

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0095c.6
(384 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
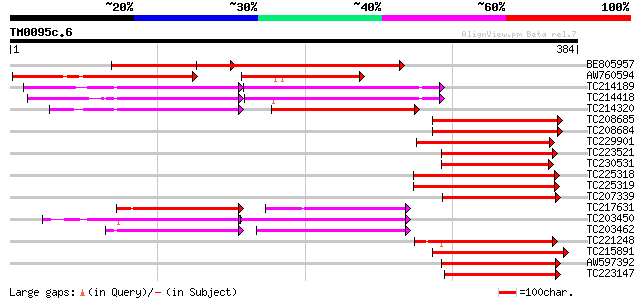
Score E
Sequences producing significant alignments: (bits) Value
BE805957 258 2e-69
AW760594 176 2e-44
TC214189 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 102 4e-22
TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 98 7e-21
TC214320 similar to UP|O22793 (O22793) Plastid protein (At2g3343... 91 7e-19
TC208685 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition mo... 91 7e-19
TC208684 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition mo... 91 7e-19
TC229901 weakly similar to UP|GRP1_SORBI (Q99069) Glycine-rich R... 88 6e-18
TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition mo... 85 6e-17
TC230531 weakly similar to UP|Q941H9 (Q941H9) RNA-binding protei... 79 3e-15
TC225318 similar to UP|P93486 (P93486) Glycine-rich RNA-binding ... 77 1e-14
TC225319 similar to UP|P93486 (P93486) Glycine-rich RNA-binding ... 77 2e-14
TC207339 similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif (RR... 76 2e-14
TC217631 similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldrich syndr... 75 6e-14
TC203450 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chlorop... 75 6e-14
TC203462 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chlorop... 73 2e-13
TC221248 weakly similar to GB|CAD40580.1|21741678|OSJN00141 oj00... 72 4e-13
TC215891 similar to GB|AAM78058.1|21928059|AY125548 AT5g61030/ma... 72 4e-13
AW597392 weakly similar to GP|7630030|emb| RNA binding protein-l... 72 5e-13
TC223147 weakly similar to UP|Q9LZT1 (Q9LZT1) RNA binding protei... 70 1e-12
>BE805957
Length = 426
Score = 258 bits (660), Expect = 2e-69
Identities = 126/141 (89%), Positives = 133/141 (93%)
Frame = +3
Query: 127 DIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSKH 186
DIDEEIS+QLASLP VL VRPD +FNSLKKDYSLSSG+AG SGL+TRTNMLFPAGNSKH
Sbjct: 3 DIDEEISAQLASLPEVLLVRPDLEFNSLKKDYSLSSGEAGHLSGLRTRTNMLFPAGNSKH 182
Query: 187 WLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCA 246
WLV+MDKPGVE VTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCA
Sbjct: 183 WLVKMDKPGVEAVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCA 362
Query: 247 QDLAGVPGVSSVQPDDNFESE 267
Q+LAGV GV SVQPD+NFESE
Sbjct: 363 QELAGVLGVLSVQPDNNFESE 425
Score = 110 bits (275), Expect = 1e-24
Identities = 44/84 (52%), Positives = 67/84 (79%)
Frame = +3
Query: 70 HNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDID 129
+++HW+V MDKP + ++K+Q++DYY + L VMG+EK+AQMC+Y SW T+FGFCC++D
Sbjct: 171 NSKHWLVKMDKPGVEAVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELD 350
Query: 130 EEISSQLASLPGVLSVRPDPDFNS 153
E+ + +LA + GVLSV+PD +F S
Sbjct: 351 EDCAQELAGVLGVLSVQPDNNFES 422
Score = 34.3 bits (77), Expect = 0.097
Identities = 26/63 (41%), Positives = 35/63 (55%), Gaps = 2/63 (3%)
Frame = +3
Query: 240 ELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPNSSEASQEASLRTK--KLFVT 297
++DE+ + LA +P V V+PD F S K+Y SLS S EA + LRT+ LF
Sbjct: 3 DIDEEISAQLASLPEVLLVRPDLEFNSLKKDY--SLS---SGEAGHLSGLRTRTNMLFPA 167
Query: 298 GLS 300
G S
Sbjct: 168 GNS 176
>AW760594
Length = 424
Score = 176 bits (446), Expect = 2e-44
Identities = 86/125 (68%), Positives = 98/125 (77%)
Frame = +1
Query: 3 TKCQKISSVNPNRSQVHQLPTNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYP 62
+KCQ +S P Q+H+LP NI LPNK+ H +L SSSSS CN ++I AAATTYP
Sbjct: 61 SKCQTLSFPKPKAPQIHELPLNIRLPNKTNHFSL--SSSSSPCCN--WSITRVAAATTYP 228
Query: 63 SFTPSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHF 122
SF PSTT N HWMVLMD PP GV SK QVIDYYVKTLQTV+GSEK+AQMC+YDASW+THF
Sbjct: 229 SFNPSTTQNPHWMVLMDTPPQGVNSKPQVIDYYVKTLQTVLGSEKDAQMCIYDASWNTHF 408
Query: 123 GFCCD 127
GFCCD
Sbjct: 409 GFCCD 423
Score = 79.7 bits (195), Expect = 2e-15
Identities = 40/90 (44%), Positives = 55/90 (60%), Gaps = 7/90 (7%)
Frame = +1
Query: 158 YSLSSGQAGLSSGLQTRTNML--FPAGN-----SKHWLVRMDKPGVEVVTKAQIVDYYAQ 210
+SLSS + + TR +P+ N + HW+V MD P V +K Q++DYY +
Sbjct: 154 FSLSSSSSPCCNWSITRVAAATTYPSFNPSTTQNPHWMVLMDTPPQGVNSKPQVIDYYVK 333
Query: 211 ILTKVMGNEKDAQMCIYHVSWKTNFGFCCE 240
L V+G+EKDAQMCIY SW T+FGFCC+
Sbjct: 334 TLQTVLGSEKDAQMCIYDASWNTHFGFCCD 423
>TC214189 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (71%)
Length = 969
Score = 102 bits (253), Expect = 4e-22
Identities = 56/136 (41%), Positives = 79/136 (57%)
Frame = +2
Query: 159 SLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGN 218
S SS + S T LFP + HWL+ MD PG E TK Q++D Y Q L KV+G+
Sbjct: 293 SASSSSSSFSDRPPTEMAPLFPGCDYNHWLIVMDHPGGEGATKQQMIDCYIQTLAKVLGS 472
Query: 219 EKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVP 278
E++A+ IY+VS + FGF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 473 EEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDPENKDYGAELFV- 649
Query: 279 NSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 650 -NGEIVQRSPERQRRV 694
Score = 92.0 bits (227), Expect = 4e-19
Identities = 55/149 (36%), Positives = 83/149 (54%)
Frame = +2
Query: 10 SVNPNRSQVHQLPTNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTT 69
++ P ++V + + S +S L S+SSSS S + T P
Sbjct: 209 AIVPPSTRVAGIRCRVNRAGDSAYSPLNSASSSSSS-------FSDRPPTEMAPLFPGCD 367
Query: 70 HNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDID 129
+N HW+++MD P +K Q+ID Y++TL V+GSE+EA+ +Y+ S + +FGF C+ID
Sbjct: 368 YN-HWLIVMDHPGGEGATKQQMIDCYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEID 544
Query: 130 EEISSQLASLPGVLSVRPDPDFNSLKKDY 158
EE S++L LPGVL V PD + KDY
Sbjct: 545 EETSNKLEGLPGVLFVLPDSYVDPENKDY 631
>TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (74%)
Length = 1130
Score = 97.8 bits (242), Expect = 7e-21
Identities = 55/137 (40%), Positives = 81/137 (58%), Gaps = 2/137 (1%)
Frame = +3
Query: 160 LSSGQAGLSSGLQTRTNM--LFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
L+SG + S + T M LFP + HWL+ M+ PG E K Q++D Y Q L KV+G
Sbjct: 72 LNSGSSSSSFSDRPPTEMAPLFPGCDYNHWLIVMENPGGEGANKQQMIDCYIQTLAKVLG 251
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + FGF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 252 SEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDPENKDYGAELFV 431
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 432 --NGEIVQRSPERQRRV 476
Score = 90.9 bits (224), Expect = 9e-19
Identities = 57/146 (39%), Positives = 84/146 (57%)
Frame = +3
Query: 13 PNRSQVHQLPTNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTTHNR 72
P ++V + + S +S L S SSSS +R T +A P F P +N
Sbjct: 3 PPSTRVAGIRCRVNRAGDSAYSPLNSGSSSSSFSDRPPTEMA-------PLF-PGCDYN- 155
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++M+ P +K Q+ID Y++TL V+GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 156 HWLIVMENPGGEGANKQQMIDCYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEET 335
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD + KDY
Sbjct: 336 SNKLEGLPGVLFVLPDSYVDPENKDY 413
>TC214320 similar to UP|O22793 (O22793) Plastid protein (At2g33430/F4P9.20),
partial (68%)
Length = 1045
Score = 91.3 bits (225), Expect = 7e-19
Identities = 45/100 (45%), Positives = 64/100 (64%)
Frame = +3
Query: 178 LFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGF 237
LFP + HWLV + KPG E TK Q++D Y Q L KV+G+E++A+ IY+VS + F F
Sbjct: 354 LFPGCDYHHWLVVIHKPGGEGATKQQMIDCYVQTLAKVLGSEEEAKKKIYNVSCERYFAF 533
Query: 238 CCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
C++DE+ + L G+PGV V PD + E+K+Y L V
Sbjct: 534 GCDIDEETSNKLEGLPGVLFVLPDSYVDPEHKDYGGELFV 653
Score = 84.7 bits (208), Expect = 6e-17
Identities = 51/131 (38%), Positives = 73/131 (54%)
Frame = +3
Query: 28 PNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTTHNRHWMVLMDKPPLGVIS 87
P S +S L + +SI + I T P ++ HW+V++ KP +
Sbjct: 261 PGDSAYSPLNTGKPTSI-----LSSITERPPTDMAPLFPGCDYH-HWLVVIHKPGGEGAT 422
Query: 88 KSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRP 147
K Q+ID YV+TL V+GSE+EA+ +Y+ S + +F F CDIDEE S++L LPGVL V P
Sbjct: 423 KQQMIDCYVQTLAKVLGSEEEAKKKIYNVSCERYFAFGCDIDEETSNKLEGLPGVLFVLP 602
Query: 148 DPDFNSLKKDY 158
D + KDY
Sbjct: 603 DSYVDPEHKDY 635
>TC208685 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (59%)
Length = 729
Score = 91.3 bits (225), Expect = 7e-19
Identities = 45/88 (51%), Positives = 62/88 (70%)
Frame = +1
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
++L + KLFV+GL T+++ L+ AF FG+LV+ KVIID+ S RSKG+AF+ YTT E A
Sbjct: 106 STLTSPKLFVSGLCRLTTDEKLKEAFSSFGQLVEAKVIIDRASGRSKGFAFVTYTTIEEA 285
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPR 374
A + MN K ++GW+I VD AK PR
Sbjct: 286 EKAREGMNAKFLDGWVIFVDPAKPREPR 369
>TC208684 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (59%)
Length = 791
Score = 91.3 bits (225), Expect = 7e-19
Identities = 45/88 (51%), Positives = 61/88 (69%)
Frame = +3
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
++L + KLFV+GLS T ++ L+ AF FG+LV+ KVI D+ S RSKG+AF+ YTT E A
Sbjct: 174 STLTSPKLFVSGLSRLTKDENLKEAFSSFGQLVEAKVITDRASGRSKGFAFVTYTTIEEA 353
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPR 374
A + MN K ++GW+I VD AK PR
Sbjct: 354 ERAREGMNAKFLDGWVIFVDPAKPREPR 437
>TC229901 weakly similar to UP|GRP1_SORBI (Q99069) Glycine-rich RNA-binding
protein 1 (Fragment), partial (28%)
Length = 673
Score = 88.2 bits (217), Expect = 6e-18
Identities = 43/94 (45%), Positives = 68/94 (71%)
Frame = +3
Query: 276 SVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGY 335
S +SS A KL+V+GLSF T+E++LR AF+ FG+LV+VK+++D+I+ R +G+
Sbjct: 219 SSTSSSTAKPSFPSPQTKLYVSGLSFRTTEESLRNAFKNFGQLVEVKLVMDRIANRPRGF 398
Query: 336 AFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAK 369
AF+ Y TEE + A++ M+GK ++G +I V+VAK
Sbjct: 399 AFLRYATEEESQKAIEGMHGKFLDGRVIFVEVAK 500
>TC223521 weakly similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (57%)
Length = 651
Score = 84.7 bits (208), Expect = 6e-17
Identities = 36/79 (45%), Positives = 57/79 (71%)
Frame = +1
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSFYT+E L AF +G++++ K++ D++S RSKG+ F+ + +++ A A+++
Sbjct: 82 KLFVGGLSFYTTENALSEAFSNYGQVIEAKIVTDRVSDRSKGFGFVTFASQDEAENAIED 261
Query: 353 MNGKIINGWMIVVDVAKTN 371
M GK +NG +I VD AK N
Sbjct: 262 MKGKTLNGRVIFVDYAKPN 318
>TC230531 weakly similar to UP|Q941H9 (Q941H9) RNA-binding protein precursor,
partial (29%)
Length = 851
Score = 79.0 bits (193), Expect = 3e-15
Identities = 39/76 (51%), Positives = 54/76 (70%)
Frame = +3
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFVTGLS+ T+E LR AF GE+++VKVI D ++ +S+GY F+ + +E A AA KE
Sbjct: 417 KLFVTGLSYDTNEPILRDAFGQHGEIIEVKVICDHVTGKSRGYGFVRFVSETTAAAARKE 596
Query: 353 MNGKIINGWMIVVDVA 368
MNG+I++G I V A
Sbjct: 597 MNGQILDGRRIRVSYA 644
>TC225318 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (86%)
Length = 749
Score = 77.0 bits (188), Expect = 1e-14
Identities = 38/99 (38%), Positives = 61/99 (61%)
Frame = +2
Query: 274 SLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSK 333
S P +S + + + KLF+ GLS+ +++L+ AF GFG++V KVI D+ S RS+
Sbjct: 116 STHAPVASMLNYIRCMSSSKLFIGGLSYGVDDQSLKDAFSGFGDVVDAKVITDRDSGRSR 295
Query: 334 GYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNP 372
G+ F+ ++ +E+A +AL M+GK +NG I V A P
Sbjct: 296 GFGFVNFSNDESASSALSAMDGKDLNGRSIRVSYANDKP 412
>TC225319 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 1167
Score = 76.6 bits (187), Expect = 2e-14
Identities = 38/99 (38%), Positives = 61/99 (61%)
Frame = +3
Query: 274 SLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSK 333
S P SS + + + KLF+ GLS+ +++L+ AF GFG++V KVI D+ S RS+
Sbjct: 312 STQAPVSSMLNYIRCMSSSKLFIGGLSYGVDDQSLKDAFSGFGDVVDAKVITDRDSGRSR 491
Query: 334 GYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNP 372
G+ F+ ++ +E+A +AL M+GK ++G I V A P
Sbjct: 492 GFGFVNFSNDESASSALSAMDGKDLDGRSIRVSYANDRP 608
>TC207339 similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (63%)
Length = 838
Score = 76.3 bits (186), Expect = 2e-14
Identities = 34/80 (42%), Positives = 53/80 (65%)
Frame = +3
Query: 294 LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEM 353
LFV+GLS T+ + LR F FGE+V +V+ D++S SKG+ F++Y T E A ++ M
Sbjct: 207 LFVSGLSKRTNTERLREEFAKFGEVVHARVVTDRVSGYSKGFGFVQYATIEDAAKGIEGM 386
Query: 354 NGKIINGWMIVVDVAKTNPP 373
+GK ++GW+I + A+ PP
Sbjct: 387 DGKFLDGWVIFAEYARPRPP 446
>TC217631 similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldrich syndrome protein
interacting protein (WASP interacting protein) (PRPL-2
protein), partial (7%)
Length = 1440
Score = 74.7 bits (182), Expect = 6e-14
Identities = 38/86 (44%), Positives = 55/86 (63%)
Frame = +1
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+V+M+KP G ++ +ID Y+KTL V+GSE+EA+M +Y S +F F + EE+
Sbjct: 364 HWLVVMEKPE-GDPTRDDIIDSYIKTLAKVIGSEEEARMKIYSVSTRHYFAFGALVSEEL 540
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S ++ LPGV V PD N +KDY
Sbjct: 541 SYKIKELPGVRWVLPDSYLNVKEKDY 618
Score = 62.4 bits (150), Expect = 3e-10
Identities = 34/98 (34%), Positives = 56/98 (56%)
Frame = +1
Query: 174 RTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKT 233
+ +L + +HWLV M+KP + T+ I+D Y + L KV+G+E++A+M IY VS +
Sbjct: 328 KETILLDGCDFEHWLVVMEKPEGDP-TRDDIIDSYIKTLAKVIGSEEEARMKIYSVSTRH 504
Query: 234 NFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
F F + E+ + + +PGV V PD + K+Y
Sbjct: 505 YFAFGALVSEELSYKIKELPGVRWVLPDSYLNVKEKDY 618
>TC203450 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (53%)
Length = 1196
Score = 74.7 bits (182), Expect = 6e-14
Identities = 50/150 (33%), Positives = 75/150 (49%), Gaps = 14/150 (9%)
Frame = +3
Query: 23 TNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTTHNR---------- 72
TN+ LP PS + + +CNR I AA + + S+++N
Sbjct: 156 TNLRLP--------PSIVTRTRNCNR----IRAALDGDFSAKRSSSSNNNDQRETIMLPG 299
Query: 73 ----HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDI 128
HW+++M+ P ++ Q+ID Y+ TL TV+GS +EA+ +Y S T+ GF C +
Sbjct: 300 CDYNHWLIVMEFPKDPAPTREQMIDTYLDTLATVLGSMEEAKKNMYAFSTTTYTGFQCTV 479
Query: 129 DEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
DE S + LPGVL V PD + KDY
Sbjct: 480 DEATSEKFKGLPGVLWVLPDSYIDVKNKDY 569
Score = 70.5 bits (171), Expect = 1e-12
Identities = 36/115 (31%), Positives = 61/115 (52%)
Frame = +3
Query: 157 DYSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVM 216
D S+ ++ S+ R ++ P + HWL+ M+ P T+ Q++D Y L V+
Sbjct: 225 DGDFSAKRSSSSNNNDQRETIMLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLDTLATVL 404
Query: 217 GNEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
G+ ++A+ +Y S T GF C +DE ++ G+PGV V PD + +NK+Y
Sbjct: 405 GSMEEAKKNMYAFSTTTYTGFQCTVDEATSEKFKGLPGVLWVLPDSYIDVKNKDY 569
>TC203462 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (53%)
Length = 708
Score = 72.8 bits (177), Expect = 2e-13
Identities = 38/93 (40%), Positives = 55/93 (58%)
Frame = +1
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW+++M+ P ++ Q+ID Y+ TL TV+GS +EA+ +Y S T+ GF
Sbjct: 271 PGCDYN-HWLIVMEFPKDPAPTREQMIDTYLDTLATVLGSMEEAKKNMYAFSTTTYTGFQ 447
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
C +DE S + LPGVL V PD + KDY
Sbjct: 448 CTVDEATSEKFKGLPGVLWVLPDSYIDVKNKDY 546
Score = 68.9 bits (167), Expect = 4e-12
Identities = 34/104 (32%), Positives = 57/104 (54%)
Frame = +1
Query: 168 SSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIY 227
S+ + R ++ P + HWL+ M+ P T+ Q++D Y L V+G+ ++A+ +Y
Sbjct: 235 SNNNEQRETIMLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLDTLATVLGSMEEAKKNMY 414
Query: 228 HVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
S T GF C +DE ++ G+PGV V PD + +NK+Y
Sbjct: 415 AFSTTTYTGFQCTVDEATSEKFKGLPGVLWVLPDSYIDVKNKDY 546
>TC221248 weakly similar to GB|CAD40580.1|21741678|OSJN00141 oj000126_13.2
{Oryza sativa (japonica cultivar-group);} , partial
(40%)
Length = 861
Score = 72.0 bits (175), Expect = 4e-13
Identities = 41/99 (41%), Positives = 62/99 (62%), Gaps = 2/99 (2%)
Frame = +1
Query: 275 LSVPNSSEASQEASLRT--KKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRS 332
L P++S SQ LRT K+FV GL+F T+E+ L AF +G +++ +I++K RS
Sbjct: 328 LKKPHASP-SQLLFLRTMTSKVFVKGLAFSTTEEELAKAFSQYGSVLKANIILNKAKNRS 504
Query: 333 KGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTN 371
KG+ ++ + EE A A +MNGKI++G +I VDV N
Sbjct: 505 KGFGYVTFAKEEEACKAQIDMNGKILHGRVIYVDVQLPN 621
>TC215891 similar to GB|AAM78058.1|21928059|AY125548 AT5g61030/maf19_30
{Arabidopsis thaliana;} , partial (41%)
Length = 782
Score = 72.0 bits (175), Expect = 4e-13
Identities = 34/92 (36%), Positives = 61/92 (65%)
Frame = +2
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
+S + KLF+ G+S+ T E++LR AF +GE+V ++I+D+ + RS+G+ FI YT+ E A
Sbjct: 158 SSAPSTKLFIGGVSYSTDEQSLREAFSKYGEVVDARIIMDRETGRSRGFGFITYTSVEEA 337
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPRFNRG 378
+A++ ++G+ ++G I V+ A P + G
Sbjct: 338 SSAIQALDGQDLHGRPIRVNYANERPRGYGGG 433
>AW597392 weakly similar to GP|7630030|emb| RNA binding protein-like
{Arabidopsis thaliana}, partial (78%)
Length = 392
Score = 71.6 bits (174), Expect = 5e-13
Identities = 35/81 (43%), Positives = 52/81 (63%)
Frame = +2
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV LSFYT+++ L+ F FG + Q + +D I+KR KG+ F+ + +E A A K
Sbjct: 80 KLFVHRLSFYTTQEQLKKLFSPFGLVTQADLALDPITKRPKGFGFVSFKSEIEAEKACKA 259
Query: 353 MNGKIINGWMIVVDVAKTNPP 373
MNG+I+NG +I+V+ A P
Sbjct: 260 MNGRIVNGRLILVEPANEK*P 322
>TC223147 weakly similar to UP|Q9LZT1 (Q9LZT1) RNA binding protein-like
(At3g46020), partial (73%)
Length = 622
Score = 70.5 bits (171), Expect = 1e-12
Identities = 33/79 (41%), Positives = 51/79 (63%)
Frame = +3
Query: 295 FVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMN 354
F +GLSFYT+++ L+ F FG + Q + +D I+KR KG+ F+ + +E A A K MN
Sbjct: 132 FFSGLSFYTTQEQLKKLFSPFGLVTQADLALDPITKRPKGFGFVSFKSEIEAEKACKAMN 311
Query: 355 GKIINGWMIVVDVAKTNPP 373
G+I+NG +I+V+ A P
Sbjct: 312 GRIVNGRLILVEPANEK*P 368
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.130 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,986,341
Number of Sequences: 63676
Number of extensions: 263250
Number of successful extensions: 2045
Number of sequences better than 10.0: 200
Number of HSP's better than 10.0 without gapping: 1959
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2034
length of query: 384
length of database: 12,639,632
effective HSP length: 99
effective length of query: 285
effective length of database: 6,335,708
effective search space: 1805676780
effective search space used: 1805676780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0095c.6


