

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
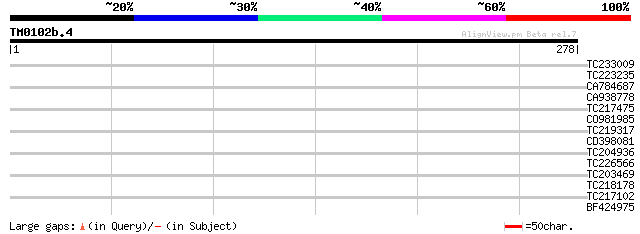
Query= TM0102b.4
(278 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC233009 35 0.049
TC223235 similar to UP|Q9XIN7 (Q9XIN7) NAM (No apical meristem)-... 30 0.92
CA784687 similar to SP|Q9JX33|DXR_N 1-deoxy-D-xylulose 5-phospha... 29 2.7
CA938778 29 2.7
TC217475 similar to UP|Q9FKX7 (Q9FKX7) Arabidopsis thaliana geno... 28 3.5
CO981985 28 4.6
TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {A... 28 4.6
CD398081 28 6.0
TC204936 homologue to UP|FD3C_SOYBN (P48621) Omega-3 fatty acid ... 28 6.0
TC226566 28 6.0
TC203469 weakly similar to UP|A4S1_HUMAN (Q9Y587) Adapter-relate... 28 6.0
TC218178 weakly similar to GB|CAD40663.1|21741487|OSJN00124 OSJN... 27 7.8
TC217102 weakly similar to UP|Q6K1Z7 (Q6K1Z7) Isochorismatase hy... 27 7.8
BF424975 similar to PIR|JQ2128|JQ212 metallothionein - soybean, ... 27 7.8
>TC233009
Length = 664
Score = 34.7 bits (78), Expect = 0.049
Identities = 13/35 (37%), Positives = 23/35 (65%)
Frame = +1
Query: 112 RVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDE 146
R+ W+ C GIPLH ++ K ++ VG+++ ID+
Sbjct: 109 RLVWILCWGIPLHA*DLENIKKIVRGVGEMLDIDD 213
>TC223235 similar to UP|Q9XIN7 (Q9XIN7) NAM (No apical meristem)-like
protein, partial (30%)
Length = 809
Score = 30.4 bits (67), Expect = 0.92
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = +1
Query: 105 SSFAPINRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDT 148
S+ P+N T+ +G P + DC+ +L D+IK+DE +
Sbjct: 463 SNQRPVNEATFTESSGQPKEGYEEDCYAEILKD--DIIKLDESS 588
>CA784687 similar to SP|Q9JX33|DXR_N 1-deoxy-D-xylulose 5-phosphate
reductoisomerase (EC 1.1.1.267) (DXP reductoisomerase),
partial (6%)
Length = 437
Score = 28.9 bits (63), Expect = 2.7
Identities = 23/93 (24%), Positives = 38/93 (40%), Gaps = 6/93 (6%)
Frame = -1
Query: 100 LKPWSSSFAPINRVT------WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVV 153
L WSS+F P N+V W R W + ++ VG I +DE T
Sbjct: 329 LSQWSSNFNP-NKVRQTHAQCWKRVHEFSQEYWDLQLLFEIVGAVGSPITLDEATKSKTF 153
Query: 154 LEFSRLLVRTTTLNFITLARKILIKGTTYTISI 186
FS +L+ TL +++++ +Y +
Sbjct: 152 GHFSCILMDVDLAK--TLHLEVMVERESYAFLV 60
>CA938778
Length = 451
Score = 28.9 bits (63), Expect = 2.7
Identities = 37/130 (28%), Positives = 58/130 (44%), Gaps = 7/130 (5%)
Frame = -3
Query: 67 LRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSS--FAPINRVTWVRCTGIPL- 123
L DD+ LLT+ + S + L++ S R + +N+ P ++S F I+ ++ + I L
Sbjct: 437 LFDDKRLLTSLRTSGINSLVRRSAEPRRQMRENISPTNNSAPFLSISSMSTLSSKIISLS 258
Query: 124 ----HMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKG 179
WT C K+ D +E V+EF L TL ILI
Sbjct: 257 SARESSWTTICLKNTC-----------DDNE--VVEFILALSGIDATTCATLDTSILI-C 120
Query: 180 TTYTISIIEE 189
T+ T+S +EE
Sbjct: 119 TSVTLSNLEE 90
>TC217475 similar to UP|Q9FKX7 (Q9FKX7) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K2A18, partial (12%)
Length = 708
Score = 28.5 bits (62), Expect = 3.5
Identities = 16/53 (30%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Frame = -1
Query: 152 VVLEFSRLLVRTTTLNFITLARKILIKGTTYTI--SIIEEGEALLHPNCCCRE 202
V +++ R +T I++ + I +TY + + EGE L CCCRE
Sbjct: 456 VSIKYYPFWTRESTA*TISIETPVEIYASTYDSGRTFLREGEKLRDNRCCCRE 298
>CO981985
Length = 903
Score = 28.1 bits (61), Expect = 4.6
Identities = 12/33 (36%), Positives = 19/33 (57%), Gaps = 2/33 (6%)
Frame = +1
Query: 118 CT--GIPLHMWTMDCFKHLLLQVGDVIKIDEDT 148
CT G P++MW ++C LL G ++ +D T
Sbjct: 184 CTLVGFPINMWNVNCHGLCLLMFGSMLMMDSYT 282
>TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {Arabidopsis
thaliana;} , partial (33%)
Length = 829
Score = 28.1 bits (61), Expect = 4.6
Identities = 16/59 (27%), Positives = 28/59 (47%)
Frame = +3
Query: 30 EEWMLRSLVREMKNLELIPSLREAFMVEGHITIQVRYLRDDRVLLTNHQESNLVELLQG 88
+ WM S+V +KNL+ P L + F +T + R + +D + ES + +G
Sbjct: 372 DRWMRESVVEIVKNLKEAPLLVQVFPKSATMTTEKRMVVEDWPAVKERWESGETPVPEG 548
>CD398081
Length = 513
Score = 27.7 bits (60), Expect = 6.0
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = -3
Query: 141 VIKIDEDTSEFVVLEFSRLLVRTTTLNF 168
++ +D+D +E +LEF LLV T +F
Sbjct: 454 LVGVDDDENELAILEFIHLLVETMDRHF 371
>TC204936 homologue to UP|FD3C_SOYBN (P48621) Omega-3 fatty acid desaturase,
chloroplast precursor , partial (46%)
Length = 637
Score = 27.7 bits (60), Expect = 6.0
Identities = 20/67 (29%), Positives = 29/67 (42%)
Frame = +2
Query: 78 QESNLVELLQGSKGWRSEFVDNLKPWSSSFAPINRVTWVRCTGIPLHMWTMDCFKHLLLQ 137
+E V+++ GS G E + P + P N +R IP H W D K +
Sbjct: 14 EEQKSVDVINGSNGVEHEKLPEFDPGAPP--PFNLAD-IRAA-IPKHCWVKDPLKSMSYV 181
Query: 138 VGDVIKI 144
V DVI +
Sbjct: 182 VRDVIAV 202
>TC226566
Length = 973
Score = 27.7 bits (60), Expect = 6.0
Identities = 18/59 (30%), Positives = 31/59 (52%)
Frame = -2
Query: 147 DTSEFVVLEFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNCCCREESS 205
D+ FVV SR L+RT + I L K + TY++++ E ++ L+ N +S+
Sbjct: 801 DSFPFVVQFSSRPLLRTYSTQLI*LQMKD*VIARTYSVALSCEVDSFLNSNSLLTSQST 625
>TC203469 weakly similar to UP|A4S1_HUMAN (Q9Y587) Adapter-related protein
complex 4 sigma 1 subunit (Sigma subunit of AP-4) (AP-4
adapter complex sigma subunit), partial (92%)
Length = 677
Score = 27.7 bits (60), Expect = 6.0
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +2
Query: 141 VIKIDEDTSEFVVLEFSRLLVRTTTLNF 168
++ +D+D +E +LEF LLV T +F
Sbjct: 230 LVGVDDDENELAILEFIHLLVETMDRHF 313
>TC218178 weakly similar to GB|CAD40663.1|21741487|OSJN00124 OSJNBb0118P14.1
{Oryza sativa (japonica cultivar-group);} , partial
(25%)
Length = 995
Score = 27.3 bits (59), Expect = 7.8
Identities = 14/31 (45%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Frame = +1
Query: 106 SFAPINRVT-WVRCTG--IPLHMWTMDCFKH 133
+F PI R T TG P H W +DCF H
Sbjct: 226 TFLPIYRTTKGAYTTGCVFPFHFWLLDCFWH 318
>TC217102 weakly similar to UP|Q6K1Z7 (Q6K1Z7) Isochorismatase hydrolase-like
protein, partial (82%)
Length = 1178
Score = 27.3 bits (59), Expect = 7.8
Identities = 13/20 (65%), Positives = 15/20 (75%), Gaps = 1/20 (5%)
Frame = +3
Query: 19 LELWSGPEIPKEEW-MLRSL 37
+ LW PEIPKE W +LRSL
Sbjct: 642 MSLWR*PEIPKELWHILRSL 701
>BF424975 similar to PIR|JQ2128|JQ212 metallothionein - soybean, partial
(92%)
Length = 238
Score = 27.3 bits (59), Expect = 7.8
Identities = 12/36 (33%), Positives = 21/36 (58%)
Frame = -2
Query: 90 KGWRSEFVDNLKPWSSSFAPINRVTWVRCTGIPLHM 125
+G + +F +L+PW S+ PI+ + TG P H+
Sbjct: 228 QGLQQQFGPHLQPWLSAGTPISAPSN*ALTGCPSHL 121
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,070,100
Number of Sequences: 63676
Number of extensions: 208513
Number of successful extensions: 1172
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1169
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1172
length of query: 278
length of database: 12,639,632
effective HSP length: 96
effective length of query: 182
effective length of database: 6,526,736
effective search space: 1187865952
effective search space used: 1187865952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0102b.4


