATMPP 387
NFLKTLIL+T+RER +S+ ++ L S +NR + DT A
Sbjct: 718 NFLKTLILRTVRERKDSVVESCPRFYLQPSVD--------NENRRILESFRQDTPAENEE 873
Query: 388 DKSEFSRMEWCVDEKVWSSEEKGTGG 413
+ F + +D S + TGG
Sbjct: 874 AQENFVLEKTALDRSPESLQNNSTGG 951
>TC229078 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (70%)
Length = 1579
Score = 181 bits (458), Expect(2) = 5e-76
Identities = 87/121 (71%), Positives = 103/121 (84%)
Frame = +1
Query: 154 ASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVR 213
+SA VFGVS +SMQ S+D RGNSVPTILL+MQ LY++GGL+AEGIFRI+A+N QEEFVR
Sbjct: 61 SSANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQAEGIFRIDAENGQEEFVR 240
Query: 214 CQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPST 273
LNRG+VP G++VHCL+GLIKAWFRELPTGVLD +PEQVM SEE+C LV+LLP T
Sbjct: 241 E*LNRGVVPDGIDVHCLAGLIKAWFRELPTGVLDPFSPEQVMQSQSEEECAQLVRLLPPT 420
Query: 274 E 274
E
Sbjct: 421 E 423
Score = 122 bits (306), Expect(2) = 5e-76
Identities = 63/107 (58%), Positives = 80/107 (73%)
Frame = +2
Query: 274 EAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTL 333
++ALLDWAINLMADV + E NKMNARNIAMVFAPNMTQM DPLTAL++AVQVMNFLKTL
Sbjct: 422 KSALLDWAINLMADVAQMENLNKMNARNIAMVFAPNMTQMADPLTALMYAVQVMNFLKTL 601
Query: 334 ILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVD 380
++K LRER+ES+ K+ + D + F D+ S+++ +D
Sbjct: 602 VVKALREREESIVKSNPVP----------DLNSFDDDGNHSNSEMLD 712
>BU926840 similar to PIR|T01383|T01 GTPase-activating protein homolog T4I9.2
- Arabidopsis thaliana, partial (32%)
Length = 443
Score = 202 bits (513), Expect = 4e-52
Identities = 99/146 (67%), Positives = 123/146 (83%), Gaps = 5/146 (3%)
Frame = +3
Query: 82 QNQNQFAILDILVAALKKSLVTCSVDR-EDVSS----LDISWPTEVRHVSHVTFDRFNGF 136
+ QNQ +++ +L+AA++KS+V+C VD EDV S ++I WPT V+H++HVTFDRFNGF
Sbjct: 6 EQQNQLSLVALLLAAIRKSMVSCRVDPPEDVISTVHHMEIGWPTNVQHITHVTFDRFNGF 185
Query: 137 LGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKA 196
LGLP E Q E+P +VPSAS VFGVSA+SMQCSYD +GNSVPTILL+MQ+RLYS+GGLKA
Sbjct: 186 LGLPYEFQVEIPARVPSASVSVFGVSAESMQCSYDPKGNSVPTILLLMQDRLYSQGGLKA 365
Query: 197 EGIFRINADNSQEEFVRCQLNRGLVP 222
EGIFRIN +NSQEE VR QLNRG+VP
Sbjct: 366 EGIFRINPENSQEEHVRDQLNRGIVP 443
>CA800979 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (25%)
Length = 421
Score = 191 bits (486), Expect = 5e-49
Identities = 94/140 (67%), Positives = 111/140 (79%)
Frame = +2
Query: 88 AILDILVAALKKSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEV 147
++L ILV L+KSL+ C+ E +++I WPT VRHV+HVTFDRFNGFLGLP E +PEV
Sbjct: 2 SLLAILVTLLRKSLIACNKSEEGHGAMEIGWPTNVRHVAHVTFDRFNGFLGLPREFEPEV 181
Query: 148 PQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNS 207
+ PSASA VFGVS +SMQ SYD RGNSVPTILL+MQ LY+ GGL+ EGIFRINADNS
Sbjct: 182 STRPPSASATVFGVSTESMQLSYDTRGNSVPTILLLMQRHLYALGGLQEEGIFRINADNS 361
Query: 208 QEEFVRCQLNRGLVPRGVEV 227
QEE VR QLNRGLVP V++
Sbjct: 362 QEESVRDQLNRGLVPEDVDI 421
>AW394648 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (24%)
Length = 361
Score = 186 bits (472), Expect = 2e-47
Identities = 89/119 (74%), Positives = 104/119 (86%)
Frame = +2
Query: 137 LGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKA 196
LGLP E +PEVP++ PSASA VFGVS +SMQ S+D RGNSVPTILL+MQ LY++GGL+A
Sbjct: 2 LGLPVEFEPEVPRRPPSASANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQA 181
Query: 197 EGIFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVM 255
EGIFRINA+N QEEFVR QLNRG+VP G++VHCL+GLIKAWFRELP VLD L+PEQVM
Sbjct: 182 EGIFRINAENGQEEFVREQLNRGIVPDGIDVHCLAGLIKAWFRELPNWVLDPLSPEQVM 358
>BE058601 similar to GP|3695061|gb|A rac GTPase activating protein 2 {Lotus
japonicus}, partial (12%)
Length = 307
Score = 97.8 bits (242), Expect = 1e-20
Identities = 44/52 (84%), Positives = 48/52 (91%)
Frame = +2
Query: 437 YRGIYDSEHWLRLRKGVRRLCQHPVFQLSKSTKKRADLGIVNTREGGGEAWA 488
YRGIYDSEHW++LR GVRRLC+HPVFQLSK TKKRA LG+VNTREGGGEA A
Sbjct: 2 YRGIYDSEHWVKLRNGVRRLCRHPVFQLSKPTKKRASLGVVNTREGGGEALA 157
>AW099373 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 416
Score = 55.1 bits (131), Expect = 7e-08
Identities = 32/68 (47%), Positives = 43/68 (63%), Gaps = 6/68 (8%)
Frame = +2
Query: 68 GSKDKGNWRGQSHNQNQNQFAILDILVAALKKSLVTCSVDR-EDVSS-----LDISWPTE 121
G KG RG + QN + + +L+AAL+KS+V CSVD +DV S ++I WPT
Sbjct: 215 GGVGKGR-RGGGGEEEQNPASPVAVLLAALRKSMVACSVDSPDDVISAVHHPMEIGWPTN 391
Query: 122 VRHVSHVT 129
V+HVSHVT
Sbjct: 392 VKHVSHVT 415
>TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%)
Length = 1495
Score = 41.6 bits (96), Expect = 8e-04
Identities = 17/30 (56%), Positives = 25/30 (82%)
Frame = +2
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F +++D+EDDED EE+++EEEE DD+DD
Sbjct: 1031 FGDLEDDEDDEDIEEDDDEEEEEDEDDDDD 1120
Score = 32.3 bits (72), Expect = 0.49
Identities = 12/33 (36%), Positives = 23/33 (69%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTP 62
EEDD+++EEE+E++++ +D+E+ S P
Sbjct: 1070 EEDDDEEEEEDEDDDDDEDDEEESKTKKKSSAP 1168
Score = 30.8 bits (68), Expect = 1.4
Identities = 10/18 (55%), Positives = 18/18 (99%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEE 45
++EE+DEDD+++E++EEE
Sbjct: 1085 EEEEEDEDDDDDEDDEEE 1138
Score = 30.0 bits (66), Expect = 2.4
Identities = 12/25 (48%), Positives = 20/25 (80%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+E+DDE++EE+E+++ DDEDD
Sbjct: 1070 EEDDDEEEEEDEDDD-----DDEDD 1129
Score = 28.1 bits (61), Expect = 9.2
Identities = 9/17 (52%), Positives = 16/17 (93%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEE 44
+++EDD+DDE++EEE +
Sbjct: 1094 EEDEDDDDDEDDEEESK 1144
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 41.6 bits (96), Expect = 8e-04
Identities = 17/30 (56%), Positives = 25/30 (82%)
Frame = +3
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F +++D+EDDED EE+++EEEE DD+DD
Sbjct: 198 FGDLEDDEDDEDIEEDDDEEEEEDEDDDDD 287
Score = 31.6 bits (70), Expect = 0.84
Identities = 13/24 (54%), Positives = 20/24 (83%)
Frame = +3
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
EEDD+++EEE+E+++ DDEDD
Sbjct: 237 EEDDDEEEEEDEDDD----DDEDD 296
Score = 30.8 bits (68), Expect = 1.4
Identities = 10/18 (55%), Positives = 18/18 (99%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEE 45
++EE+DEDD+++E++EEE
Sbjct: 252 EEEEEDEDDDDDEDDEEE 305
Score = 28.1 bits (61), Expect = 9.2
Identities = 9/17 (52%), Positives = 16/17 (93%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEE 44
+++EDD+DDE++EEE +
Sbjct: 261 EEDEDDDDDEDDEEESK 311
>TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly protein 1,
partial (33%)
Length = 729
Score = 41.2 bits (95), Expect = 0.001
Identities = 20/59 (33%), Positives = 36/59 (60%), Gaps = 4/59 (6%)
Frame = +1
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVST----PLINPAGSKDKGNWRGQ 78
F +++D+ED+E+DE+E+E+EE Y++DEDD + T + +G G+ G+
Sbjct: 178 FEDLEDDEDEEEDEDEDEDEE--YDEDEDDEEEDDTKTKKKLSAVKKSGKAQAGDGDGE 348
>TC235094 similar to UP|PRT3_SCYCA (P30258) Protamine Z3 (Scylliorhinine Z3),
partial (76%)
Length = 466
Score = 40.4 bits (93), Expect = 0.002
Identities = 23/43 (53%), Positives = 29/43 (66%)
Frame = -2
Query: 18 SPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVS 60
SPP+P +N EEDDE DE E+E+EEE +DD+DD S S
Sbjct: 378 SPPNP--NNA--EEDDEIDEPEDEDEEEEEDDDDDDVVSQEQS 262
>TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembly protein,
partial (20%)
Length = 777
Score = 40.4 bits (93), Expect = 0.002
Identities = 16/30 (53%), Positives = 26/30 (86%)
Frame = +2
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F +++D+EDDED EE+++E+EE +DD+DD
Sbjct: 77 FGDLEDDEDDEDIEEDDDEDEEDDDDDDDD 166
Score = 31.6 bits (70), Expect = 0.84
Identities = 10/18 (55%), Positives = 18/18 (99%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEE 45
+DEEDD+DD++++++EEE
Sbjct: 131 EDEEDDDDDDDDDDDEEE 184
>TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%)
Length = 609
Score = 40.4 bits (93), Expect = 0.002
Identities = 16/30 (53%), Positives = 26/30 (86%)
Frame = +3
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F +++D+EDDED EE+++E+EE +DD+DD
Sbjct: 126 FGDLEDDEDDEDIEEDDDEDEEDDDDDDDD 215
Score = 31.6 bits (70), Expect = 0.84
Identities = 10/18 (55%), Positives = 18/18 (99%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEE 45
+DEEDD+DD++++++EEE
Sbjct: 180 EDEEDDDDDDDDDDDEEE 233
Score = 31.2 bits (69), Expect = 1.1
Identities = 11/33 (33%), Positives = 23/33 (69%)
Frame = +3
Query: 30 EEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTP 62
EEDD++DEE+++++++ +D+E+ S P
Sbjct: 165 EEDDDEDEEDDDDDDDDDDDEEESKTKKKSSAP 263
>TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete
Length = 1507
Score = 38.9 bits (89), Expect = 0.005
Identities = 15/30 (50%), Positives = 27/30 (90%)
Frame = +1
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
F +++D+ED+E+DE+E+E+EE+ ++DEDD
Sbjct: 991 FEDLEDDEDEEEDEDEDEDEED--DEDEDD 1074
Score = 37.0 bits (84), Expect = 0.020
Identities = 13/26 (50%), Positives = 24/26 (92%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+DEE+DED++E+EE++E+ +++EDD
Sbjct: 1012 EDEEEDEDEDEDEEDDEDEDDEEEDD 1089
Score = 32.7 bits (73), Expect = 0.38
Identities = 13/25 (52%), Positives = 20/25 (80%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE +D +D+E+EEE+E+ D+EDD
Sbjct: 985 DEFEDLEDDEDEEEDEDEDEDEEDD 1059
>CK606020
Length = 340
Score = 38.5 bits (88), Expect = 0.007
Identities = 19/52 (36%), Positives = 34/52 (64%), Gaps = 11/52 (21%)
Frame = +1
Query: 13 VEFNPS--PPSPFFHN---------VKDEEDDEDDEEEEEEEEELYNDDEDD 53
++FNP P SP ++ ++EE++E++EEEEEEEEE+ ++E++
Sbjct: 127 IQFNPMSLPDSPVANDDEIDPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEE 282
Score = 32.7 bits (73), Expect = 0.38
Identities = 13/26 (50%), Positives = 22/26 (84%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E +EEEEEEEEE+ + E++
Sbjct: 238 EEEEEEEIEEEEEEEEEEVEEEVEEE 315
Score = 30.8 bits (68), Expect = 1.4
Identities = 12/26 (46%), Positives = 21/26 (80%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E+ EEEEEEEEE ++ ++
Sbjct: 235 EEEEEEEEIEEEEEEEEEEVEEEVEE 312
Score = 30.0 bits (66), Expect = 2.4
Identities = 12/26 (46%), Positives = 21/26 (80%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EE EEEEEE + E++
Sbjct: 226 EEEEEEEEEEEIEEEEEEEEEEVEEE 303
Score = 29.3 bits (64), Expect = 4.1
Identities = 11/26 (42%), Positives = 21/26 (80%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEE EEEEE ++ ++
Sbjct: 223 EEEEEEEEEEEEIEEEEEEEEEEVEE 300
Score = 29.3 bits (64), Expect = 4.1
Identities = 12/24 (50%), Positives = 19/24 (79%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDE 51
++EE +E++EEEEEE EE ++E
Sbjct: 247 EEEEIEEEEEEEEEEVEEEVEEEE 318
Score = 28.5 bits (62), Expect = 7.1
Identities = 12/25 (48%), Positives = 20/25 (80%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDED 52
++EE+ E++EEEEEEE E ++E+
Sbjct: 244 EEEEEIEEEEEEEEEEVEEEVEEEE 318
Score = 28.5 bits (62), Expect = 7.1
Identities = 11/25 (44%), Positives = 19/25 (76%)
Frame = +1
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDE 51
+++EE++E++E EEE EEE +E
Sbjct: 259 IEEEEEEEEEEVEEEVEEEEVEGEE 333
>TC221531 similar to UP|Q84N48 (Q84N48) CRS2-associated factor 2, partial
(5%)
Length = 922
Score = 37.7 bits (86), Expect = 0.012
Identities = 11/28 (39%), Positives = 26/28 (92%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWS 56
+EE++E+++E+E+E+E+ Y+D++DD ++
Sbjct: 640 EEEEEEEEDEDEDEDEDYYDDEDDDLYA 723
Score = 35.0 bits (79), Expect = 0.076
Identities = 14/26 (53%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E+DE+E+E+E+ Y DDEDD
Sbjct: 640 EEEEEEEEDEDEDEDED--YYDDEDD 711
Score = 30.0 bits (66), Expect = 2.4
Identities = 18/51 (35%), Positives = 25/51 (48%), Gaps = 8/51 (15%)
Frame = +1
Query: 15 FNPSPPSPFFHNVKDEEDDEDD--------EEEEEEEEELYNDDEDDYWSN 57
F S S H +D ED EEEEEEEE+ D+++DY+ +
Sbjct: 550 FENSYSSDGLHTEEDYAKFEDSYSSDALQTEEEEEEEEDEDEDEDEDYYDD 702
Score = 28.1 bits (61), Expect = 9.2
Identities = 12/36 (33%), Positives = 24/36 (66%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPL 63
+DE++DED++ ++E+++LY D + P S P+
Sbjct: 661 EDEDEDEDEDYYDDEDDDLYAVDTS---AQPGSLPV 759
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 37.4 bits (85), Expect = 0.015
Identities = 15/27 (55%), Positives = 22/27 (80%)
Frame = +2
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
V D EDDED+EE+++++E NDDE+D
Sbjct: 320 VDDAEDDEDEEEDDDDDEGDDNDDEED 400
Score = 35.0 bits (79), Expect = 0.076
Identities = 14/58 (24%), Positives = 31/58 (53%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGNWRGQSHNQNQNQ 86
D +DD DD+E+E+EE+E +E+D + + PL + ++ + + + + +
Sbjct: 494 DSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEENGEEEEE 667
Score = 34.3 bits (77), Expect = 0.13
Identities = 17/55 (30%), Positives = 28/55 (50%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGNWRGQSHNQN 83
D +DDED++EE+EEE+ D +Y P+ T A S + G+ ++
Sbjct: 506 DSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEENGEEEEED 670
Score = 30.0 bits (66), Expect = 2.4
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE DD DDEE++ + +DDEDD
Sbjct: 368 DEGDDNDDEEDDAPDGGDDDDDEDD 442
Score = 29.6 bits (65), Expect = 3.2
Identities = 12/25 (48%), Positives = 18/25 (72%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D++DDEDDEEE + + DD+D+
Sbjct: 419 DDDDDEDDEEEGDVQRGGEPDDDDN 493
Score = 29.6 bits (65), Expect = 3.2
Identities = 10/26 (38%), Positives = 19/26 (72%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D++DDE D+ ++EE++ D+DD
Sbjct: 353 EDDDDDEGDDNDDEEDDAPDGGDDDD 430
Score = 28.1 bits (61), Expect = 9.2
Identities = 15/33 (45%), Positives = 20/33 (60%), Gaps = 9/33 (27%)
Frame = +2
Query: 30 EEDDED---------DEEEEEEEEELYNDDEDD 53
EEDDE+ D E++E+EEE +DDE D
Sbjct: 281 EEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGD 379
>TC212752 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein
T2 precursor, partial (35%)
Length = 533
Score = 37.4 bits (85), Expect = 0.015
Identities = 19/61 (31%), Positives = 30/61 (49%)
Frame = +3
Query: 30 EEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGNWRGQSHNQNQNQFAI 89
EEDD++ EEEEEEEEE +E++ W +W G+ + +++F
Sbjct: 60 EEDDDESEEEEEEEEE----EEEEVWD-----------------DWEGEDEGERESEFVC 176
Query: 90 L 90
L
Sbjct: 177L 179
Score = 35.8 bits (81), Expect = 0.044
Identities = 14/25 (56%), Positives = 22/25 (88%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D+E +E++EEEEEEEEE+++D E +
Sbjct: 69 DDESEEEEEEEEEEEEEVWDDWEGE 143
>TC213554 weakly similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1,
partial (18%)
Length = 451
Score = 37.0 bits (84), Expect = 0.020
Identities = 25/55 (45%), Positives = 35/55 (63%), Gaps = 9/55 (16%)
Frame = +2
Query: 27 VKDEEDDEDDEEEEEEEEELYND---------DEDDYWSNPVSTPLINPAGSKDK 72
V++EE++E+DEE+EEEEEE D DEDD +P+S L+ P SKD+
Sbjct: 239 VEEEEEEEEDEEDEEEEEEDQKDQYQHQQQQQDEDD---DPIS-KLLEPL-SKDQ 388
Score = 33.1 bits (74), Expect = 0.29
Identities = 17/45 (37%), Positives = 27/45 (59%), Gaps = 3/45 (6%)
Frame = +2
Query: 12 LVEFNPSPPSP---FFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
L + P P + + V++E ++E+ EEE EEEEE D+ED+
Sbjct: 146 LKQHQPKPENAAQEYEEEVEEEVEEEEVEEEVEEEEEEEEDEEDE 280
Score = 32.7 bits (73), Expect = 0.38
Identities = 12/27 (44%), Positives = 21/27 (77%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDY 54
++ E++E++EE+EE+EEE D +D Y
Sbjct: 233 EEVEEEEEEEEDEEDEEEEEEDQKDQY 313
Score = 32.0 bits (71), Expect = 0.64
Identities = 13/27 (48%), Positives = 23/27 (85%)
Frame = +2
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
V++EE +E+ EEEEEEEE+ +++E++
Sbjct: 212 VEEEEVEEEVEEEEEEEEDEEDEEEEE 292
Score = 30.4 bits (67), Expect = 1.9
Identities = 12/23 (52%), Positives = 19/23 (82%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEELYNDDED 52
EE+ E++EEEEE+EE+ ++ED
Sbjct: 230 EEEVEEEEEEEEDEEDEEEEEED 298
Score = 29.6 bits (65), Expect = 3.2
Identities = 11/26 (42%), Positives = 21/26 (80%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++ E++ ++EEEEEE+EE ++E+D
Sbjct: 221 EEVEEEVEEEEEEEEDEEDEEEEEED 298
Score = 28.9 bits (63), Expect = 5.4
Identities = 11/26 (42%), Positives = 21/26 (80%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++E ++E +EEEEEEE+E ++E++
Sbjct: 218 EEEVEEEVEEEEEEEEDEEDEEEEEE 295
>BI699342
Length = 325
Score = 36.6 bits (83), Expect = 0.026
Identities = 14/27 (51%), Positives = 23/27 (84%)
Frame = -1
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDED 52
N ++EE++E++EEEEEEEEE ++E+
Sbjct: 271 NAEEEEEEEEEEEEEEEEEEEEEEEEE 191
Score = 34.7 bits (78), Expect = 0.099
Identities = 13/25 (52%), Positives = 22/25 (88%)
Frame = -1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+EE++E++EEEEEEEEE ++E++
Sbjct: 265 EEEEEEEEEEEEEEEEEEEEEEEEE 191
Score = 32.0 bits (71), Expect = 0.64
Identities = 12/21 (57%), Positives = 19/21 (90%)
Frame = -1
Query: 28 KDEEDDEDDEEEEEEEEELYN 48
++EE++E++EEEEEEEE+ N
Sbjct: 241 EEEEEEEEEEEEEEEEEKAIN 179
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,215,039
Number of Sequences: 63676
Number of extensions: 320379
Number of successful extensions: 5393
Number of sequences better than 10.0: 225
Number of HSP's better than 10.0 without gapping: 2309
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3734
length of query: 488
length of database: 12,639,632
effective HSP length: 101
effective length of query: 387
effective length of database: 6,208,356
effective search space: 2402633772
effective search space used: 2402633772
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0105.17


