

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0096b.3
(223 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
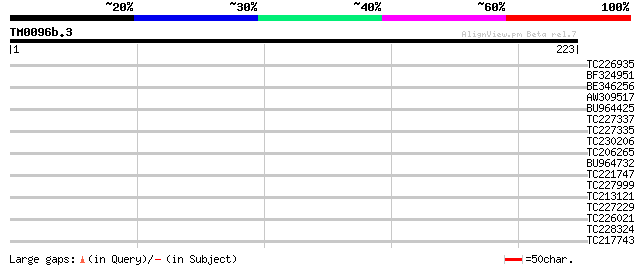
Score E
Sequences producing significant alignments: (bits) Value
TC226935 similar to UP|Q9LNV5 (Q9LNV5) F22G5.30, partial (11%) 31 0.51
BF324951 similar to GP|28416657|gb| At5g60800 {Arabidopsis thali... 30 0.87
BE346256 similar to SP|Q41387|PSBW_ Photosystem II reaction cent... 29 1.9
AW309517 28 4.3
BU964425 homologue to GP|3287679|gb T22J18.6 {Arabidopsis thalia... 28 4.3
TC227337 similar to UP|Q9LP69 (Q9LP69) T1N15.17, partial (37%) 28 4.3
TC227335 similar to UP|Q9LP69 (Q9LP69) T1N15.17, partial (16%) 28 4.3
TC230206 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {A... 27 5.6
TC206265 weakly similar to UP|PLB2_YEAST (Q03674) Lysophospholip... 27 5.6
BU964732 27 7.4
TC221747 weakly similar to UP|Q8L6T7 (Q8L6T7) Ring-H2 zinc finge... 27 7.4
TC227999 27 7.4
TC213121 27 7.4
TC227229 similar to UP|Q712P1 (Q712P1) BZIP protein BZ3, partial... 27 9.6
TC226021 similar to UP|Q9LRZ0 (Q9LRZ0) Arabidopsis thaliana geno... 27 9.6
TC228324 UP|LGUL_SOYBN (P46417) Lactoylglutathione lyase (Methy... 27 9.6
TC217743 similar to UP|Q8L751 (Q8L751) AT4g08180/T12G13_20, part... 27 9.6
>TC226935 similar to UP|Q9LNV5 (Q9LNV5) F22G5.30, partial (11%)
Length = 584
Score = 30.8 bits (68), Expect = 0.51
Identities = 12/34 (35%), Positives = 22/34 (64%)
Frame = +1
Query: 166 PINPHPIHILEFLGSIIIHLLMCYSLLLSNSSHI 199
PI +H+ + SII+H+++C+ L NS+H+
Sbjct: 295 PIQ*CHLHLANIIISIILHMVICHQFPLINSTHL 396
>BF324951 similar to GP|28416657|gb| At5g60800 {Arabidopsis thaliana},
partial (21%)
Length = 316
Score = 30.0 bits (66), Expect = 0.87
Identities = 17/57 (29%), Positives = 26/57 (44%), Gaps = 4/57 (7%)
Frame = -2
Query: 142 VFCIGFIIKTFYQPPPPNDYSCKEPINPHPIHI----LEFLGSIIIHLLMCYSLLLS 194
+ CI I +F PPPP + +P +P P + L F G L+ +S+ S
Sbjct: 276 ILCIPCSIFSFLSPPPPLLFFLPDPFSPPPPSLLSPSLSFWGGTTSTFLLSFSMRFS 106
>BE346256 similar to SP|Q41387|PSBW_ Photosystem II reaction center W protein
chloroplast precursor (PSII 6.1 kDa protein).
[Spinach], partial (53%)
Length = 403
Score = 28.9 bits (63), Expect = 1.9
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 2/43 (4%)
Frame = -2
Query: 51 RGMLSPHIHHRMQDVGALVELDPGTPVPEVLALDS--SLILPD 91
R +SP +R+ G L+ G+PVP+V++L S ++I PD
Sbjct: 348 RAQMSPKTRNRIHPRGLLIS-PKGSPVPQVISLSSTKAMIEPD 223
>AW309517
Length = 453
Score = 27.7 bits (60), Expect = 4.3
Identities = 12/31 (38%), Positives = 21/31 (67%)
Frame = +1
Query: 192 LLSNSSHIVMFSKLTSLEKGKSEVIGESQRR 222
LLS+ ++ + L S EKG ++V+G+ +RR
Sbjct: 253 LLSSD*KLITMATLNSTEKGPAKVVGQVKRR 345
>BU964425 homologue to GP|3287679|gb T22J18.6 {Arabidopsis thaliana}, partial
(9%)
Length = 313
Score = 27.7 bits (60), Expect = 4.3
Identities = 22/74 (29%), Positives = 33/74 (43%)
Frame = -2
Query: 8 GHEEPILEVLVRMARVVSLVVVDDVVYHLQLLSGNTNGLSIQTRGMLSPHIHHRMQDVGA 67
GH EP++ VL++ V +VV+ L S N I +L P I R
Sbjct: 309 GHPEPMMGVLLK---------VQEVVFELH*ESQWHNICEINVHMVLEPVIQQRDAHPKL 157
Query: 68 LVELDPGTPVPEVL 81
L+E + G P ++L
Sbjct: 156 LLEEEVGVPGQQLL 115
>TC227337 similar to UP|Q9LP69 (Q9LP69) T1N15.17, partial (37%)
Length = 965
Score = 27.7 bits (60), Expect = 4.3
Identities = 12/31 (38%), Positives = 21/31 (67%)
Frame = -3
Query: 192 LLSNSSHIVMFSKLTSLEKGKSEVIGESQRR 222
LLS+ ++ + L S EKG ++V+G+ +RR
Sbjct: 705 LLSSD*KLITMATLNSTEKGPAKVVGQVKRR 613
>TC227335 similar to UP|Q9LP69 (Q9LP69) T1N15.17, partial (16%)
Length = 713
Score = 27.7 bits (60), Expect = 4.3
Identities = 12/31 (38%), Positives = 21/31 (67%)
Frame = -1
Query: 192 LLSNSSHIVMFSKLTSLEKGKSEVIGESQRR 222
LLS+ ++ + L S EKG ++V+G+ +RR
Sbjct: 179 LLSSD*KLITMATLNSTEKGPAKVVGQVKRR 87
>TC230206 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;} , partial (28%)
Length = 1305
Score = 27.3 bits (59), Expect = 5.6
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = +1
Query: 156 PPPNDYSCKEPINPHPIHILEFLGSIIIHLLM--CYSLLLSNSSHIVMFSKL 205
PP DYS PI P+H+L L I + L+ +S + S S H++M + +
Sbjct: 463 PPACDYSSFPPILFLPLHLLLLLHPIRLSNLITSIFSSVSSLSHHLLMSNSM 618
>TC206265 weakly similar to UP|PLB2_YEAST (Q03674) Lysophospholipase 2
precursor (Phospholipase B 2) , partial (3%)
Length = 742
Score = 27.3 bits (59), Expect = 5.6
Identities = 12/31 (38%), Positives = 18/31 (57%), Gaps = 2/31 (6%)
Frame = +2
Query: 163 CKEPINPHPIHI--LEFLGSIIIHLLMCYSL 191
C + + P+ I L F+G +IIH +CY L
Sbjct: 560 CSQVSSQQPVEICMLHFMGIVIIHKSLCYIL 652
>BU964732
Length = 451
Score = 26.9 bits (58), Expect = 7.4
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +3
Query: 152 FYQPPPPNDYSCKEPINPHPIHILEFLGSIIIHL 185
+ PPPP++ C I PI L F G I+I +
Sbjct: 333 YLPPPPPHNTQCFSCIVKCPIFNLCFTGEILIFI 434
>TC221747 weakly similar to UP|Q8L6T7 (Q8L6T7) Ring-H2 zinc finger protein,
partial (16%)
Length = 613
Score = 26.9 bits (58), Expect = 7.4
Identities = 17/48 (35%), Positives = 24/48 (49%)
Frame = +3
Query: 151 TFYQPPPPNDYSCKEPINPHPIHILEFLGSIIIHLLMCYSLLLSNSSH 198
T PPPP+ S K P + L +G+ I +L Y+L+L SH
Sbjct: 57 TISSPPPPSSPSTKSNSMPMLYYGLVVVGTAAI-VLAIYNLVLLRRSH 197
>TC227999
Length = 1046
Score = 26.9 bits (58), Expect = 7.4
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = -1
Query: 59 HHRMQDVGALVELDPGTPVPEVLALDSSLIL 89
HH A+ L P TP+PE L+L +L
Sbjct: 533 HH*SHPESAIPSLSPATPLPETLSLKPKHLL 441
>TC213121
Length = 713
Score = 26.9 bits (58), Expect = 7.4
Identities = 15/35 (42%), Positives = 21/35 (59%), Gaps = 1/35 (2%)
Frame = +2
Query: 119 TDEEA-IESYAIHLVANCHGGVYLVFCIGFIIKTF 152
TDE A I + +H++A GGVY ++ G KTF
Sbjct: 413 TDEHASIFNKIMHMIATQSGGVYFLYGYGGTGKTF 517
>TC227229 similar to UP|Q712P1 (Q712P1) BZIP protein BZ3, partial (24%)
Length = 460
Score = 26.6 bits (57), Expect = 9.6
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +3
Query: 155 PPPPNDYSCKEPINPHP 171
PPPP YS + I+P+P
Sbjct: 216 PPPPTSYSWRSMISPNP 266
>TC226021 similar to UP|Q9LRZ0 (Q9LRZ0) Arabidopsis thaliana genomic DNA,
chromosome 3, TAC clone:K20I9, partial (29%)
Length = 1585
Score = 26.6 bits (57), Expect = 9.6
Identities = 14/43 (32%), Positives = 23/43 (52%), Gaps = 5/43 (11%)
Frame = +3
Query: 121 EEAIESYAI-----HLVANCHGGVYLVFCIGFIIKTFYQPPPP 158
EEA+ A+ ++++ C G V L + F + T + PPPP
Sbjct: 498 EEAVPCIALSKNDSYVMSACGGKVSLFNMMTFKVMTTFMPPPP 626
>TC228324 UP|LGUL_SOYBN (P46417) Lactoylglutathione lyase (Methylglyoxalase)
(Aldoketomutase) (Glyoxalase I) , complete
Length = 995
Score = 26.6 bits (57), Expect = 9.6
Identities = 26/87 (29%), Positives = 39/87 (43%), Gaps = 2/87 (2%)
Frame = -3
Query: 9 HEEPILEVLVRMARVVSLVVVDDVVYHLQLLSGNTNGLSI--QTRGMLSPHIHHRMQDVG 66
H E L LVR+ +V+ D++ L TNG +I Q R +L +H + Q
Sbjct: 366 HPESGLRSLVRIGGHEWRLVIPDLINVLHYYCRFTNGFAIMDQNRNLLVNWVHLQKQ--- 196
Query: 67 ALVELDPGTPVPEVLALDSSLILPDAL 93
TPV E+L L + +P+ L
Sbjct: 195 -------RTPVHEILFL---VFIPNTL 145
>TC217743 similar to UP|Q8L751 (Q8L751) AT4g08180/T12G13_20, partial (23%)
Length = 994
Score = 26.6 bits (57), Expect = 9.6
Identities = 14/40 (35%), Positives = 19/40 (47%), Gaps = 8/40 (20%)
Frame = +2
Query: 160 DYSCKEPI--------NPHPIHILEFLGSIIIHLLMCYSL 191
+YSCK NPH + + + SI HL+ CY L
Sbjct: 8 EYSCKLKFKEQSIIDRNPHQVLYICCMDSIFSHLIYCYFL 127
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.142 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,214,178
Number of Sequences: 63676
Number of extensions: 182804
Number of successful extensions: 1371
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1217
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1370
length of query: 223
length of database: 12,639,632
effective HSP length: 94
effective length of query: 129
effective length of database: 6,654,088
effective search space: 858377352
effective search space used: 858377352
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0096b.3


