

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
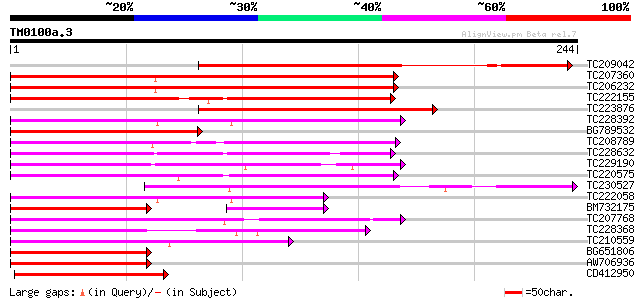
Score E
Sequences producing significant alignments: (bits) Value
TC209042 weakly similar to UP|JOIN_LYCES (Q9FUY6) MADS-box JOINT... 145 2e-35
TC207360 similar to UP|Q8VWZ3 (Q8VWZ3) C-type MADS box protein, ... 115 1e-26
TC206232 homologue to UP|Q8H6F8 (Q8H6F8) MADS box protein GHMADS... 112 1e-25
TC222155 weakly similar to UP|O81662 (O81662) Transcription acti... 110 4e-25
TC223876 weakly similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral re... 104 4e-23
TC228392 similar to UP|Q8LLR0 (Q8LLR0) MADS-box protein 4, parti... 102 1e-22
BG789532 similar to GP|13384052|gb| MADS-box transcription facto... 100 5e-22
TC208789 similar to UP|Q9SBQ1 (Q9SBQ1) MADS box transcription fa... 100 1e-21
TC228632 similar to UP|Q9ZTQ9 (Q9ZTQ9) MADS box protein 26, part... 99 2e-21
TC229190 similar to UP|Q9SEG4 (Q9SEG4) CAGL2, partial (86%) 99 2e-21
TC220575 similar to UP|Q8LLR1 (Q8LLR1) MADS-box protein 3, parti... 91 4e-19
TC230527 similar to UP|Q6V0J7 (Q6V0J7) Short vegetative phase pr... 91 6e-19
TC222058 homologue to UP|AGL9_PETHY (Q03489) Agamous-like MADS b... 88 3e-18
BM732175 similar to GP|15022157|gb MADS box protein-like protein... 74 3e-18
TC207768 similar to UP|Q9ST53 (Q9ST53) MADS-box protein 4, parti... 88 4e-18
TC228368 similar to UP|O49173 (O49173) MADS-box protein NMH 7, p... 86 2e-17
TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragmen... 86 2e-17
BG651806 84 6e-17
AW706936 84 6e-17
CD412950 weakly similar to SP|Q9S7Q7|FLC_A FLOWERING LOCUS C pro... 83 1e-16
>TC209042 weakly similar to UP|JOIN_LYCES (Q9FUY6) MADS-box JOINTLESS protein
(LeMADS), partial (29%)
Length = 700
Score = 145 bits (365), Expect = 2e-35
Identities = 89/161 (55%), Positives = 103/161 (63%)
Frame = +2
Query: 82 PSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVK 141
PS +Q E+DSN+ LRKKVE+K ELRQ+NGEDLQGLTL +L KLEE LKR L +VS+VK
Sbjct: 11 PSIELQIENDSNEILRKKVEDKNRELRQMNGEDLQGLTLQELHKLEEHLKRGLINVSKVK 190
Query: 142 DEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNEYLCSIILSYGGIDSQHILMKY 201
DEK MQEISTLKRK VEL+EENQ+LKQV
Sbjct: 191 DEKLMQEISTLKRKGVELMEENQRLKQV-------------------------------- 274
Query: 202 M*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLKLGL 242
VPSL + RQS ES +S+SS L E+ GSDTSLKLGL
Sbjct: 275 ----VPSLI-HVHRQSSESILSNSSNLPEDGGSDTSLKLGL 382
>TC207360 similar to UP|Q8VWZ3 (Q8VWZ3) C-type MADS box protein, partial
(88%)
Length = 1075
Score = 115 bits (289), Expect = 1e-26
Identities = 64/168 (38%), Positives = 102/168 (60%), Gaps = 1/168 (0%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL+VFS +L+EYA++
Sbjct: 95 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVVFSTRGRLYEYANN 274
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S R + AN+A + + QF + LR+++ + + R + GE L L+
Sbjct: 275 SVRATIERYKKANAAASNAESVSEANTQFYQQESSKLRRQIRDIQNLNRHILGEALGSLS 454
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L +L+ LE L++ L+ V K E ++ ++++E+EL N L+
Sbjct: 455 LKELKNLEGRLEKGLSRVRSRKHETLFADVEFMQKREIELQNHNNYLR 598
>TC206232 homologue to UP|Q8H6F8 (Q8H6F8) MADS box protein GHMADS-2, partial
(95%)
Length = 960
Score = 112 bits (281), Expect = 1e-25
Identities = 64/168 (38%), Positives = 102/168 (60%), Gaps = 1/168 (0%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EY+++
Sbjct: 75 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNN 254
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+ R + A S +T Q+ + LR++++ + R L G+ L LT
Sbjct: 255 NIRSTIERYKKACSDHSSASTTTEINAQYYQQESAKLRQQIQMLQNSNRHLMGDALSTLT 434
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+ +L++LE L+R + + K E + EI +++E+EL EN L+
Sbjct: 435 VKELKQLENRLERGITRIRSKKHEMLLAEIEYFQKREIELENENLCLR 578
>TC222155 weakly similar to UP|O81662 (O81662) Transcription activator,
partial (65%)
Length = 915
Score = 110 bits (276), Expect = 4e-25
Identities = 67/171 (39%), Positives = 103/171 (60%), Gaps = 5/171 (2%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R ++Q+KKI++ +SRQV FSKRR GL KKA ELS LCDA++A+IVFS +L+E++SS
Sbjct: 152 MARGKVQLKKIEDTTSRQVAFSKRRSGLLKKAYELSVLCDAEVAVIVFSQNGRLYEFSSS 331
Query: 61 SRGGLLPLPDANSAT*DKTTPPSP-----TMQFESDSNDTLRKKVEEKTHELRQLNGEDL 115
+L ++ K P S Q + DS +L KK+E H R+L G+ +
Sbjct: 332 DMTKILERYREHT----KDVPASKFGDDYIQQLKLDS-ASLAKKIELLEHSKRELLGQSV 496
Query: 116 QGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
+ +L+ +EE L+ SL V + K + + ++I L+ +E L++EN KL
Sbjct: 497 SSCSYDELKGIEEQLQISLQRVRQRKTQLYTEQIDQLRSQESNLLKENAKL 649
>TC223876 weakly similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral repressor,
partial (12%)
Length = 713
Score = 104 bits (259), Expect = 4e-23
Identities = 52/103 (50%), Positives = 74/103 (71%)
Frame = +2
Query: 82 PSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVK 141
PS Q SDS LRK++E KT+E+ QLNGE++QGLT+ +LQK+EE+L+R ++S++K
Sbjct: 47 PSIGQQLGSDSLGILRKEIEHKTNEMSQLNGEEIQGLTIKELQKVEELLQRRWTTISKIK 226
Query: 142 DEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNEYLCS 184
DEK +QEI+ LK KE +L+EENQKLKQ + + + CS
Sbjct: 227 DEKIIQEINHLKTKEAKLMEENQKLKQSFVRDPRQPYESFTCS 355
>TC228392 similar to UP|Q8LLR0 (Q8LLR0) MADS-box protein 4, partial (98%)
Length = 755
Score = 102 bits (255), Expect = 1e-22
Identities = 65/175 (37%), Positives = 94/175 (53%), Gaps = 5/175 (2%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALI+FS K +E+ S
Sbjct: 40 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKQYEFCSG 219
Query: 61 SR--GGLLPLPDANSAT*DKTTPPSPTMQFESDSND---TLRKKVEEKTHELRQLNGEDL 115
S L N + + E S L+ + E R L GEDL
Sbjct: 220 SSMLKTLERYQKCNYGAPEDNVATKEALVLELSSQQEYLRLKARYEALQRSQRNLMGEDL 399
Query: 116 QGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
L+ +L+ LE L SL + ++ + + ++S L+RKE L E N+ L+Q L
Sbjct: 400 GPLSSKELESLERQLDSSLKQIRSIRTQFMLDQLSDLQRKEHFLGESNRDLRQRL 564
>BG789532 similar to GP|13384052|gb| MADS-box transcription factor FBP13
{Petunia x hybrida}, partial (37%)
Length = 414
Score = 100 bits (250), Expect = 5e-22
Identities = 51/83 (61%), Positives = 65/83 (77%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
MTR RI+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDA++ LIVFS+T KLF+Y+SS
Sbjct: 156 MTRTRIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDAEVGLIVFSSTGKLFDYSSS 335
Query: 61 SRGGLLPLPDANSAT*DKTTPPS 83
S ++ +S +K PS
Sbjct: 336 SMNDIVTKYSTHSHGINKLDKPS 404
>TC208789 similar to UP|Q9SBQ1 (Q9SBQ1) MADS box transcription factor,
partial (72%)
Length = 995
Score = 99.8 bits (247), Expect = 1e-21
Identities = 65/175 (37%), Positives = 93/175 (53%), Gaps = 7/175 (4%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N QVTFSKRR GL KKA+E+S LCDAD+ALIVFS KL +Y++
Sbjct: 219 MGRGRVELKRIENKIFMQVTFSKRRSGLLKKAREISVLCDADVALIVFSTKGKLLDYSNQ 398
Query: 61 -------SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGE 113
R + D+ PP+ E ++ L+ +VE R GE
Sbjct: 399 PCTERILERYERYSYAERQLVGDDQ--PPNENWVIE---HEKLKARVEVLQRNQRNFMGE 563
Query: 114 DLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
DL L L LQ LE+ L +L + K++ + IS L++K+ L E N L +
Sbjct: 564 DLDSLNLRGLQSLEQQLDSALKHIRSRKNQAMNESISELQKKDRTLREHNNLLSK 728
>TC228632 similar to UP|Q9ZTQ9 (Q9ZTQ9) MADS box protein 26, partial (82%)
Length = 880
Score = 99.0 bits (245), Expect = 2e-21
Identities = 61/166 (36%), Positives = 97/166 (57%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N S+RQVT+SKR+ G+ KKA+E++ LCDA ++LI+F+A+ K+ +Y S
Sbjct: 55 MGRGKIEIKRIENSSNRQVTYSKRKNGILKKAKEITVLCDAQVSLIIFAASGKMHDYISP 234
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S L+ + + T K + + + L+K+ + ELR L G+D+ L
Sbjct: 235 ST-TLIDILERYHKTSGKRLWDAKHENLNGEI-ERLKKENDSMQIELRHLKGDDINSLNY 408
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
+L LE+ L+ L SV EK M L+R + L EEN++L
Sbjct: 409 KELMALEDALETGLVSVR----EKQMDVYRMLRRNDKILEEENREL 534
>TC229190 similar to UP|Q9SEG4 (Q9SEG4) CAGL2, partial (86%)
Length = 1062
Score = 99.0 bits (245), Expect = 2e-21
Identities = 67/173 (38%), Positives = 94/173 (53%), Gaps = 3/173 (1%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALI+FS KL+E+ S+
Sbjct: 313 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFCST 492
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKV--EEKTHELRQLNGEDLQGL 118
+ L L + P + ES + L+ K E R L GEDL L
Sbjct: 493 N-SMLKTLERYQKCSYGAVEVSKPGKELESSYREYLKLKARFESLQRTQRNLLGEDLGPL 669
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFM-QEISTLKRKEVELIEENQKLKQVL 170
L++LE L SL K +FM +++ L+ KE L+E N+ L L
Sbjct: 670 NTKDLEQLERQLDSSL------KQTQFMLDQLADLQNKEHMLVEANRSLTMKL 810
>TC220575 similar to UP|Q8LLR1 (Q8LLR1) MADS-box protein 3, partial (44%)
Length = 851
Score = 91.3 bits (225), Expect = 4e-19
Identities = 62/171 (36%), Positives = 92/171 (53%), Gaps = 4/171 (2%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R ++ +++I N +RQVTFSKRR GL KKA ELS LCDA+IAL++FS+ KLF+Y+S+
Sbjct: 158 MGRGKVVLERIQNKINRQVTFSKRRNGLLKKAFELSVLCDAEIALVIFSSRGKLFQYSST 337
Query: 61 SRGGLLPLPDA----NSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQ 116
++ S T D S ++ LR K E R L GE+L+
Sbjct: 338 DINRIIEKYRQCCFNMSQTGDVAEHQSEQCLYQELL--ILRVKHESLQRTQRNLLGEELE 511
Query: 117 GLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L++ +L LE+ L R+L + +K + I L K L + N+ L+
Sbjct: 512 PLSMKELHSLEKQLDRTLGQARKHLTQKLISRIDELHGKVHNLEQVNKHLE 664
>TC230527 similar to UP|Q6V0J7 (Q6V0J7) Short vegetative phase protein,
partial (60%)
Length = 588
Score = 90.5 bits (223), Expect = 6e-19
Identities = 67/189 (35%), Positives = 100/189 (52%), Gaps = 3/189 (1%)
Frame = +3
Query: 59 SSSRGGLLPLPDANSAT*DKTTPPSPTMQFESDSN-DTLRKKVEEKTHELRQLNGEDLQG 117
SSS +L +S + PS +Q +SN L K+V EK+H+LRQL GEDLQG
Sbjct: 3 SSSMKEILERHHLHSKNLARMEQPSLELQLVENSNCSRLSKEVAEKSHQLRQLRGEDLQG 182
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLL 177
L + +LQ+LE L+ L V K EK M EI+ L+RK + L+EEN++LK+
Sbjct: 183 LNIEELQQLEMSLETGLGRVIEKKGEKIMSEIADLQRKGMLLMEENERLKR--------- 335
Query: 178 FNEYLCSII--LSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSD 235
++ II +GG +S++ +M + +S+ +S+ + + SD
Sbjct: 336 ---HVAGIINGQRHGGAESENFVM----------DEGQSSESVTYVCNSTGPPQDFESSD 476
Query: 236 TSLKLGLVY 244
TSLKLGL Y
Sbjct: 477 TSLKLGLPY 503
>TC222058 homologue to UP|AGL9_PETHY (Q03489) Agamous-like MADS box protein
AGL9 homolog (Floral homeotic protein FBP2) (Floral
binding protein 2), partial (60%)
Length = 495
Score = 88.2 bits (217), Expect = 3e-18
Identities = 55/140 (39%), Positives = 76/140 (54%), Gaps = 3/140 (2%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALI+FS KL+E+ SS
Sbjct: 60 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLYEFCSS 239
Query: 61 SR--GGLLPLPDANSAT*DKTTPPSPTMQFESDSND-TLRKKVEEKTHELRQLNGEDLQG 117
S L N + ++ S L+ + E R L GEDL
Sbjct: 240 SSMLKTLERYQKCNYGAPEANVSTREALELSSQQEYLKLKARYEALQRSQRNLMGEDLGP 419
Query: 118 LTLHQLQKLEEVLKRSLASV 137
L+ +L+ LE L SL +
Sbjct: 420 LSSKELESLERQLDSSLKQI 479
>BM732175 similar to GP|15022157|gb MADS box protein-like protein NGL9
{Medicago sativa}, partial (60%)
Length = 421
Score = 73.9 bits (180), Expect(2) = 3e-18
Identities = 32/61 (52%), Positives = 50/61 (81%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N S+RQVT+SKR+ G+ KKA+E++ LCDA ++LI+F+A+ K+ +Y S
Sbjct: 39 MGRGKIEIKRIENSSNRQVTYSKRKNGILKKAKEITVLCDAQVSLIIFAASGKMHDYISP 218
Query: 61 S 61
S
Sbjct: 219S 221
Score = 34.7 bits (78), Expect(2) = 3e-18
Identities = 17/44 (38%), Positives = 25/44 (56%)
Frame = +2
Query: 94 DTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASV 137
+ L+K+ + ELR L GED+ L +L LE+ L+ L SV
Sbjct: 245 ERLKKENDSMQIELRHLKGEDINSLNYKELMALEDALETGLVSV 376
>TC207768 similar to UP|Q9ST53 (Q9ST53) MADS-box protein 4, partial (27%)
Length = 951
Score = 87.8 bits (216), Expect = 4e-18
Identities = 55/173 (31%), Positives = 94/173 (53%), Gaps = 3/173 (1%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M +K+++IK+I+N S+RQ+TFSKRR GL KKA+ELS LCDA +AL++FS+T KL+E +
Sbjct: 111 MGKKKVEIKRIENKSTRQITFSKRRNGLMKKARELSILCDAKVALLIFSSTGKLYELCNG 290
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESD---SNDTLRKKVEEKTHELRQLNGEDLQG 117
+ + T S + FE S + ++ R +L+
Sbjct: 291 DSLAEVVQQYWDHLGASGTDTKSQELCFEIADIWSGSAFSQMIK------RHFGVSELEH 452
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
L++ L +LE++ +L+ + K M+ + LK+K +E +E+ + V+
Sbjct: 453 LSVSDLMELEKLTHAALSRIRSAKMRLMMESVVNLKKK-IEALEKTNDVNNVV 608
>TC228368 similar to UP|O49173 (O49173) MADS-box protein NMH 7, partial (97%)
Length = 1042
Score = 85.5 bits (210), Expect = 2e-17
Identities = 60/174 (34%), Positives = 87/174 (49%), Gaps = 19/174 (10%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +IQIK+I+N ++RQVT+SKRR GLFKKA EL+ LCDA +++I+FS+T KL +Y S
Sbjct: 81 MARGKIQIKRIENNTNRQVTYSKRRNGLFKKANELTVLCDAKVSIIMFSSTGKLHQYIS- 257
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTL---------------RKKVEEKTH 105
P + T QF TL KK++E
Sbjct: 258 --------------------PSTSTKQFFDQYQMTLGVDLWNSHYENMQENLKKLKEVNR 377
Query: 106 ----ELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRK 155
E+RQ G+ L L + L+ LEE + ++ V K + +I T ++K
Sbjct: 378 NLRKEIRQRMGDCLNELGMEDLKLLEEEMDKAAKVVRERKYKVITNQIDTQRKK 539
>TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragment), partial
(64%)
Length = 710
Score = 85.5 bits (210), Expect = 2e-17
Identities = 52/123 (42%), Positives = 68/123 (55%), Gaps = 1/123 (0%)
Frame = +1
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R Q+++I+N +SRQVTFSKRR GL KKA ELS LCDA++ALI+FS KL+E+ASS
Sbjct: 340 MVRGETQMRRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIIFSPRGKLYEFASS 519
Query: 61 SRGGLLP-LPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S + N + MQ L KK+E R+L GE L +
Sbjct: 520 SMQDTIERYRRHNRSAQTVNRSDEQNMQHLKQETANLMKKIELLEASKRKLLGEGLGSCS 699
Query: 120 LHQ 122
L +
Sbjct: 700 LEE 708
>BG651806
Length = 309
Score = 84.0 bits (206), Expect = 6e-17
Identities = 40/61 (65%), Positives = 52/61 (84%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I++IDN +SRQVTFSKRR GL KKA+ELS LCDA++ L+VFS+T KL++YAS+
Sbjct: 120 MGRGKIAIRRIDNSTSRQVTFSKRRNGLLKKARELSILCDAEVGLMVFSSTGKLYDYAST 299
Query: 61 S 61
S
Sbjct: 300 S 302
>AW706936
Length = 422
Score = 84.0 bits (206), Expect = 6e-17
Identities = 40/61 (65%), Positives = 52/61 (84%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I++IDN +SRQVTFSKRR GL KKA+ELS LCDA++ L+VFS+T KL++YAS+
Sbjct: 131 MGRGKIAIRRIDNSTSRQVTFSKRRNGLLKKARELSILCDAEVGLMVFSSTGKLYDYAST 310
Query: 61 S 61
S
Sbjct: 311 S 313
>CD412950 weakly similar to SP|Q9S7Q7|FLC_A FLOWERING LOCUS C protein (MADS
box protein FLOWERING LOCUS F). [Mouse-ear cress],
partial (41%)
Length = 674
Score = 82.8 bits (203), Expect = 1e-16
Identities = 38/66 (57%), Positives = 53/66 (79%)
Frame = -3
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
+K+++IK+I+N S+RQ+TFSKRRKGL KKA+ELS LCDA +AL++FS+T KL+E + R
Sbjct: 669 KKKLEIKRIENKSNRQITFSKRRKGLMKKARELSILCDAKLALLIFSSTGKLYELCNGDR 490
Query: 63 GGLLPL 68
L L
Sbjct: 489 DSALKL 472
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,176,111
Number of Sequences: 63676
Number of extensions: 93595
Number of successful extensions: 595
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 591
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 593
length of query: 244
length of database: 12,639,632
effective HSP length: 95
effective length of query: 149
effective length of database: 6,590,412
effective search space: 981971388
effective search space used: 981971388
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0100a.3


