

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0101.12
(310 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
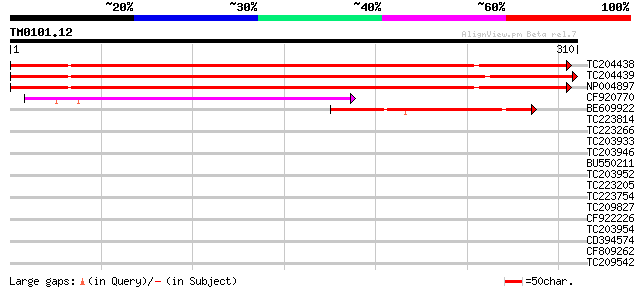
Sequences producing significant alignments: (bits) Value
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 359 e-100
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 353 7e-98
NP004897 gag-protease polyprotein 352 9e-98
CF920770 107 7e-24
BE609922 similar to GP|29423273|g gag-pol polyprotein {Glycine m... 91 7e-19
TC223814 35 0.033
TC223266 32 0.48
TC203933 elongation factor 1A SMV resistance-related protein [Gl... 30 1.4
TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%) 29 3.1
BU550211 29 3.1
TC203952 homologue to UP|O81921 (O81921) Elongation factor 1-alp... 28 4.1
TC223205 28 4.1
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 28 4.1
TC209827 28 4.1
CF922226 28 5.3
TC203954 homologue to UP|EF1A_LYCES (P17786) Elongation factor 1... 27 9.0
CD394574 weakly similar to PIR|T10862|T108 phaseolin G-box bindi... 27 9.0
CF809262 27 9.0
TC209542 27 9.0
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 359 bits (922), Expect = e-100
Identities = 179/309 (57%), Positives = 242/309 (77%), Gaps = 2/309 (0%)
Frame = +1
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 82 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TNELKPEEDWTKEEDELALGNSKALNA 258
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 259 LFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 438
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 439 DFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 618
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 619 SLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 798
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
RR +P+V++I D K S+ + K ++K +KG+QCH CEGYGHI++EC T+LKKQ+K
Sbjct: 799 RRQKPHVRNIPFDIRK--GSEYQKKSDEKPSHSKGIQCHGCEGYGHIKAECPTHLKKQRK 972
Query: 299 GMVVTWSDE 307
G+ V SD+
Sbjct: 973 GLSVCRSDD 999
Score = 32.0 bits (71), Expect = 0.37
Identities = 34/134 (25%), Positives = 55/134 (40%), Gaps = 4/134 (2%)
Frame = +1
Query: 160 KVIA-IEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEG-DDADSESE 217
KVIA +E ++ ++ EL G + KL E TKSI ++ ++ D+ +
Sbjct: 1180 KVIANLEAEKEAHEEEISELKGEVGFLNSKL----ENMTKSIKMLNKGSDMLDEVLQLGK 1347
Query: 218 NLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKE--EDKSGQNKGVQ 275
N+ L FN + R T K+ + + R + K + K +
Sbjct: 1348 NVGNQRGL---GFNH--KSAGRTTMTEFVPAKNSTGATMSQHRSRHHGTQQKKSKRKKWR 1512
Query: 276 CHECEGYGHIRSEC 289
CH C YGHI+ C
Sbjct: 1513 CHYCGKYGHIKPFC 1554
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 353 bits (905), Expect = 7e-98
Identities = 177/312 (56%), Positives = 239/312 (75%), Gaps = 2/312 (0%)
Frame = +1
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 82 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TDELKPEEDWTKEEDELALGNSKALNA 258
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILK THEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 259 LFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKMSRLQLLATKFENLKMKEEECIH 438
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE ++DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 439 DFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 618
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 619 SLQTFELGLSDRAEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 798
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+R +P+VQ+I D K Q+ + + K +KG+QCH CEGYGHI +EC T+LKK +K
Sbjct: 799 KRQKPHVQNIPFDIRKGSKYQK--RSDVKPSHSKGIQCHGCEGYGHIIAECPTHLKKHRK 972
Query: 299 GMVVTWSDEDSE 310
G+ V SD +SE
Sbjct: 973 GLSVCQSDTESE 1008
Score = 31.2 bits (69), Expect = 0.63
Identities = 37/138 (26%), Positives = 54/138 (38%), Gaps = 8/138 (5%)
Frame = +1
Query: 160 KVIA-IEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESEN 218
KVIA +E ++ ++ EL G + KL E TKSI ++ S+
Sbjct: 1177 KVIADLEAEKEAHKEEISELKGEVGFLNSKL----ENMTKSIKMLNKG---------SDT 1317
Query: 219 LSEALALLARKFN-RAL----RKIDRRTRPNVQDIKSDNSKSYNSQRKVKE--EDKSGQN 271
L E L L N R L + R T K+ + + R + K +
Sbjct: 1318 LDEVLLLGKNAGNQRGLGFNPKSAGRTTMTEFVPAKNRTGATMSQHRSRHHGMQQKKSKR 1497
Query: 272 KGVQCHECEGYGHIRSEC 289
K +CH C YGHI+ C
Sbjct: 1498 KKWRCHYCGKYGHIKPFC 1551
>NP004897 gag-protease polyprotein
Length = 1923
Score = 352 bits (904), Expect = 9e-98
Identities = 177/309 (57%), Positives = 239/309 (77%), Gaps = 2/309 (0%)
Frame = +1
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+K W+HP EG T KPE++ T E+E +L NSKA+NA
Sbjct: 82 MVAFLKSLDSRTWKAVIKDWEHPKMLDTEGKP-TDGLKPEEDWTKEEDELALGNSKALNA 258
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 259 LFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 438
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +L+VDELIG
Sbjct: 439 DFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNLRVDELIG 618
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 619 SLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 798
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
RR +P+V++I D K S+ + + ++K +KG QCH CEGYGHI++EC T+LKKQ+K
Sbjct: 799 RRQKPHVRNIPFDIRK--GSEYQKRSDEKPSHSKGFQCHGCEGYGHIKAECPTHLKKQRK 972
Query: 299 GMVVTWSDE 307
G+ V SD+
Sbjct: 973 GLSVCRSDD 999
Score = 32.0 bits (71), Expect = 0.37
Identities = 34/132 (25%), Positives = 54/132 (40%), Gaps = 2/132 (1%)
Frame = +1
Query: 160 KVIA-IEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEG-DDADSESE 217
KVIA +E ++ ++ EL G + KL E TKSI ++ ++ D+ +
Sbjct: 1180 KVIANLEAEKEAHEEEISELKGEVGFLNSKL----ENMTKSIKMLNKGSDMLDEVLQLGK 1347
Query: 218 NLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCH 277
N+ L FN T I + + S + R + K + K +CH
Sbjct: 1348 NVGNQRGL---GFNHKSAGRITMTEFVPAKISTGATMSQHRSRHHGTQQKKSKRKKWRCH 1518
Query: 278 ECEGYGHIRSEC 289
C YGHI+ C
Sbjct: 1519 YCGKYGHIKPFC 1554
>CF920770
Length = 581
Score = 107 bits (267), Expect = 7e-24
Identities = 60/187 (32%), Positives = 102/187 (54%), Gaps = 6/187 (3%)
Frame = -2
Query: 9 DIKTWKAVVKGWKHPV---KAQAEGSTSTPE---EKPEDELTSAEEEASLENSKAMNAIF 62
D+ W+A+ G P + +GS+S+ EKP D + + + N KA N I
Sbjct: 574 DLNIWEAIEIGPYIPTTVERVSIDGSSSSESITIEKPRDRWSEEDRKRVQYNLKAKNIIT 395
Query: 63 NAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEY 122
+A+ + + ++ C AK+ W+ L+ THEGTT V+ SR+ LT +E M +E I
Sbjct: 394 SALGMDEYFRVSNCKSAKEMWDTLRLTHEGTTDVKRSRINALTHEYELFRMNTNENIQSM 215
Query: 123 HMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSL 182
R + N ALG+ +E L+ K+LR ++ ++ KV AI E++D+S++ + L G L
Sbjct: 214 QKRFTHIVNHLAALGKEFQNEDLINKVLRCLSREWQPKVTAISESRDLSNMSLATLFGKL 35
Query: 183 QTYKLKL 189
Q ++++L
Sbjct: 34 QEHEMEL 14
>BE609922 similar to GP|29423273|g gag-pol polyprotein {Glycine max}, partial
(7%)
Length = 340
Score = 90.9 bits (224), Expect = 7e-19
Identities = 49/116 (42%), Positives = 76/116 (65%), Gaps = 3/116 (2%)
Frame = +2
Query: 176 DELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSE---SENLSEALALLARKFNR 232
DEL+GSLQT++L L ++ EKK+K++AF+SN EG++ + + E L+ A LL ++F++
Sbjct: 2 DELMGSLQTFELGLSDRAEKKSKNLAFMSN-DEGEEGEYDLDTDEGLTNAGVLLGKQFSK 178
Query: 233 ALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSE 288
+ +IDRR +P+VQ + D KS QRK G +G+QCH CEG H+ +E
Sbjct: 179 VMNRIDRRQKPHVQSRRGDVRKSSKYQRKWGVRPMHG--RGMQCHGCEGVWHVIAE 340
>TC223814
Length = 607
Score = 35.4 bits (80), Expect = 0.033
Identities = 22/57 (38%), Positives = 39/57 (67%), Gaps = 1/57 (1%)
Frame = +1
Query: 146 VRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGE-KPEKKTKSIA 201
+ KIL+S+ + VIA+ +++D+ SL V+E G+LQ ++L+L E + ++K K IA
Sbjct: 292 ITKILQSLLIQ*RP*VIALCDSKDLKSLPVEEFDGTLQVHELELMEDEGQRKGKFIA 462
>TC223266
Length = 454
Score = 31.6 bits (70), Expect = 0.48
Identities = 25/69 (36%), Positives = 36/69 (51%)
Frame = +1
Query: 115 EDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLK 174
E+ER E RVR+L +LGE +S L K+L + + A++ A+D S
Sbjct: 130 EEERCKESEARVRELEKQVASLGEGVS---LEAKLLSRKEAALRQREAALKNAKD-SKDG 297
Query: 175 VDELIGSLQ 183
VD+ I SLQ
Sbjct: 298 VDKEITSLQ 324
>TC203933 elongation factor 1A SMV resistance-related protein [Glycine max]
Length = 1323
Score = 30.0 bits (66), Expect = 1.4
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Frame = +2
Query: 160 KVIAIEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVS-NTTEGDDADSESEN 218
K + +E + S + DE++ + +Y K+G P+K I FV + EGD+ S N
Sbjct: 47 KPMVVETFSEYSKARYDEIVKEVSSYLKKVGYNPDK----IPFVPISGFEGDNMIERSTN 214
Query: 219 L 219
L
Sbjct: 215 L 217
>TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%)
Length = 1858
Score = 28.9 bits (63), Expect = 3.1
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 5/83 (6%)
Frame = +3
Query: 171 SSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVS-NTTEGDDADSESENL----SEALAL 225
S + DE++ + +Y K+G P+K I FV + EGD+ S NL L
Sbjct: 528 SKARYDEIVKEVSSYMKKVGYNPDK----IPFVPISGFEGDNMIERSTNLDWYKGPTLLE 695
Query: 226 LARKFNRALRKIDRRTRPNVQDI 248
+ N R D+ R +QD+
Sbjct: 696 ALDQINEPKRPSDKPLRLPLQDV 764
>BU550211
Length = 625
Score = 28.9 bits (63), Expect = 3.1
Identities = 23/70 (32%), Positives = 35/70 (49%), Gaps = 4/70 (5%)
Frame = -2
Query: 1 MIAFLKSIDIKTWKAVVK---GWKHPVKAQAEGST-STPEEKPEDELTSAEEEASLENSK 56
M F K DI + ++ K GW ++ T STP+E+P+D ++S + E SK
Sbjct: 384 MCCFTKQFDI*S*VSISKCYNGWXXRCRSNNFKIT*STPQEQPKD*MSSISIYLTQEISK 205
Query: 57 AMNAIFNAVD 66
N I A+D
Sbjct: 204 KHNIITLALD 175
>TC203952 homologue to UP|O81921 (O81921) Elongation factor 1-alpha (EF1-a)
(Fragment), partial (85%)
Length = 995
Score = 28.5 bits (62), Expect = 4.1
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 5/83 (6%)
Frame = +2
Query: 171 SSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVS-NTTEGDDADSESENL----SEALAL 225
S + DE++ + +Y K+G P+K I FV + EGD+ S NL L
Sbjct: 107 SKARYDEIVKEVSSYLKKVGYNPDK----IPFVPISGFEGDNMIERSTNLDWYKGPTLLE 274
Query: 226 LARKFNRALRKIDRRTRPNVQDI 248
+ N R D+ R +QD+
Sbjct: 275 ALDQINEPKRPSDKPLRLPLQDV 343
>TC223205
Length = 448
Score = 28.5 bits (62), Expect = 4.1
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Frame = +1
Query: 109 ENLTMTEDERISEYHMRVRDLSNASFAL--GEPMSDEKLVRKILRSVTSKFAMKVIAIEE 166
E + M + S M + A FAL P+S+++++ ++ VT +FAM+ I EE
Sbjct: 25 ETMKMDLHNKKSMPSMSPKKKLTAKFALMKSRPLSEQEILTEVTALVTKQFAMQEILSEE 204
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 28.5 bits (62), Expect = 4.1
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = +2
Query: 269 GQNKGVQCHECEGYGHIRSECAT 291
G G C C G+GH+ +CAT
Sbjct: 209 GGGGGGSCFRCGGFGHMARDCAT 277
>TC209827
Length = 761
Score = 28.5 bits (62), Expect = 4.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Frame = -3
Query: 228 RKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQ----RKVKEEDKSGQNKGV--QCHECEG 281
RK+NR+L + RRT +D++ ++ + S+ EED+ + G+ H EG
Sbjct: 369 RKYNRSLHFLGRRTTA-FEDLEVHEAQRHESEGETGHDASEEDEETGDAGIGEPPHRSEG 193
Query: 282 YGHIRSECATYLKKQ 296
Y + + + +Q
Sbjct: 192 YAFVAKDRERVVPRQ 148
>CF922226
Length = 667
Score = 28.1 bits (61), Expect = 5.3
Identities = 13/51 (25%), Positives = 24/51 (46%)
Frame = -3
Query: 249 KSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKKG 299
K D+ Q+ +++ G ++C+ C+ GH R C ++QK G
Sbjct: 323 KKDSEFDKKKQKPENQKNGEGNIFKIRCYHCKKEGHTRKVCP---ERQKNG 180
>TC203954 homologue to UP|EF1A_LYCES (P17786) Elongation factor 1-alpha
(EF-1-alpha), partial (61%)
Length = 1423
Score = 27.3 bits (59), Expect = 9.0
Identities = 26/83 (31%), Positives = 40/83 (47%), Gaps = 3/83 (3%)
Frame = +3
Query: 171 SSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVS-NTTEGDDADSESENLS--EALALLA 227
S + DE++ + +Y K+G P+K I FV + EGD+ S NL + LL
Sbjct: 918 SKARYDEIVKEVSSYLKKVGYNPDK----IPFVPISGFEGDNMIERSTNLDWYKGPTLL- 1082
Query: 228 RKFNRALRKIDRRTRPNVQDIKS 250
AL +I+ RP+ Q K+
Sbjct: 1083----EALDQINEPKRPSRQASKA 1139
>CD394574 weakly similar to PIR|T10862|T108 phaseolin G-box binding protein
PG2 - kidney bean (fragment), partial (8%)
Length = 702
Score = 27.3 bits (59), Expect = 9.0
Identities = 9/27 (33%), Positives = 16/27 (58%)
Frame = +1
Query: 16 VVKGWKHPVKAQAEGSTSTPEEKPEDE 42
++ GW P + QA T TP++ +D+
Sbjct: 88 IIAGWSQPHRHQAPRKTKTPQKNKKDK 168
>CF809262
Length = 656
Score = 27.3 bits (59), Expect = 9.0
Identities = 32/138 (23%), Positives = 61/138 (44%), Gaps = 4/138 (2%)
Frame = +1
Query: 173 LKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNR 232
+K+ E ++ +L L EKPEK + F+ AD + L A + ++
Sbjct: 250 MKIVEAGAVVELIELAL-EKPEKNMTELIFILLAHLCSCADGREQFLQHAAGIAV--VSK 420
Query: 233 ALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATY 292
+ ++ T I S SK S V+E + G + C ++++CA+Y
Sbjct: 421 RILRVSPTTDDRALHIFSLVSKFSASNEVVQEMLRVGAVSKL-CMV------LQADCASY 579
Query: 293 LKKQKKGMV----VTWSD 306
LK++ +G++ TW++
Sbjct: 580 LKEKARGVLRLHSKTWNN 633
>TC209542
Length = 1445
Score = 27.3 bits (59), Expect = 9.0
Identities = 19/78 (24%), Positives = 38/78 (48%), Gaps = 1/78 (1%)
Frame = +2
Query: 180 GSLQTYKLKLG-EKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKID 238
G+++T+ +++G E + + + VS + +G+ +LA + LR +D
Sbjct: 488 GTVRTFDIRIGRETSDNLGQPVNCVSMSNDGN-------------CILAGCLDSTLRLLD 628
Query: 239 RRTRPNVQDIKSDNSKSY 256
R T +Q+ K +KSY
Sbjct: 629 RSTGELLQEYKGHTNKSY 682
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.126 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,923,108
Number of Sequences: 63676
Number of extensions: 92949
Number of successful extensions: 537
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 513
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 525
length of query: 310
length of database: 12,639,632
effective HSP length: 97
effective length of query: 213
effective length of database: 6,463,060
effective search space: 1376631780
effective search space used: 1376631780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0101.12


