

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
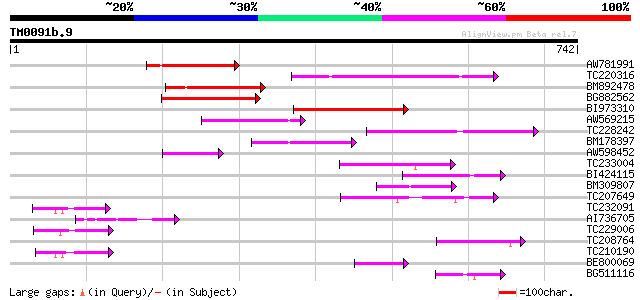
Query= TM0091b.9
(742 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW781991 156 3e-38
TC220316 152 4e-37
BM892478 149 6e-36
BG882562 130 2e-30
BI973310 119 5e-27
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 99 7e-21
TC228242 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired r... 87 3e-17
BM178397 similar to GP|4914326|gb| F14N23.12 {Arabidopsis thalia... 84 2e-16
AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {A... 76 5e-14
TC233004 weakly similar to PIR|G96565|G96565 F6D8.26 [imported] ... 72 7e-13
BI424115 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidop... 69 8e-12
BM309807 similar to GP|5764395|gb| far-red impaired response pro... 63 4e-10
TC207649 similar to UP|Q9SY66 (Q9SY66) F14N23.12 (At1g10240/F14N... 60 4e-09
TC232091 57 2e-08
AI736705 similar to GP|11994736|db far-red impaired response pro... 54 3e-07
TC229006 52 7e-07
TC208764 similar to UP|Q9ZVC9 (Q9ZVC9) Mutator-like transposase ... 45 2e-04
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 45 2e-04
BE800069 44 3e-04
BG511116 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidop... 44 3e-04
>AW781991
Length = 419
Score = 156 bits (395), Expect = 3e-38
Identities = 69/122 (56%), Positives = 92/122 (74%)
Frame = +3
Query: 179 FTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDA 238
FT DC NY+RN R ++L+ GD Q V DY ++ EN NFFYA+Q DED + N+FW D
Sbjct: 54 FTEVDCRNYMRNNRLRSLE-GDIQLVLDYLRQMHAENPNFFYAVQGDEDQSITNVFWADP 230
Query: 239 RSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLF 298
++R++Y FGD +TFDTTY++N+Y +PFA F G+N+H Q +L+GCA L +ESEASFVWLF
Sbjct: 231 KARMNYTFFGDTVTFDTTYRSNRYRLPFAFFTGVNHHGQPVLFGCAFLINESEASFVWLF 410
Query: 299 KT 300
KT
Sbjct: 411 KT 416
>TC220316
Length = 988
Score = 152 bits (385), Expect = 4e-37
Identities = 81/270 (30%), Positives = 142/270 (52%)
Frame = +1
Query: 370 SHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINA 429
S ++ W +Y LK++ WL+ +Y+ R SW+P+Y + TFFAG+ +ES+++
Sbjct: 16 SQTVDEXXATWNXXXNKYGLKDDAWLKEMYQKRASWVPLYLKGTFFAGI---PMNESLDS 186
Query: 430 FFDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTR 489
FF + ++A T L EF+ ++E+ +E R E ERKE++ + + IL T +EE +YT
Sbjct: 187 FFGALLNAQTPLMEFIPRYERGLERRREEERKEDFNTSNFQPILQTKEPVEEQCRRLYTL 366
Query: 490 NIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELY 549
+F F+ EL + + KI +G Y V C + + V + + C +++
Sbjct: 367 TVFKIFQKELLQCFSYLGFKIFEEGGLSRYMVRRCGNDMEKHVVTFNASNISISCSCQMF 546
Query: 550 EFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRR 609
E+ G+LCRH+L +FQ + ++PS +I+ RWT++ G+ + D + S LK L
Sbjct: 547 EYEGVLCRHVLRVFQILQLREVPSRYILHRWTRNTEDGV---FPDMESWSSSQELKNLML 717
Query: 610 VHVQREASFLADLAEESEEAYNFIISEMKQ 639
++ AS D S E Y +++
Sbjct: 718 WSLRETASKYIDAGATSIEKYKLAYEILRE 807
>BM892478
Length = 429
Score = 149 bits (375), Expect = 6e-36
Identities = 67/132 (50%), Positives = 96/132 (71%)
Frame = +3
Query: 204 VFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYS 263
+ +Y + +Q E++ FFYA++ D + M N+FW D RSR S FGDV+ DT+Y+ Y
Sbjct: 30 LLEYFQSRQAEDTGFFYAMEVDNGNCM-NIFWADGRSRYSCSHFGDVLVLDTSYRKTVYL 206
Query: 264 MPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKA 323
+PFA FVG+N+H Q +L GCAL+ DESE SF WLF+T L+AM G+ P+++I DQD+A++
Sbjct: 207 VPFATFVGVNHHKQPVLLGCALIADESEESFTWLFQTWLRAMSGRLPLTVIADQDIAIQR 386
Query: 324 ALAKVFPESRHR 335
A+AKVFP + HR
Sbjct: 387 AIAKVFPVTHHR 422
>BG882562
Length = 416
Score = 130 bits (328), Expect = 2e-30
Identities = 57/130 (43%), Positives = 90/130 (68%)
Frame = +3
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
GD + + +Y R Q+ N NFFY + ++D ++ N+FW+D+RSR +Y FGDV+ FD+T
Sbjct: 27 GDPELISNYFCRIQLMNPNFFYVMDLNDDGQLRNVFWIDSRSRAAYSYFGDVVAFDSTCL 206
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
+N Y +P FVG+N+H +S+L GC LL DE+ +++WLF+ L M G+ P +IIT+
Sbjct: 207 SNNYEIPLVAFVGVNHHGKSVLLGCGLLADETFETYIWLFRAWLTCMTGRPPQTIITNXC 386
Query: 319 LAMKAALAKV 328
AM++A+A+V
Sbjct: 387 KAMQSAIAEV 416
>BI973310
Length = 457
Score = 119 bits (298), Expect = 5e-27
Identities = 60/151 (39%), Positives = 92/151 (60%)
Frame = +3
Query: 372 SIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFF 431
+I FE W ++ + + +NEWLQ +Y R+ W+P+Y R TFF ++ + +E + +FF
Sbjct: 3 TIDEFESYWHSLLERFYVMDNEWLQSIYNSRQHWVPVYLRETFFGEISLNEGNEYLISFF 182
Query: 432 DSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNI 491
D +V +STTLQ V ++EKAV S E E K +Y++ + S +L T S +E+ AAS+YTR I
Sbjct: 183 DGYVFSSTTLQVLVRQYEKAVSSWHEKELKADYDTTNSSPVLKTPSPMEKQAASLYTRKI 362
Query: 492 FAKFKGELEKINRFTRHKIRRDGPKYVYQVS 522
F KF+ EL + KI G Y+V+
Sbjct: 363 FMKFQEELVETLANPAIKIDDSGTITTYRVA 455
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 99.0 bits (245), Expect = 7e-21
Identities = 46/138 (33%), Positives = 80/138 (57%), Gaps = 1/138 (0%)
Frame = +1
Query: 251 ITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKP 310
+ FDTTY+ Y M ++G++N+ + + CALL+DE+ SF W K L M GK P
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGKAP 189
Query: 311 ISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQ-SIFKRDMKRCIRG 369
+I+TD ++ +K A+A P+++H C+WHI+ KF + + + Q +K + R +
Sbjct: 190 QTILTDHNMWLKEAIAVELPQTKHAFCIWHILSKFSDWFSLLLGSQYDEWKAEFHR-LYN 366
Query: 370 SHSIQSFEEEWMRIMVEY 387
++ FEE W +++ +Y
Sbjct: 367 LEQVEDFEEGWRQMVDQY 420
>TC228242 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired response
protein; Mutator-like transposase-like protein;
phytochrome A signaling protein-like, partial (25%)
Length = 1354
Score = 86.7 bits (213), Expect = 3e-17
Identities = 61/227 (26%), Positives = 108/227 (46%), Gaps = 2/227 (0%)
Frame = +2
Query: 468 HKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDA 527
+K L T S LE+ A I+T +F K + E+ R+D V++V +
Sbjct: 17 NKVATLKTPSPLEKSVAGIFTHAVFKKIQAEVIGAVACHPKADRQDDTTIVHRVHDMETN 196
Query: 528 RDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRG 587
+D F V + S+++ C L+E+ G LCRH L + Q G PS +I++RWTKDA
Sbjct: 197 KDFFVVVNQVKSELS-CICRLFEYRGYLCRHALFVLQYSGQSVFPSQYILKRWTKDAK-- 367
Query: 588 LEDTYNDNDLGEKSDTL--KILRRVHVQREASFLADLAEESEEAYNFIISEMKQTHKSAA 645
N +GE+S+ + ++ R + + A L++ S+E+Y + + HKS
Sbjct: 368 -----VRNIMGEESEHMLTRVQRYNDLCQRALKLSEEGSLSQESYGIAFHALHEAHKSCV 532
Query: 646 AMKTSEGVVLLESSEKNVNQVCSSEQVSEPPQLTIGNPHVSQTKGRK 692
++ S E+ + S+E+ ++ ++ N + TK +K
Sbjct: 533 SVNNSSKSSPTEAGTPGAHGQLSTEEDTQSRNMSKSNKKKNPTKKKK 673
>BM178397 similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana}, partial
(20%)
Length = 422
Score = 84.3 bits (207), Expect = 2e-16
Identities = 47/138 (34%), Positives = 72/138 (52%), Gaps = 1/138 (0%)
Frame = +1
Query: 317 QDLAMKAALAKVFPESRHRLCLWHIIKKFPEKL-AHIYHKQSIFKRDMKRCIRGSHSIQS 375
Q++ +K AL+ P ++H C+W I+ KFP A + + + +K + R + S++
Sbjct: 1 QNICLKEALSTEMPTTKHAFCIWMIVAKFPSWFNAVLGERYNDWKAEFYR-LYNLESVED 177
Query: 376 FEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFV 435
FE W + + L N + LY R W + RS F AGM TT +S+SINAF F+
Sbjct: 178 FELGWREMACSFGLHSNRHMVNLYSSRSLWALPFLRSHFLAGMTTTGQSKSINAFIQRFL 357
Query: 436 DASTTLQEFVLKFEKAVE 453
A T L FV + AV+
Sbjct: 358 SAQTRLAHFVEQVAVAVD 411
>AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (7%)
Length = 256
Score = 76.3 bits (186), Expect = 5e-14
Identities = 35/80 (43%), Positives = 48/80 (59%)
Frame = +2
Query: 200 DAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKT 259
DA + +YCKR Q EN FYAIQ DED R+ N+ RSR SY+ GD + D Y
Sbjct: 17 DAHNLLEYCKRMQAENHGCFYAIQLDEDDRVSNVL*AGTRSRTSYKHMGDAESLDAAYGK 196
Query: 260 NKYSMPFAPFVGLNNHSQSI 279
++Y++ +AP+ G N H + I
Sbjct: 197 HRYAVSYAPYTGRNCHGRMI 256
>TC233004 weakly similar to PIR|G96565|G96565 F6D8.26 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(8%)
Length = 551
Score = 72.4 bits (176), Expect = 7e-13
Identities = 44/159 (27%), Positives = 75/159 (46%), Gaps = 7/159 (4%)
Frame = +1
Query: 432 DSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNI 491
D +V T+ +EFV K++ + + E + E+R+ S L T E A +YT+ I
Sbjct: 1 DGYVHKHTSFKEFVDKYDLVLHRKHLKEAMADLETRNVSFELKTRCNFEVQLAKVYTKEI 180
Query: 492 FAKFKGELEKI-NRFTRHKIRRDGPKYVYQVSNCYDARD------TFTVDIDLDSQIAKC 544
F KF+ E+E + + F ++ +G Y V + +F V + +C
Sbjct: 181 FQKFQSEVEGMYSCFNTRQVSVNGSIITYVVKERVEVEGNEKGVKSFEVLYETTELDIRC 360
Query: 545 GYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKD 583
L+ + G LCR L + G+ +IPS +I+ RW +D
Sbjct: 361 ICSLFNYKGYLCRRALNVLNYNGIEEIPSRYILHRWRRD 477
>BI424115 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (9%)
Length = 402
Score = 68.9 bits (167), Expect = 8e-12
Identities = 38/136 (27%), Positives = 72/136 (52%), Gaps = 2/136 (1%)
Frame = +2
Query: 515 PKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSH 574
P ++V+ D + + V ++ A C +++E+ GILC+HIL +F V+ +P H
Sbjct: 2 PNSTFRVAKFEDDQKAYMVTLNHSELKANCSCQMFEYAGILCKHILTVFTVTNVLTLPPH 181
Query: 575 FIMERWTKDA--NRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNF 632
+I++RWT++A + GL++ ++ E R ++ +EA A+ + E YN
Sbjct: 182 YILKRWTRNAKNSAGLDEHTGESHAQESLTA----RYGNLCKEAIRYAEEGSVTVETYNA 349
Query: 633 IISEMKQTHKSAAAMK 648
IS +++ K A +K
Sbjct: 350 AISGLREGVKKVANVK 397
>BM309807 similar to GP|5764395|gb| far-red impaired response protein
{Arabidopsis thaliana}, partial (15%)
Length = 428
Score = 63.2 bits (152), Expect = 4e-10
Identities = 36/105 (34%), Positives = 55/105 (52%)
Frame = +3
Query: 480 EEHAASIYTRNIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDS 539
E+ +++YT IF KF+ E+ + DG + V + Y+ + F V + S
Sbjct: 48 EKQMSTVYTHAIFKKFQVEVLGVAGCQSRIEAGDGTIAKFIVQD-YEKDEEFLVTWNELS 224
Query: 540 QIAKCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDA 584
C L+E+ G LCRH L + Q G +PSH+I++RWTKDA
Sbjct: 225 SEVSCFCRLFEYKGFLCRHALSVLQRCGCSCVPSHYILKRWTKDA 359
>TC207649 similar to UP|Q9SY66 (Q9SY66) F14N23.12 (At1g10240/F14N23_12),
partial (30%)
Length = 971
Score = 60.1 bits (144), Expect = 4e-09
Identities = 49/221 (22%), Positives = 96/221 (43%), Gaps = 15/221 (6%)
Frame = +2
Query: 434 FVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFA 493
F+ A T L FV + AV+ + + ++ + ++ L TG+ +E HAA+I T F+
Sbjct: 116 FLSAQTRLAHFVEQVAVAVDFKDQTG*QQTMQQNLQNVCLKTGAPMESHAATILTPFAFS 295
Query: 494 KFKGELEKINRFT----------RHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAK 543
K + +L + RH + +G + VY I
Sbjct: 296 KLQEQLVLAAHYASFSIEDGFLVRHHTKAEGGRKVYWAPQ---------------EGIIS 430
Query: 544 CGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTK---DANRGLEDTYNDNDLGEK 600
C +EF GILCRH L + QIP ++ RW + +++ L+ ND+
Sbjct: 431 CSCHQFEFTGILCRHSLRVLSTGNCFQIPDRYLPIRWRRINMPSSKLLQSAPNDH----- 595
Query: 601 SDTLKILRRV--HVQREASFLADLAEESEEAYNFIISEMKQ 639
++ +K+L+ + + E++ + + + E ++S +++
Sbjct: 596 AERVKLLQNMVSSLMTESAKSKERLDIATEQVTLLLSRIRE 718
>TC232091
Length = 530
Score = 57.4 bits (137), Expect = 2e-08
Identities = 33/109 (30%), Positives = 57/109 (52%), Gaps = 7/109 (6%)
Frame = +1
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV----RSSKPKRAV---LVCCNEGQHKVKISRT 83
+GMEF+S E + FY +A++ GF VR+ RS K+ + C +G H
Sbjct: 142 VGMEFQSEEAAKNFYEEYARREGFVVRLDRCHRSEVDKQIISRRFSCNKQGFH------- 300
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVM 132
+++ T + ++IR GCEA + V + T KW++ F +H+H++
Sbjct: 301 VRVRNKTKPVHKPRASIREGCEAMMYV-KVNTCGKWVVTKFVKEHSHLL 444
>AI736705 similar to GP|11994736|db far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (14%)
Length = 383
Score = 53.5 bits (127), Expect = 3e-07
Identities = 41/137 (29%), Positives = 65/137 (46%), Gaps = 1/137 (0%)
Frame = +3
Query: 87 QDSTNQT-KRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHK 145
QDS N T +R CS ++ C+AS+ V R + KW+I SF +HNH ++ ++VS + +
Sbjct: 39 QDSENSTGRRSCS--KTDCKASMHVKR-RSDGKWVIHSFVKEHNHELLPAQAVSE-QTRR 206
Query: 146 KMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVF 205
+ A+ E G+ K N R L+ G+A+ +
Sbjct: 207 MYAAMARQFAEYKTVVGLKNEK------------------NPFDKGRNLGLESGEAKLML 332
Query: 206 DYCKRKQVENSNFFYAI 222
D+ + Q NSNFFYA+
Sbjct: 333 DFFIQMQNMNSNFFYAV 383
>TC229006
Length = 991
Score = 52.4 bits (124), Expect = 7e-07
Identities = 37/112 (33%), Positives = 61/112 (54%), Gaps = 7/112 (6%)
Frame = +2
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRS-SKPKR-----AVLVCCNEGQHKVKISRTEE 85
GMEFES + + FY+ +A++ GF +RV S +P+R A + CN+ + V
Sbjct: 110 GMEFESEDAAKIFYDEYARRLGFVMRVMSCRRPERDGRILARRLGCNKEGYCV------S 271
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNH-VMVSPK 136
I+ ++ ++ R GC+A +I + KW+I F DHNH ++VSP+
Sbjct: 272 IRGKFASVRKPRASTREGCKA-MIHIKYNKSGKWVITKFVKDHNHPLVVSPR 424
>TC208764 similar to UP|Q9ZVC9 (Q9ZVC9) Mutator-like transposase
(At2g27110/T20P8.16), partial (4%)
Length = 774
Score = 44.7 bits (104), Expect = 2e-04
Identities = 30/127 (23%), Positives = 56/127 (43%), Gaps = 11/127 (8%)
Frame = +2
Query: 559 ILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASF 618
+L +F+ V+ +PSH+I++RWT++A + + D+ I+R ++ EA
Sbjct: 2 VLAVFRVTNVLTLPSHYILKRWTRNAKSNVILEEHACDVYTYYLESHIVRYNTLRHEAFK 181
Query: 619 LADLAEESEEAYNFIISEMKQTHKSAAAMKTSEGV-----------VLLESSEKNVNQVC 667
D S E Y+ + +++ K + +EG VL + S N C
Sbjct: 182 FVDEGARSAETYDVAMDALQEAAKRVSQGMQNEGKIPINNGKVRSHVLNDESHANYTSGC 361
Query: 668 SSEQVSE 674
E +S+
Sbjct: 362 QEESLSQ 382
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 44.7 bits (104), Expect = 2e-04
Identities = 34/110 (30%), Positives = 50/110 (44%), Gaps = 7/110 (6%)
Frame = +1
Query: 34 EFESIEKVREFYNSFAKKNGFGVRV----RSSKPKRAV---LVCCNEGQHKVKISRTEEI 86
EF S FYN++A + GF VRV RS + A+ LVC EG +
Sbjct: 4 EFGSEAAAHAFYNAYATEVGFIVRVSKLSRSRRDGTAIGRTLVCNKEG---------FRM 156
Query: 87 QDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPK 136
D + R+ + R GC A ++V R + KW++ F +H H + K
Sbjct: 157 ADKREKIVRQRAETRVGCRAMIMV-RKLSSGKWVVAKFVKEHTHPLTPGK 303
>BE800069
Length = 233
Score = 43.9 bits (102), Expect = 3e-04
Identities = 24/71 (33%), Positives = 38/71 (52%)
Frame = +1
Query: 452 VESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIR 511
+ESR E E + +Y++ + +L T S +E+ A+ +YTR IF +F+ EL K
Sbjct: 1 LESRNEKEVRADYDTMNTLPVLRTPSPMEKQASELYTRKIFMRFQEELVGTLTLMASKAD 180
Query: 512 RDGPKYVYQVS 522
DG Y V+
Sbjct: 181 DDGEVITYHVA 213
>BG511116 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (7%)
Length = 296
Score = 43.9 bits (102), Expect = 3e-04
Identities = 30/95 (31%), Positives = 49/95 (51%), Gaps = 4/95 (4%)
Frame = +1
Query: 558 HILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKI----LRRVHVQ 613
HIL +F V+ +PSH+I++RWT +A + TY + +D L I +R +
Sbjct: 1 HILTVFTVTNVLTLPSHYILKRWTTNAKSDIR-TYE-----KITDPLDIENLTVRFNSLC 162
Query: 614 REASFLADLAEESEEAYNFIISEMKQTHKSAAAMK 648
REA LA+ + E YN ++ +++ K MK
Sbjct: 163 REAIKLAEEGAIAVETYNATMNALREGAKRVGIMK 267
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,721,853
Number of Sequences: 63676
Number of extensions: 421549
Number of successful extensions: 2284
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 2235
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2274
length of query: 742
length of database: 12,639,632
effective HSP length: 104
effective length of query: 638
effective length of database: 6,017,328
effective search space: 3839055264
effective search space used: 3839055264
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0091b.9


