

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094b.3
(337 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
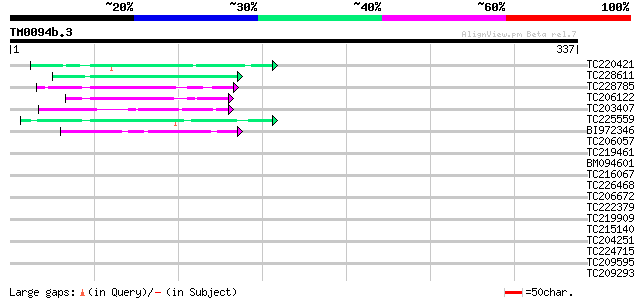
Score E
Sequences producing significant alignments: (bits) Value
TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell... 53 2e-07
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 47 9e-06
TC228785 weakly similar to GB|AAP42749.1|30984572|BT008736 At3g5... 45 6e-05
TC206122 weakly similar to PIR|C86473|C86473 arabinogalactan-pro... 44 8e-05
TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pr... 43 2e-04
TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 43 2e-04
BI972346 41 9e-04
TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin, d... 40 0.001
TC219461 homologue to UP|Q9AR19 (Q9AR19) Histone acetyltransfera... 40 0.002
BM094601 40 0.002
TC216067 weakly similar to UP|RH2A_ARATH (Q9ZT50) RHA2a protein ... 39 0.004
TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich prote... 39 0.004
TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyra... 39 0.004
TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, par... 38 0.006
TC219909 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-... 38 0.006
TC215140 similar to GB|AAL32703.1|17065098|AY062625 nucleotide s... 38 0.007
TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Sy... 37 0.010
TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 37 0.010
TC209595 homologue to GB|AAL62011.1|18252261|AY072620 AT5g08330/... 37 0.013
TC209293 homologue to UP|RN12_HUMAN (Q9NVW2) RING finger protein... 37 0.013
>TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell wall
protein gp1 precursor (Hydroxyproline-rich glycoprotein
1), partial (13%)
Length = 1409
Score = 53.1 bits (126), Expect = 2e-07
Identities = 49/151 (32%), Positives = 58/151 (37%), Gaps = 4/151 (2%)
Frame = +3
Query: 13 KRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACD----PLASDHPSE 68
K+Q S P R+ P P T P + T P P T S P S PS
Sbjct: 186 KQQPLNPSTSPPHRTPAPSP-PSTMPPSAT-----PPSPSATSSPSASPTAPPTSPSPSP 347
Query: 69 VNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPT 128
++ P AP P + A S S P PSP SA NSP G S P+
Sbjct: 348 PSSPSPPSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASA-NSPASGSPTSRTSPTSPS 524
Query: 129 GPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
S+S PP P S S P K +PS P
Sbjct: 525 PTSTSKPPAPSS---SSPA*PSSKPSPSPTP 608
Score = 33.5 bits (75), Expect = 0.14
Identities = 39/128 (30%), Positives = 51/128 (39%), Gaps = 2/128 (1%)
Frame = +3
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P+ T + PL +PS +TP AP SP S PP +P PS
Sbjct: 162 PHPLWTQNKQQPL---NPSTSPPHRTP-AP----------SPPSTMPPSATPPS----PS 287
Query: 109 APNSPGVG--GEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
A +SP + S + P+ PS S P P S S P +PS +P A A
Sbjct: 288 ATSSPSASPTAPPTSPSPSPPSSPSPPSAPSPSSTAPSPPSASP-PTSPSPSPPSPASTA 464
Query: 167 VLNLLRAG 174
N +G
Sbjct: 465 SANSPASG 488
Score = 28.1 bits (61), Expect = 5.9
Identities = 23/82 (28%), Positives = 31/82 (37%), Gaps = 3/82 (3%)
Frame = +3
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASD---HPSEVNASQ 73
SP S S G P +T+P +P P T P +S S+ + S
Sbjct: 447 SPASTASANSPASGSPTSRTSP--------TSPSPTSTSKPPAPSSSSPA*PSSKPSPSP 602
Query: 74 TPDAPRPVTLAEGQRSPSSIRP 95
TP +P P S +SI P
Sbjct: 603 TPISPDPSPATSTPTSHTSISP 668
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 47.4 bits (111), Expect = 9e-06
Identities = 35/114 (30%), Positives = 44/114 (37%), Gaps = 1/114 (0%)
Frame = +3
Query: 26 RSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAE 85
RS P P + + T +P V S+ P S PS S +P P T +
Sbjct: 162 RSPAPPPPPPRSASSSTTAS--SPTAVTPPSSSPPRPSPGPSATPTSTSPSPATPSTSSS 335
Query: 86 GQRSPSSIRPPIPSPVKV-TLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
R P S P PSP + P A +SP S+A T S PPYP
Sbjct: 336 APRPPLSSPAPTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYP 497
>TC228785 weakly similar to GB|AAP42749.1|30984572|BT008736 At3g52060
{Arabidopsis thaliana;} , partial (65%)
Length = 860
Score = 44.7 bits (104), Expect = 6e-05
Identities = 39/121 (32%), Positives = 50/121 (41%), Gaps = 1/121 (0%)
Frame = +1
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL-ASDHPSEVNASQTP 75
+P+S SP P P T T T P P T A PL +S + S NA P
Sbjct: 79 TPIST-SPPYGTSSSPTPPPTFSTSTS----TPTPPSTSPALCPLFSSTNSSPPNAPSAP 243
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
P A PSS PP P+ PS+P++ + + + PT PSSS P
Sbjct: 244 PLP*SPPPAASSPPPSSTTPPTPT------SPSSPSTASL-----STPSPTPTSPSSSPP 390
Query: 136 P 136
P
Sbjct: 391 P 393
>TC206122 weakly similar to PIR|C86473|C86473 arabinogalactan-protein -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(40%)
Length = 797
Score = 44.3 bits (103), Expect = 8e-05
Identities = 30/100 (30%), Positives = 42/100 (42%)
Frame = +3
Query: 34 PQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSI 93
P TTP T AP P T + P A+ P+ + P +P P T P++
Sbjct: 153 PSTTPSPSTS----APAPSPTANTPPPAAAPAPATPTPTPAPSSPTPTTAPSQAPGPTTS 320
Query: 94 RPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
PP P GPS P +PG G + A++P + S
Sbjct: 321 SPPAP-------GPSGP-APGPSGPAPDQPASEPPSAAFS 416
Score = 33.9 bits (76), Expect = 0.11
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Frame = +3
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS-APNSPGVGGEGENLSAAQPTGPSSSSPP 136
P P T A ++ PP +P T P+ AP+SP + +Q GP++SSPP
Sbjct: 165 PSPSTSAPAPSPTANTPPPAAAPAPATPTPTPAPSSPT-----PTTAPSQAPGPTTSSPP 329
Query: 137 YP 138
P
Sbjct: 330 AP 335
>TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(AtAGP4) (AT5g10430/F12B17_220), partial (50%)
Length = 966
Score = 42.7 bits (99), Expect = 2e-04
Identities = 36/116 (31%), Positives = 48/116 (41%)
Frame = +2
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P SA +P + P P T P T P P A+ TP A
Sbjct: 248 PRSAPAPAPTTPATPPPATPPPAATPPPAATPPPA------------------ATPTP-A 370
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
P P T A S + PP PSP VT P++PN+P G + +A+P PS++
Sbjct: 371 PAPPTAAPTPASSPAASPPSPSPT-VTPSPTSPNTPPGPSPGPS-GSAEPPPPSAA 532
Score = 33.1 bits (74), Expect = 0.18
Identities = 29/96 (30%), Positives = 38/96 (39%), Gaps = 5/96 (5%)
Frame = +2
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
AP P T A P A+ P A+ P A P A +P+ PP +P +
Sbjct: 257 APAPAPTTPATPPPATPPP----AATPPPAATPPPAATPTPAPA---PPTAAPTPASSPA 415
Query: 108 SAPNSPGVGGEGENLSAAQPTGPS-----SSSPPYP 138
++P SP S P GPS S+ PP P
Sbjct: 416 ASPPSPSPTVTPSPTSPNTPPGPSPGPSGSAEPPPP 523
>TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (47%)
Length = 870
Score = 42.7 bits (99), Expect = 2e-04
Identities = 46/157 (29%), Positives = 63/157 (39%), Gaps = 4/157 (2%)
Frame = +3
Query: 7 VAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHP 66
VA VGG+ SP +A S + P P T +P PV + + P A+
Sbjct: 117 VAGVGGQ---SPAAAPSNTPATPAAATPAQAPSTVPK----SPAPVASPKSSPPAATPAS 275
Query: 67 SEV-NASQTPDAPRPVTLAEGQRSPSSIRPPI---PSPVKVTLGPSAPNSPGVGGEGENL 122
+ + + TP AP PVT P + PP P+PV V+ S P V +
Sbjct: 276 TPAASPTTTPAAPAPVTKPPAASPPPATPPPATPPPAPVPVS---SPPAPVPVSSPPAPV 446
Query: 123 SAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
A PT P + SP H K GAP+ +P
Sbjct: 447 PVAAPTTPVAPSP------APKHKKKGKKHGAPAPSP 539
>BI972346
Length = 421
Score = 40.8 bits (94), Expect = 9e-04
Identities = 34/109 (31%), Positives = 48/109 (43%), Gaps = 1/109 (0%)
Frame = +2
Query: 31 EPDPQT-TPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRS 89
EPD + TP T T +P SA D AS S A+++P P L +
Sbjct: 44 EPDQASLTPSTSTSNPARSPASATLASAPDLTASTPSS---ATRSPSTALP--LRRRSPT 208
Query: 90 PSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
P+S+RPP P P+ + S ++ G ++ P PSSS P P
Sbjct: 209 PTSLRPPPPPPIPASRSSSLGSTARAARSGR---SSWPPSPSSSFPSIP 346
>TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin,
delta-subunit, complete
Length = 1930
Score = 40.0 bits (92), Expect = 0.001
Identities = 45/154 (29%), Positives = 55/154 (35%), Gaps = 8/154 (5%)
Frame = +3
Query: 14 RQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
R SSP CS P P T PL + +AP +S S AS
Sbjct: 306 RSSSPPPRCSSSSPSPRTPPPATAPLPSSSS--LAPSS----------SSASSSSPTAST 449
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPIPSPVK---VTLGPSAPNSPGVGGEGENL---SAAQP 127
P +P P T + PS+ PP PSP T S P P N S P
Sbjct: 450 PPSSPTPSTRLP*R--PSTFSPPWPSPSSSPTATPS*SPPAHPSTARSSANTPRSSLPSP 623
Query: 128 TGPSSSS--PPYPFSFLLSHPYLEKLKGAPSQNP 159
+ P S S PP P + + + APS P
Sbjct: 624 STPFSPSWMPPSPTWSISAMSRS*RNSAAPSTTP 725
>TC219461 homologue to UP|Q9AR19 (Q9AR19) Histone acetyltransferase GCN5
(At3g54610) (Expressed protein), partial (12%)
Length = 853
Score = 39.7 bits (91), Expect = 0.002
Identities = 34/129 (26%), Positives = 52/129 (39%), Gaps = 6/129 (4%)
Frame = +2
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
S+ S+C P+ +++ P P + P T P P T S P A+ P+ +TP
Sbjct: 206 STSASSC-PRTTRLPSPPPPSPPTPAT-----VPSPPTTTSRASPPAAPTPTPTPNPRTP 367
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP------SAPNSPGVGGEGENLSAAQPTG 129
+ T + + P P P TL P S P +P + L+A +
Sbjct: 368 SSTTTRTTTTTTTTTTPPCAPSPPPGSATLPPLPATPSSRPKTP--PSKLRILTAPKTPE 541
Query: 130 PSSSSPPYP 138
PS +PP P
Sbjct: 542 PSDLAPPPP 568
>BM094601
Length = 429
Score = 39.7 bits (91), Expect = 0.002
Identities = 31/102 (30%), Positives = 45/102 (43%), Gaps = 5/102 (4%)
Frame = +1
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
P + S +AS P P ++ ++ S SS+ P+ SP ++ PS SP
Sbjct: 76 PPSPSSASGGSASTVPPQPPSLSASQPLTSLSSLLSPLSSPTTLSSPPSLSPSPHPSSLS 255
Query: 120 ENLSAAQ-----PTGPSSSSPPYPFSFLLSHPYLEKLKGAPS 156
+ S+ PT S SPP+PF LS PY AP+
Sbjct: 256 PHTSSRSSSLPFPTSSRSLSPPHPF---LSPPYSLSPPPAPA 372
>TC216067 weakly similar to UP|RH2A_ARATH (Q9ZT50) RHA2a protein (RING-H2
zinc finger protein), partial (37%)
Length = 733
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/78 (30%), Positives = 35/78 (44%), Gaps = 1/78 (1%)
Frame = +1
Query: 36 TTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSS-IR 94
T + T DP D +C P++S PS N+S AP P T +P+S R
Sbjct: 208 TPTRSSTSASSPPSDPASPDLSCSPISS--PSAANSSTITTAPPPATTTTTTTTPASCAR 381
Query: 95 PPIPSPVKVTLGPSAPNS 112
PP + + P+A +S
Sbjct: 382 PPSRTATRSACSPAATSS 435
>TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich protein, partial
(20%)
Length = 1041
Score = 38.5 bits (88), Expect = 0.004
Identities = 38/162 (23%), Positives = 54/162 (32%), Gaps = 7/162 (4%)
Frame = -2
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
++ P S+P++ P P T P T VAP P + +
Sbjct: 947 TLPPTTPTATSPSSSTPLAVTQPPTVVASPPTTSTPPTTSQPPANVAPKPAPVKPSAPKV 768
Query: 62 ASDHPSEVNASQTP----DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGG 117
A + Q P P PV+ P + PP+P P P SP
Sbjct: 767 APASSPKAPPPQIPIPQPPKPSPVSPPPLPPPPVASPPPLPPPKIAPTPAKTPPSP-APA 591
Query: 118 EGENLSAAQPTGPS---SSSPPYPFSFLLSHPYLEKLKGAPS 156
+ A PT P+ S PP P + P +E AP+
Sbjct: 590 KATPAPAPAPTKPAPTPSPVPPPPAATPAPAPVIEVPAPAPA 465
>TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyrate
dehydrogenase , partial (76%)
Length = 1230
Score = 38.5 bits (88), Expect = 0.004
Identities = 34/127 (26%), Positives = 42/127 (32%)
Frame = +2
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
G ++S S C+P S P P +T P P T P
Sbjct: 98 GSARASWASPCAPTSSVPATPSPSSTAPPPR------PSPSSTSGPTSPPPPTPSQPAPT 259
Query: 72 SQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS 131
S +P + P T A+ +P PP PSP PSAP P PS
Sbjct: 260 SSSPSSATPRTSAQSSSTP----PPAPSP------PSAP-----AASSST*PPPNPPSPS 394
Query: 132 SSSPPYP 138
S P P
Sbjct: 395 KSRTPLP 415
>TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, partial (84%)
Length = 950
Score = 38.1 bits (87), Expect = 0.006
Identities = 31/83 (37%), Positives = 40/83 (47%), Gaps = 4/83 (4%)
Frame = +2
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIP---SPVKVTLGPSAPNSPGVG 116
PL PS + S P P V+ A S S PP SPV VT PS+ +P V
Sbjct: 86 PLPGTRPS-ASGSSPPARPPAVSSATTTTSFPSASPPTTVRSSPVPVT-APSSCGTPSVT 259
Query: 117 GEGENLSAAQPTG-PSSSSPPYP 138
+ + A P+G P+S+SPP P
Sbjct: 260 ASTPSPTRATPSGSPASASPPTP 328
>TC219909 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (21%)
Length = 943
Score = 38.1 bits (87), Expect = 0.006
Identities = 35/128 (27%), Positives = 45/128 (34%), Gaps = 4/128 (3%)
Frame = +1
Query: 13 KRQSSPVSACSPKRSK--VGEPDPQTTPLTGTGGGFVA--PDPVVTDSACDPLASDHPSE 68
K SP + SPK + V P P +P + A P P V A P S
Sbjct: 247 KTSPSPSPSQSPKANPPTVSPPLPSPSPSVHSKSLSPASSPVPAVGTPAISPAISIPTLA 426
Query: 69 VNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPT 128
P + P + + PSS PP P P + P+ P SP N +A P
Sbjct: 427 PETGTPPPSLGPSSSSPPSPGPSSSSPPSPGPSSSSPPPAGPTSPPSLAPSSNSTAPSPK 606
Query: 129 GPSSSSPP 136
S P
Sbjct: 607 NSGFSVAP 630
Score = 35.0 bits (79), Expect = 0.048
Identities = 36/107 (33%), Positives = 44/107 (40%)
Frame = +1
Query: 32 PDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPS 91
P P+T+P A P V+ PL S PS + S +P A PV
Sbjct: 238 PFPKTSPSPSPSQSPKANPPTVSP----PLPSPSPSVHSKSLSP-ASSPVPAV----GTP 390
Query: 92 SIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+I P I P TL P P G + S+ GPSSSSPP P
Sbjct: 391 AISPAISIP---TLAPETGTPPP--SLGPSSSSPPSPGPSSSSPPSP 516
>TC215140 similar to GB|AAL32703.1|17065098|AY062625 nucleotide sugar
epimerase-like protein {Arabidopsis thaliana;} , partial
(91%)
Length = 1885
Score = 37.7 bits (86), Expect = 0.007
Identities = 36/123 (29%), Positives = 49/123 (39%), Gaps = 2/123 (1%)
Frame = +1
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
++PV CS TTP ++P P + S LAS P +TP
Sbjct: 478 NAPVRQCSGNTRFSSWKVTLTTPPCLRSSSTLSPSPTSSTSPLR-LASATPC-----RTP 639
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENL--SAAQPTGPSSS 133
+ P TL S PPIP+P L P +P + L S+ PT P +S
Sbjct: 640 NPTSPPTLLASSTSSRLPNPPIPNPP*SGLHP----APSTASTPKTLSPSSTAPTNPPAS 807
Query: 134 SPP 136
+PP
Sbjct: 808 TPP 816
>TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Syndrome
protein homolog, partial (8%)
Length = 1045
Score = 37.4 bits (85), Expect = 0.010
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 2/71 (2%)
Frame = +3
Query: 70 NASQTPDAPRPVTLAEGQRSPSS-IRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAA-QP 127
+ + TP AP A SPS+ + P PSP +L P+ SPG AA +P
Sbjct: 192 STAPTPTAPTFPPSAPPSTSPSTPLTAPSPSPSPPSLPPAPAGSPGASTSSAMACAAPKP 371
Query: 128 TGPSSSSPPYP 138
+ PS PP P
Sbjct: 372 SSPSPPPPPPP 404
>TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1270
Score = 37.4 bits (85), Expect = 0.010
Identities = 25/93 (26%), Positives = 36/93 (37%), Gaps = 12/93 (12%)
Frame = +2
Query: 56 SACDPLASDHPSEVNASQTPDAPR------------PVTLAEGQRSPSSIRPPIPSPVKV 103
S+ P +S P++ AS T +P P + SPS P P+
Sbjct: 578 SSSQPSSSCSPAQAPASPTTSSPTTALPPLPASSPPPSPMHSPSSSPSPSAPTSPAATST 757
Query: 104 TLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
PSAP+S S + P+ + SSPP
Sbjct: 758 PPSPSAPSSAATSPSSAASSTSSPSSSAPSSPP 856
>TC209595 homologue to GB|AAL62011.1|18252261|AY072620 AT5g08330/F8L15_60
{Arabidopsis thaliana;} , partial (56%)
Length = 869
Score = 37.0 bits (84), Expect = 0.013
Identities = 35/114 (30%), Positives = 48/114 (41%), Gaps = 9/114 (7%)
Frame = +2
Query: 32 PDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPS 91
P P T T VA S+ P +S S +++ +P A RP + + +R+P
Sbjct: 80 PSRNRLPRTATARSTVAGAAFACRSSAPPASSS--SRASSATSPTA-RPSSGSSARRNPP 250
Query: 92 SIRPPIP------SPVKVTLGPSA-PNSPGVGGEGENLSAAQP--TGPSSSSPP 136
S P P SP+ T PS+ P P L+ P T PS SSPP
Sbjct: 251 SSPLPAPAPLRRASPLSPTTSPSSLPLPPSSSANAFALTTTPPPKTTPSRSSPP 412
>TC209293 homologue to UP|RN12_HUMAN (Q9NVW2) RING finger protein 12 (LIM
domain interacting RING finger protein) (RING finger LIM
domain-binding protein) (R-LIM) (NY-REN-43 antigen),
partial (4%)
Length = 957
Score = 37.0 bits (84), Expect = 0.013
Identities = 33/119 (27%), Positives = 50/119 (41%), Gaps = 13/119 (10%)
Frame = -3
Query: 30 GEP--DPQTTPLTG--TGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAE 85
G+P DP TG +G GF SA +S HPS ++S +P + ++
Sbjct: 448 GDPKLDPSEGETTGGESGQGFSRLSSAARASAIASQSSKHPSSSSSSSSPSSSSSTSVTF 269
Query: 86 GQRSPSSIR---------PPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
++ P + PP+ PVK N+P G++ S + PSSSSP
Sbjct: 268 PEQHPPLLELLLLLLLLLPPLLLPVKFVPESVPHNTPAGQLVGDSESLS*WINPSSSSP 92
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,291,831
Number of Sequences: 63676
Number of extensions: 210197
Number of successful extensions: 2176
Number of sequences better than 10.0: 294
Number of HSP's better than 10.0 without gapping: 1867
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2049
length of query: 337
length of database: 12,639,632
effective HSP length: 98
effective length of query: 239
effective length of database: 6,399,384
effective search space: 1529452776
effective search space used: 1529452776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0094b.3


