

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
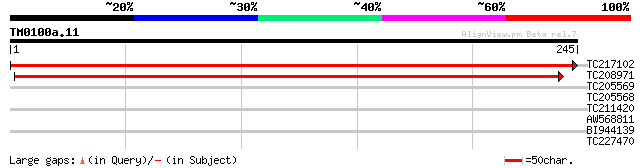
Score E
Sequences producing significant alignments: (bits) Value
TC217102 weakly similar to UP|Q6K1Z7 (Q6K1Z7) Isochorismatase hy... 454 e-128
TC208971 327 3e-90
TC205569 similar to UP|MPPA_SOLTU (P29677) Mitochondrial process... 30 1.3
TC205568 similar to UP|MPPA_SOLTU (P29677) Mitochondrial process... 29 1.7
TC211420 similar to UP|Q8L470 (Q8L470) Small heat shock protein,... 28 3.8
AW568811 28 3.8
BI944139 27 8.5
TC227470 similar to UP|Q8L7V7 (Q8L7V7) AT4g02010/T10M13_2, parti... 27 8.5
>TC217102 weakly similar to UP|Q6K1Z7 (Q6K1Z7) Isochorismatase hydrolase-like
protein, partial (82%)
Length = 1178
Score = 454 bits (1169), Expect = e-128
Identities = 219/245 (89%), Positives = 232/245 (94%)
Frame = +2
Query: 1 MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 60
MVSQ +ELLKNEIPLEQE VVLAED VNGLVLVDIINGFCTVGAGNLAPRESN QIS MI
Sbjct: 50 MVSQTVELLKNEIPLEQESVVLAEDAVNGLVLVDIINGFCTVGAGNLAPRESNTQISGMI 229
Query: 61 NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT 120
+ESARLAR+FCEKNLPVMAFLDSHHPNKPEDPYPPHCI G+DESNLVPALRWLENE NVT
Sbjct: 230 SESARLARVFCEKNLPVMAFLDSHHPNKPEDPYPPHCIVGSDESNLVPALRWLENEPNVT 409
Query: 121 LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 180
+RRK+CFDGY+GS +EDGSNVFVDWVKKNKI TLLVVG+CTDICVLDFVCSTMSAKNRGF
Sbjct: 410 IRRKDCFDGYLGSIQEDGSNVFVDWVKKNKITTLLVVGVCTDICVLDFVCSTMSAKNRGF 589
Query: 181 LKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLFG 240
L+PLENVVVYS CATF+VPLEVARNTKGALAHPQEF+HH+GLYMAKERGAKIANEVLFG
Sbjct: 590 LEPLENVVVYSRACATFNVPLEVARNTKGALAHPQEFMHHVGLYMAKERGAKIANEVLFG 769
Query: 241 AVEKV 245
A KV
Sbjct: 770 AAGKV 784
>TC208971
Length = 956
Score = 327 bits (838), Expect = 3e-90
Identities = 147/237 (62%), Positives = 188/237 (79%)
Frame = +2
Query: 3 SQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINE 62
S ELL+ EIP++Q+ + L+ D + GLVLVD++NGFCTVGAGNLAP+E + +IS+M+ E
Sbjct: 26 SNTAELLREEIPVKQQPLTLSADIITGLVLVDVVNGFCTVGAGNLAPKEPDERISQMVKE 205
Query: 63 SARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLR 122
S RL++ F E+ P+ AFLD HHP+KPE PYPPHCI G+ E LVP L WLEN+ N TLR
Sbjct: 206 SLRLSKAFSERKWPIFAFLDCHHPDKPEPPYPPHCIIGSGEEKLVPDLLWLENDPNATLR 385
Query: 123 RKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLK 182
+KEC DG++GS E+DGSNVF+DWVK N+I+ +LV GICTDICVLDFV S +S +NRGFL
Sbjct: 386 QKECIDGFLGSTEKDGSNVFIDWVKNNQIKQILVAGICTDICVLDFVSSVLSVRNRGFLT 565
Query: 183 PLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
PLENV+V S CAT+D+PL VA+ K ++HPQE +HH+GLY+A RGA IA+EVLF
Sbjct: 566 PLENVIVSSQACATYDLPLHVAKTNKDFVSHPQELMHHVGLYIASGRGAHIASEVLF 736
>TC205569 similar to UP|MPPA_SOLTU (P29677) Mitochondrial processing
peptidase alpha subunit, mitochondrial precursor
(Alpha-MPP) (Ubiquinol-cytochrome-c reductase subunit
II) , partial (19%)
Length = 497
Score = 29.6 bits (65), Expect = 1.3
Identities = 23/58 (39%), Positives = 28/58 (47%), Gaps = 2/58 (3%)
Frame = -3
Query: 85 HPNKPEDPYPPHCIAGTDESNLVPA--LRWLENETNVTLRRKECFDGYIGSAEEDGSN 140
HPNKP P C+A T+ NL A L L +TLRR+ F GS D S+
Sbjct: 243 HPNKPPKPEEDCCVA-TELENLALAGILLLLHERALMTLRREAAFLYMSGSRIFDFSS 73
>TC205568 similar to UP|MPPA_SOLTU (P29677) Mitochondrial processing
peptidase alpha subunit, mitochondrial precursor
(Alpha-MPP) (Ubiquinol-cytochrome-c reductase subunit
II) , partial (93%)
Length = 1943
Score = 29.3 bits (64), Expect = 1.7
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 3/46 (6%)
Frame = -3
Query: 85 HPNKPEDPYPPHCIAGTDESNLVPA---LRWLENETNVTLRRKECF 127
HPNKP P C T+ +NL A L L + + LRR+ F
Sbjct: 306 HPNKPPKPEEEDCCVATELANLALAGTLLLLLHDGALMALRREAAF 169
>TC211420 similar to UP|Q8L470 (Q8L470) Small heat shock protein, partial
(33%)
Length = 514
Score = 28.1 bits (61), Expect = 3.8
Identities = 15/40 (37%), Positives = 22/40 (54%), Gaps = 7/40 (17%)
Frame = -2
Query: 66 LARLFCEKNLPVMAFLDSHH-------PNKPEDPYPPHCI 98
LA L+C +NLP+ F+ S + N+ P+P HCI
Sbjct: 252 LAFLYCPRNLPLTVFIISRNYHR*SSTINQEILPHPLHCI 133
>AW568811
Length = 394
Score = 28.1 bits (61), Expect = 3.8
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -2
Query: 128 DGYIGSAEEDGSNVFVDWVKKNK 150
D +GSA + SNVF+ W K NK
Sbjct: 294 DRLVGSAPYNESNVFLSWQKTNK 226
>BI944139
Length = 405
Score = 26.9 bits (58), Expect = 8.5
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = -3
Query: 106 LVPALRWLENETNVTLRRKECFDGYIGSAEEDGS 139
L+ L WL T R++ CF GY+ + +GS
Sbjct: 199 LLQLLSWLYFFTLPCFRQRSCFKGYLDISASEGS 98
>TC227470 similar to UP|Q8L7V7 (Q8L7V7) AT4g02010/T10M13_2, partial (34%)
Length = 1061
Score = 26.9 bits (58), Expect = 8.5
Identities = 18/72 (25%), Positives = 35/72 (48%)
Frame = +1
Query: 8 LLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLA 67
+L+++ LE+ LA+ + G + CT+ A +AP S R + +S ++
Sbjct: 397 ILRDKDRLEE----LADPRLGGRYPKEDFVRVCTIAAACVAPEASQRPTMGEVVQSLKMV 564
Query: 68 RLFCEKNLPVMA 79
+ E + PV+A
Sbjct: 565 QRITESHDPVLA 600
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,714,838
Number of Sequences: 63676
Number of extensions: 170740
Number of successful extensions: 745
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 739
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 745
length of query: 245
length of database: 12,639,632
effective HSP length: 95
effective length of query: 150
effective length of database: 6,590,412
effective search space: 988561800
effective search space used: 988561800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0100a.11


