

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094a.6
(990 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
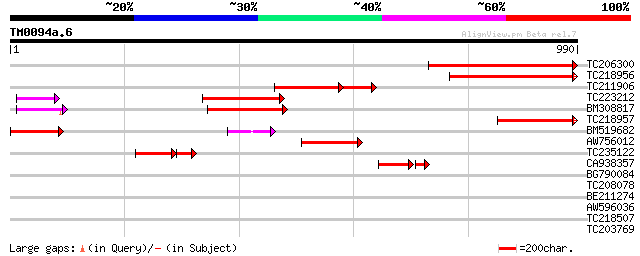
Score E
Sequences producing significant alignments: (bits) Value
TC206300 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 462 e-130
TC218956 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 373 e-103
TC211906 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 196 1e-72
TC223212 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 263 2e-70
BM308817 similar to GP|13111324|db 110 kDa 4SNc-Tudor domain pro... 252 5e-67
TC218957 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 233 4e-61
BM519682 similar to GP|10176871|dbj transcription factor-like pr... 177 3e-44
AW756012 similar to GP|13111324|db 110 kDa 4SNc-Tudor domain pro... 173 4e-43
TC235122 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain... 119 3e-42
CA938357 similar to GP|18086332|gb| AT5g07350/T2I1_60 {Arabidops... 115 6e-34
BG790084 similar to GP|24421095|dbj receptor-like protein kinase... 35 0.21
TC208078 31 3.1
BE211274 similar to PIR|T06699|T0 zinc finger protein T29H11.50 ... 30 5.3
AW596036 30 5.3
TC218507 similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like protei... 30 5.3
TC203769 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate ... 30 6.9
>TC206300 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (26%)
Length = 1165
Score = 462 bits (1189), Expect = e-130
Identities = 226/260 (86%), Positives = 245/260 (93%)
Frame = +1
Query: 731 QQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCY 790
QQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLA+LNLK+APVLGAF+PKKGDIVLCY
Sbjct: 1 QQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLASLNLKDAPVLGAFNPKKGDIVLCY 180
Query: 791 FHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQ 850
FHADKSWYRAMVVNTPRGPVESP D+FEVFY+DYGNQE V YSQLRP+D SVSAAPGLAQ
Sbjct: 181 FHADKSWYRAMVVNTPRGPVESPNDLFEVFYVDYGNQEVVPYSQLRPVDPSVSAAPGLAQ 360
Query: 851 LCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAV 910
LCSLAYIK P+LEEDFGQEAAEYLSELTL+SGKEFRA+VEE+DTSGGKVKGQGTG ILAV
Sbjct: 361 LCSLAYIKIPNLEEDFGQEAAEYLSELTLNSGKEFRAKVEEKDTSGGKVKGQGTGAILAV 540
Query: 911 TLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGD 970
TLVAVDAEISVNAAMLQEGLAR EKRNRWDRK+R+ LD+L+ FQD+A+ RRGMWQYGD
Sbjct: 541 TLVAVDAEISVNAAMLQEGLARTEKRNRWDRKDRQTALDNLENFQDEAKTSRRGMWQYGD 720
Query: 971 VESDDEDTAPPARKAGAGRR 990
++SDDEDTAPP RK G GR+
Sbjct: 721 IQSDDEDTAPPPRKTGGGRK 780
>TC218956 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (22%)
Length = 889
Score = 373 bits (957), Expect = e-103
Identities = 182/223 (81%), Positives = 204/223 (90%)
Frame = +3
Query: 768 LNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQ 827
LNL+EAP+LGAF+PKKGD VLC F ADKSWYRAMVVN PRGPVESP D+FEVFYIDYGNQ
Sbjct: 9 LNLQEAPLLGAFNPKKGDTVLCLFGADKSWYRAMVVNGPRGPVESPNDMFEVFYIDYGNQ 188
Query: 828 EQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRA 887
E+V YSQLRP+D SVSAAPG+AQLCSLAY+K P+LEEDFG+EAAEYLSELTL+SGKEFRA
Sbjct: 189 EEVPYSQLRPIDPSVSAAPGIAQLCSLAYVKVPNLEEDFGEEAAEYLSELTLNSGKEFRA 368
Query: 888 QVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVG 947
+VEERDTSGGK KGQGTG +LAVTLVAVD++ISVNAAMLQEGLAR+EKRNRWDRKER+
Sbjct: 369 KVEERDTSGGKAKGQGTGPVLAVTLVAVDSDISVNAAMLQEGLARLEKRNRWDRKERQQA 548
Query: 948 LDSLQKFQDDARKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
LD+L FQ +AR R GMWQYGD++SDDEDTAPPARKAG GR+
Sbjct: 549 LDNLDPFQGEARTNRCGMWQYGDIQSDDEDTAPPARKAG-GRK 674
>TC211906 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (15%)
Length = 536
Score = 196 bits (498), Expect(2) = 1e-72
Identities = 99/120 (82%), Positives = 104/120 (86%)
Frame = +1
Query: 463 TDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFG 522
TDGS VPS AADS VMDFGSVFLLS K D+DD PSS P AGSQ GVNV EL+VGRGFG
Sbjct: 1 TDGSVVPS-AADS*VMDFGSVFLLSGAKVDNDDAPSSAPPAGSQQNGVNVAELIVGRGFG 177
Query: 523 TVIRHRDFEERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPF 582
TVIRHRDFEERSNYYD+LL AESRA+SGRKG HSAKDPPVMHITDLT SAKKA+DFLPF
Sbjct: 178 TVIRHRDFEERSNYYDSLLAAESRAISGRKGTHSAKDPPVMHITDLTMASAKKARDFLPF 357
Score = 97.1 bits (240), Expect(2) = 1e-72
Identities = 49/68 (72%), Positives = 54/68 (79%), Gaps = 1/68 (1%)
Frame = +2
Query: 574 KKAKDFLPFLHRS-RRIPAVVEYVLSGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEE 632
+K + FLH + AVVEYVL GHRFKLLIPKETCSIAF+ SGVRCPGR EPYS+E
Sbjct: 332 RKPETSCXFLHXK*KEFRAVVEYVLRGHRFKLLIPKETCSIAFSFSGVRCPGRDEPYSDE 511
Query: 633 AIALMRRK 640
AIALMRRK
Sbjct: 512 AIALMRRK 535
>TC223212 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (13%)
Length = 431
Score = 263 bits (673), Expect = 2e-70
Identities = 127/143 (88%), Positives = 137/143 (94%)
Frame = +3
Query: 337 NMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVA 396
NMMEEEAKR+LKTAEL+AKK RLRMWTNYVPP SNSKAIHNQNF+GKVVEVVSGDCI+VA
Sbjct: 3 NMMEEEAKRKLKTAELQAKKDRLRMWTNYVPPPSNSKAIHNQNFSGKVVEVVSGDCIVVA 182
Query: 397 DDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEY 456
DDSIPYGSPLAERRVNLSSIRCPK+GNPRRDEKPAPYAREAKEFLRT L+GRQV+V+MEY
Sbjct: 183 DDSIPYGSPLAERRVNLSSIRCPKMGNPRRDEKPAPYAREAKEFLRTHLIGRQVNVQMEY 362
Query: 457 SRKIVPTDGSAVPSPAADSRVMD 479
SRK+ PTDGS VPS A+DSRVMD
Sbjct: 363 SRKVSPTDGSVVPSTASDSRVMD 431
Score = 52.0 bits (123), Expect = 1e-06
Identities = 26/78 (33%), Positives = 47/78 (59%), Gaps = 2/78 (2%)
Frame = +3
Query: 12 YRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLA--RRGGVDEPFAWESRE 69
+ G+V V SGDC+V+ + PL E+ + L+S+ P++ RR P+A E++E
Sbjct: 132 FSGKVVEVVSGDCIVVADDSIPYGSPLAERRVNLSSIRCPKMGNPRRDEKPAPYAREAKE 311
Query: 70 YLRKLCIGKEVTFRVDYN 87
+LR IG++V +++Y+
Sbjct: 312 FLRTHLIGRQVNVQMEYS 365
>BM308817 similar to GP|13111324|db 110 kDa 4SNc-Tudor domain protein {Pisum
sativum}, partial (19%)
Length = 428
Score = 252 bits (644), Expect = 5e-67
Identities = 121/141 (85%), Positives = 136/141 (95%)
Frame = +2
Query: 345 RRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGS 404
R+LKT+EL+AKK+RL++WTNYVPPA+NSKAIH+QNFTGKVVEVVSGDCIIVADD IPYGS
Sbjct: 2 RKLKTSELQAKKNRLKIWTNYVPPATNSKAIHDQNFTGKVVEVVSGDCIIVADDLIPYGS 181
Query: 405 PLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTD 464
PLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRL+GRQV+V+MEYSRK+ P D
Sbjct: 182 PLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLIGRQVNVQMEYSRKVGPAD 361
Query: 465 GSAVPSPAADSRVMDFGSVFL 485
GSAVPS A+++R MDFGSVFL
Sbjct: 362 GSAVPSGASEARAMDFGSVFL 424
Score = 51.2 bits (121), Expect = 2e-06
Identities = 34/106 (32%), Positives = 56/106 (52%), Gaps = 17/106 (16%)
Frame = +2
Query: 12 YRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLA--RRGGVDEPFAWESRE 69
+ G+V V SGDC+++ PL E+ + L+S+ P++ RR P+A E++E
Sbjct: 107 FTGKVVEVVSGDCIIVADDLIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKE 286
Query: 70 YLRKLCIGKEVTFRVDY-------------NVASINR--DFGTVFL 100
+LR IG++V +++Y + AS R DFG+VFL
Sbjct: 287 FLRTRLIGRQVNVQMEYSRKVGPADGSAVPSGASEARAMDFGSVFL 424
Score = 29.6 bits (65), Expect = 6.9
Identities = 17/54 (31%), Positives = 27/54 (49%), Gaps = 8/54 (14%)
Frame = +2
Query: 617 LSGVRCPGRGE--------PYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGS 662
LS +RCP G PY+ EA +R +++ R V +++E + G GS
Sbjct: 206 LSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLIGRQVNVQMEYSRKVGPADGS 367
>TC218957 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (14%)
Length = 629
Score = 233 bits (593), Expect = 4e-61
Identities = 115/139 (82%), Positives = 128/139 (91%)
Frame = +2
Query: 852 CSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVT 911
CSLAY+K P+LEEDFGQEAAEYLSELTL+SGKEFRA+VEERDTSGGK KGQGTGT+LAVT
Sbjct: 2 CSLAYVKVPNLEEDFGQEAAEYLSELTLNSGKEFRAKVEERDTSGGKAKGQGTGTVLAVT 181
Query: 912 LVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDV 971
LVAVD+EISVNAAMLQEGLAR+EKRNRWD KER+ LD+L FQ +AR RRGMWQYGD+
Sbjct: 182 LVAVDSEISVNAAMLQEGLARLEKRNRWDGKERQQALDNLVPFQGEARTSRRGMWQYGDI 361
Query: 972 ESDDEDTAPPARKAGAGRR 990
+SDDEDTAPPARKAG GR+
Sbjct: 362 QSDDEDTAPPARKAG-GRK 415
>BM519682 similar to GP|10176871|dbj transcription factor-like protein
{Arabidopsis thaliana}, partial (7%)
Length = 434
Score = 177 bits (448), Expect = 3e-44
Identities = 84/94 (89%), Positives = 92/94 (97%)
Frame = +3
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD 60
MASAA+GATGWYRGRVKAVPSGDCLVIVA++S+KPGPLPEK+ITL+SLI PRLARRGGVD
Sbjct: 153 MASAASGATGWYRGRVKAVPSGDCLVIVAISSTKPGPLPEKTITLSSLIAPRLARRGGVD 332
Query: 61 EPFAWESREYLRKLCIGKEVTFRVDYNVASINRD 94
EPFAWESRE+LRKLCIGKEVTFRVDYNV SI+RD
Sbjct: 333 EPFAWESREFLRKLCIGKEVTFRVDYNVPSISRD 434
Score = 55.1 bits (131), Expect = 2e-07
Identities = 28/85 (32%), Positives = 49/85 (56%)
Frame = +3
Query: 380 FTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKE 439
+ G+V V SGDC+++ S PL E+ + LSS+ P++ RR P+A E++E
Sbjct: 186 YRGRVKAVPSGDCLVIVAISSTKPGPLPEKTITLSSLIAPRLA--RRGGVDEPFAWESRE 359
Query: 440 FLRTRLLGRQVSVEMEYSRKIVPTD 464
FLR +G++V+ ++Y+ + D
Sbjct: 360 FLRKLCIGKEVTFRVDYNVPSISRD 434
>AW756012 similar to GP|13111324|db 110 kDa 4SNc-Tudor domain protein {Pisum
sativum}, partial (15%)
Length = 319
Score = 173 bits (438), Expect = 4e-43
Identities = 86/106 (81%), Positives = 92/106 (86%)
Frame = +2
Query: 510 VNVGELVVGRGFGTVIRHRDFEERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLT 569
VNVGEL+V RGFGTVIRHRDFEERSNYYDALLTAES A+SG+KGIHSAKD P MHITDLT
Sbjct: 2 VNVGELIVSRGFGTVIRHRDFEERSNYYDALLTAESLAISGKKGIHSAKDSPSMHITDLT 181
Query: 570 TTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLLIPKETCSIAF 615
T+SAKKAKD L FL+RS IPA VEY L GHRFKLL+P ETC I F
Sbjct: 182 TSSAKKAKDSLSFLYRST*IPARVEYALGGHRFKLLVPDETCIIVF 319
>TC235122 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-Tudor domain protein,
partial (11%)
Length = 599
Score = 119 bits (298), Expect(2) = 3e-42
Identities = 58/72 (80%), Positives = 65/72 (89%)
Frame = +3
Query: 220 QSPQMGRRAAPETVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAK 279
Q+PQMGRRAAPE+VVE EL +D+ +GDVPGEP+ PLTSAQRLAVS+S ETAADPF DAK
Sbjct: 81 QAPQMGRRAAPESVVEPELVSDDTNGDVPGEPQAPLTSAQRLAVSTSAETAADPFAHDAK 260
Query: 280 FFTEMRVLNRDV 291
FFTEMRVLNRDV
Sbjct: 261 FFTEMRVLNRDV 296
Score = 72.4 bits (176), Expect(2) = 3e-42
Identities = 35/36 (97%), Positives = 36/36 (99%)
Frame = +2
Query: 291 VRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVEN 326
VR+VLEGVDKFSNLIGSVYYPDGESAKDLALELVEN
Sbjct: 365 VRLVLEGVDKFSNLIGSVYYPDGESAKDLALELVEN 472
>CA938357 similar to GP|18086332|gb| AT5g07350/T2I1_60 {Arabidopsis
thaliana}, partial (7%)
Length = 267
Score = 115 bits (289), Expect(2) = 6e-34
Identities = 57/61 (93%), Positives = 60/61 (97%)
Frame = +3
Query: 645 DVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSA 704
DVEIEVETVDR GTFLGSLWESRTNVA+TLLEAGLAKLQTSFGSDRIP+FHLLD+AEQSA
Sbjct: 3 DVEIEVETVDRTGTFLGSLWESRTNVAITLLEAGLAKLQTSFGSDRIPDFHLLDQAEQSA 182
Query: 705 K 705
K
Sbjct: 183 K 185
Score = 48.1 bits (113), Expect(2) = 6e-34
Identities = 22/24 (91%), Positives = 23/24 (95%)
Frame = +2
Query: 709 LKIWENFVEGEEVSNGANVESKQQ 732
LKIWENFVEGEEVSNGA VE+KQQ
Sbjct: 194 LKIWENFVEGEEVSNGAAVENKQQ 265
>BG790084 similar to GP|24421095|dbj receptor-like protein kinase {Nicotiana
tabacum}, partial (8%)
Length = 397
Score = 34.7 bits (78), Expect = 0.21
Identities = 16/49 (32%), Positives = 29/49 (58%)
Frame = +1
Query: 228 AAPETVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGP 276
++P+T++ +++ DGDVP P P ++Q+ ++ST AA GP
Sbjct: 163 SSPDTILSRRFLSEDGDGDVPA-PAPAPNNSQKKKSNASTHAAAAAPGP 306
>TC208078
Length = 1117
Score = 30.8 bits (68), Expect = 3.1
Identities = 19/56 (33%), Positives = 29/56 (50%)
Frame = -2
Query: 919 ISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESD 974
IS+N ++L E KR R + E + +D Q+F RK RRG+ Q + E +
Sbjct: 174 ISINGSLLSEEPK*QPKRCR*IKTEGRRRVDQRQRFVASGRKTRRGIEQNDE*EGE 7
>BE211274 similar to PIR|T06699|T0 zinc finger protein T29H11.50 -
Arabidopsis thaliana, partial (9%)
Length = 420
Score = 30.0 bits (66), Expect = 5.3
Identities = 24/61 (39%), Positives = 28/61 (45%)
Frame = +1
Query: 18 AVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEPFAWESREYLRKLCIG 77
+VPSG + ASS P P P + T S TPR AR AW + RK C G
Sbjct: 208 SVPSGRAAIATPSASSSPRPSPSRRHT--SNATPRRARA----PARAW--ARWRRKHCSG 363
Query: 78 K 78
K
Sbjct: 364 K 366
>AW596036
Length = 434
Score = 30.0 bits (66), Expect = 5.3
Identities = 21/58 (36%), Positives = 27/58 (46%)
Frame = -2
Query: 401 PYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSR 458
P PL ERR ++ R K+G RR E + RE FLR LG + + Y R
Sbjct: 277 PRSIPLRERRRRSTTERLEKIGKERRQENHENH-REC-GFLRLVQLGENLKLHCSYKR 110
>TC218507 similar to UP|Q8L9I2 (Q8L9I2) Nodulin MtN21-like protein, partial
(24%)
Length = 859
Score = 30.0 bits (66), Expect = 5.3
Identities = 16/39 (41%), Positives = 21/39 (53%)
Frame = +3
Query: 232 TVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETA 270
TV+ + T + D DVPG+ PP T L S T+TA
Sbjct: 339 TVMWGKATEENVDEDVPGQQSPPTTENVPLLQSYKTDTA 455
>TC203769 similar to UP|RCA_PHAAU (O98997) Ribulose bisphosphate
carboxylase/oxygenase activase, chloroplast precursor
(RuBisCO activase) (RA), partial (42%)
Length = 1336
Score = 29.6 bits (65), Expect = 6.9
Identities = 19/49 (38%), Positives = 26/49 (52%)
Frame = +2
Query: 40 EKSITLASLITPRLARRGGVDEPFAWESREYLRKLCIGKEVTFRVDYNV 88
+K+ L + P AR VD P E+ EY+ +L + EVT RV NV
Sbjct: 872 KKNSRLVLSLVPNSARGRIVDLPEIRETGEYIAELKLHPEVTARVRVNV 1018
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,739,738
Number of Sequences: 63676
Number of extensions: 434185
Number of successful extensions: 1663
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1647
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1661
length of query: 990
length of database: 12,639,632
effective HSP length: 106
effective length of query: 884
effective length of database: 5,889,976
effective search space: 5206738784
effective search space used: 5206738784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0094a.6


