

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.13
(267 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
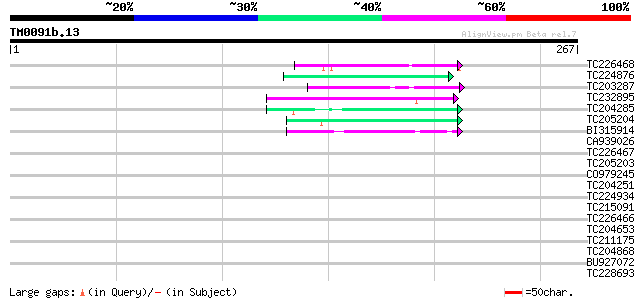
Score E
Sequences producing significant alignments: (bits) Value
TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich prote... 46 2e-05
TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial ... 42 2e-04
TC203287 weakly similar to UP|Q41645 (Q41645) Extensin (Fragment... 42 4e-04
TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfat... 42 4e-04
TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 41 5e-04
TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein pr... 40 8e-04
BI315914 similar to PIR|T09854|T098 proline-rich cell wall prote... 40 8e-04
CA939026 40 0.001
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 40 0.001
TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall pr... 40 0.001
CO979245 40 0.001
TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Sy... 40 0.001
TC224934 similar to UP|Q9SII5 (Q9SII5) Expressed protein (At2g17... 39 0.002
TC215091 UP|CUT2_CAEEL (P34682) Cuticlin 2 precursor, partial (10%) 39 0.002
TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {... 39 0.002
TC204653 similar to UP|Q9AT20 (Q9AT20) Histone H1 (Fragment), pa... 39 0.002
TC211175 weakly similar to UP|Q9LIE8 (Q9LIE8) Similarity to cell... 39 0.002
TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, par... 39 0.002
BU927072 39 0.002
TC228693 similar to UP|Q8S9A6 (Q8S9A6) Glucosyltransferase-3, pa... 39 0.003
>TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich protein, partial
(20%)
Length = 1041
Score = 45.8 bits (107), Expect = 2e-05
Identities = 34/85 (40%), Positives = 41/85 (48%), Gaps = 6/85 (7%)
Frame = -2
Query: 135 KPSPSMPAQPKT---DSPK--PPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
KP+P P+ PK SPK PPQI QP P + PP P A+ PP AP
Sbjct: 803 KPAPVKPSAPKVAPASSPKAPPPQIPIPQPPKPSPVSPPPLPPPPVASPPPLPPPKIAP- 627
Query: 190 KKALSQHVSAPKKVTPSKRPA-TQP 213
A + AP K TP+ PA T+P
Sbjct: 626 TPAKTPPSPAPAKATPAPAPAPTKP 552
Score = 29.6 bits (65), Expect = 1.5
Identities = 19/77 (24%), Positives = 27/77 (34%)
Frame = -2
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P P + P PP P P P K++P+ A + P +P
Sbjct: 692 PPPLPPPPVASPPPLPPPKIAPTPAKTPPSPAPAKATPAPAPAPTKPAPTPSP------- 534
Query: 196 HVSAPKKVTPSKRPATQ 212
V P TP+ P +
Sbjct: 533 -VPPPPAATPAPAPVIE 486
>TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial (66%)
Length = 1340
Score = 42.4 bits (98), Expect = 2e-04
Identities = 23/80 (28%), Positives = 30/80 (36%)
Frame = +1
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP+ PSP P + SP PP + PP +P +PP P
Sbjct: 181 SPPYHYPSPPPPPKYYYHSPPPPSPSPPKKPYYYHSPPPPSPTPPKKPYYYHSPPPPPPK 360
Query: 190 KKALSQHVSAPKKVTPSKRP 209
KK H P P K+P
Sbjct: 361 KKPYYYHSPPPPSPPPPKKP 420
Score = 35.4 bits (80), Expect = 0.027
Identities = 27/84 (32%), Positives = 34/84 (40%), Gaps = 3/84 (3%)
Frame = +1
Query: 129 YNPPFKKPSPSMPAQPKT-DSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPP--N 185
++PP PSPS P +P SP PP PK PP P +PP +
Sbjct: 232 HSPP--PPSPSPPKKPYYYHSPPPPS--PTPPKKPYYYHSPPPPPPKKKPYYYHSPPPPS 399
Query: 186 TAPMKKALSQHVSAPKKVTPSKRP 209
P KK H P +P K+P
Sbjct: 400 PPPPKKPYYYHSPPPPSPSPPKKP 471
Score = 30.0 bits (66), Expect = 1.1
Identities = 23/83 (27%), Positives = 33/83 (39%), Gaps = 2/83 (2%)
Frame = +1
Query: 129 YNPPFKKPS-PSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
++PP P+ P P + P PP+ K + P + PP P PP+ +
Sbjct: 283 HSPPPPSPTPPKKPYYYHSPPPPPPKKKPYYYHSPPPPSPPPPKKPYYYHSP--PPPSPS 456
Query: 188 PMKKALSQHVSAPKKVTPS-KRP 209
P KK H P P K+P
Sbjct: 457 PPKKPYYYHSPPPPSPPPPYKKP 525
>TC203287 weakly similar to UP|Q41645 (Q41645) Extensin (Fragment), partial
(14%)
Length = 967
Score = 41.6 bits (96), Expect = 4e-04
Identities = 25/74 (33%), Positives = 34/74 (45%)
Frame = +3
Query: 141 PAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAP 200
P QP T +P PP + P A PP ++P AA PP T+P A + +
Sbjct: 138 PTQPPTTTPSPPPPRSAPAPAPTPPATPPPATPPPAAAT--PPPATSP--PATTPTPTPT 305
Query: 201 KKVTPSKRPATQPL 214
TP+ PAT P+
Sbjct: 306 AAPTPASSPATSPV 347
Score = 35.0 bits (79), Expect = 0.035
Identities = 29/89 (32%), Positives = 37/89 (40%), Gaps = 1/89 (1%)
Frame = +3
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
P T PP P+P+ P P T P PP P P A P +P+ AA
Sbjct: 150 PTTTPSPPPPRSAPAPA-PTPPATPPPATPPPAAATPPPATSPPATTPTPTPT-AAPTPA 323
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPA 210
+ P T+P+ A S AP P+ PA
Sbjct: 324 SSPATSPVPSASSP--PAPGTAGPAPGPA 404
>TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (14%)
Length = 907
Score = 41.6 bits (96), Expect = 4e-04
Identities = 25/91 (27%), Positives = 39/91 (42%), Gaps = 1/91 (1%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
+P + R N P PSPS + SP P +P P + P ++P+ A
Sbjct: 249 SPSSPPRRNTPPPPPSPSSSTPLRPSSPPPTTSPTSRPSRPTPSSPSPSTAPAPAPTTST 428
Query: 182 APPNTAPMK-KALSQHVSAPKKVTPSKRPAT 211
PP + K K S + P + +P RP++
Sbjct: 429 TPPTPSKSKAKPTSPSPTTPTRPSPRVRPSS 521
Score = 33.5 bits (75), Expect = 0.10
Identities = 30/105 (28%), Positives = 44/105 (41%), Gaps = 6/105 (5%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE--- 178
T T+ R+ PP SPS P + T P PP P + P + PP +SP++
Sbjct: 213 TSATVSRHAPP---TSPSSPPRRNTPPP-PPSPSSSTP--LRPSSPPPTTSPTSRPSRPT 374
Query: 179 AIRAPPNTAPM---KKALSQHVSAPKKVTPSKRPATQPLRRSNRL 220
P+TAP + + + K P+ T P R S R+
Sbjct: 375 PSSPSPSTAPAPAPTTSTTPPTPSKSKAKPTSPSPTTPTRPSPRV 509
>TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (32%)
Length = 1539
Score = 41.2 bits (95), Expect = 5e-04
Identities = 30/95 (31%), Positives = 35/95 (36%), Gaps = 3/95 (3%)
Frame = +1
Query: 122 TPITMIRYNPP---FKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
TP + Y PP KKP P +P +P PP C PK V P PP P
Sbjct: 151 TPPPVPVYTPPPVPEKKPPPPVPEKP------PP----CPPKVVPPPKPPPVKPPPKPPV 300
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP P V P P K+P + P
Sbjct: 301 KPPCPPKVEPPHPPKPPKVKPPHPPHPPKKPPSPP 405
Score = 39.3 bits (90), Expect = 0.002
Identities = 30/100 (30%), Positives = 37/100 (37%), Gaps = 7/100 (7%)
Frame = +1
Query: 121 CTPITMIRYNPPFKKPSPSMPAQP----KTDS---PKPPQIKGCQPKNVLPKADPPKSSP 173
C P + PP KP P P +P K + PKPP++K P + PPK P
Sbjct: 235 CPPKVVPPPKPPPVKPPPKPPVKPPCPPKVEPPHPPKPPKVKPPHPPH------PPKKPP 396
Query: 174 SNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
S PP P V P P K+P P
Sbjct: 397 S--------PPKVKPPHPPKPPKVEPPHPPKPPKKPPCPP 492
Score = 37.7 bits (86), Expect = 0.005
Identities = 30/97 (30%), Positives = 37/97 (37%), Gaps = 13/97 (13%)
Frame = +1
Query: 130 NPPFKKPSPSM------PAQPKTDS---PKPPQIKGCQPKNVLP-KADPPKSSPSNAAEA 179
+PP K PSP P PK + PKPP+ C PK P PPK P + +
Sbjct: 376 HPPKKPPSPPKVKPPHPPKPPKVEPPHPPKPPKKPPCPPKVKPPHPPKPPKVEPPHPPKP 555
Query: 180 IR---APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ +PP P P P K P T P
Sbjct: 556 PKKPPSPPKVVPPLPPKPPKNPPPLPPKPPKEPPTPP 666
Score = 36.2 bits (82), Expect = 0.016
Identities = 28/99 (28%), Positives = 33/99 (33%), Gaps = 5/99 (5%)
Frame = +1
Query: 121 CTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
C P + P K P P P P PP++K P PK +PP P +
Sbjct: 310 CPPKVEPPHPPKPPKVKPPHPPHPPKKPPSPPKVKPPHPPKP-PKVEPPH--PPKPPKKP 480
Query: 181 RAPPNTAPMK-----KALSQHVSAPKKVTPSKRPATQPL 214
PP P K H P K PS PL
Sbjct: 481 PCPPKVKPPHPPKPPKVEPPHPPKPPKKPPSPPKVVPPL 597
Score = 32.0 bits (71), Expect = 0.30
Identities = 24/75 (32%), Positives = 29/75 (38%), Gaps = 5/75 (6%)
Frame = +1
Query: 121 CTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQI----KGCQPKNVLP-KADPPKSSPSN 175
C P + P K P P +P P PP++ PKN P PPK P+
Sbjct: 484 CPPKVKPPHPPKPPKVEPPHPPKPPKKPPSPPKVVPPLPPKPPKNPPPLPPKPPKEPPTP 663
Query: 176 AAEAIRAPPNTAPMK 190
E PP T P K
Sbjct: 664 PKE----PPPTPPKK 696
Score = 31.6 bits (70), Expect = 0.39
Identities = 26/89 (29%), Positives = 34/89 (37%), Gaps = 10/89 (11%)
Frame = +1
Query: 131 PP--FKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKAD---PPKSSPSNAAEAIRAPPN 185
PP F KP P + P PP + +P +P+ PPK P ++ PP
Sbjct: 112 PPLIFHKPPPPVYTPPPVPVYTPPPVPEKKPPPPVPEKPPPCPPKVVPPPKPPPVKPPPK 291
Query: 186 TAPMK-----KALSQHVSAPKKVTPSKRP 209
P+K K H P KV P P
Sbjct: 292 -PPVKPPCPPKVEPPHPPKPPKVKPPHPP 375
Score = 30.8 bits (68), Expect = 0.66
Identities = 28/84 (33%), Positives = 31/84 (36%), Gaps = 4/84 (4%)
Frame = +1
Query: 130 NPPFKKPSPSM--PAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP-PNT 186
NPP P P P PK P PP+ K C P P PK P + P P
Sbjct: 616 NPPPLPPKPPKEPPTPPKEPPPTPPK-KPCPPIFKKPLPPIPKLPPLPPVPIYKKPLPPL 792
Query: 187 APMKKALSQH-VSAPKKVTPSKRP 209
P+ K H V KK P P
Sbjct: 793 PPLPKLPPLHPVPIYKKPLPPLPP 864
>TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (44%)
Length = 996
Score = 40.4 bits (93), Expect = 8e-04
Identities = 24/84 (28%), Positives = 33/84 (38%), Gaps = 1/84 (1%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP + P +P+ P SP P P + P A PP SSP A+ +PP +P
Sbjct: 226 PPIQSSPPPVPSSPPPAQSPPPASTPPPAPLSSPPPASPPPSSPPPASPPPASPPPASPP 405
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ P P+ P P
Sbjct: 406 PASPPPASPPPASPPPATPPPASP 477
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/83 (30%), Positives = 33/83 (39%), Gaps = 2/83 (2%)
Frame = +1
Query: 130 NPPFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
+PP P P+ P A P SP P P P A PP +SP + PP
Sbjct: 352 SPPPASPPPASPPPASPPPASPPPASPPPASP----PPATPPPASPPPFSPPPATPPPAT 519
Query: 188 PMKKALSQHVSAPKKVTPSKRPA 210
P +S+P +P+ PA
Sbjct: 520 PPPALTPTPLSSPPATSPAPAPA 588
Score = 37.0 bits (84), Expect = 0.009
Identities = 26/90 (28%), Positives = 33/90 (35%), Gaps = 4/90 (4%)
Frame = +1
Query: 130 NPPFKKPSPSMP--AQPKTDSPKP--PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
+PP P PS P A P SP P P P + P + PP + P + PP
Sbjct: 322 SPPPASPPPSSPPPASPPPASPPPASPPPASPPPASPPPASPPPATPPPASPPPFSPPPA 501
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPATQPLR 215
T P P P+ PA P +
Sbjct: 502 TPPPATPPPALTPTPLSSPPATSPAPAPAK 591
Score = 36.2 bits (82), Expect = 0.016
Identities = 27/86 (31%), Positives = 34/86 (39%), Gaps = 3/86 (3%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP-- 188
PP P P+ + P SP P P P A PP +SP A+ +PP +P
Sbjct: 286 PPASTPPPAPLSSPPPASPPPSSPPPASP----PPASPPPASPPPASPPPASPPPASPPP 453
Query: 189 -MKKALSQHVSAPKKVTPSKRPATQP 213
S +P TP PAT P
Sbjct: 454 ATPPPASPPPFSPPPATPP--PATPP 525
Score = 35.8 bits (81), Expect = 0.021
Identities = 26/87 (29%), Positives = 36/87 (40%)
Frame = +1
Query: 127 IRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT 186
I+ +PP SP P S PP P P + PP +SP A+ +PP
Sbjct: 232 IQSSPPPVPSSPPPAQSPPPASTPPPAPLSSPPPASPPPSSPPPASPPPASPPPASPPPA 411
Query: 187 APMKKALSQHVSAPKKVTPSKRPATQP 213
+P S ++P TP PA+ P
Sbjct: 412 SP--PPASPPPASPPPATPP--PASPP 480
Score = 32.0 bits (71), Expect = 0.30
Identities = 23/67 (34%), Positives = 30/67 (44%), Gaps = 8/67 (11%)
Frame = +1
Query: 130 NPPFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVL---PKADPPKSSPSNAAEAIRAP- 183
+PP P P+ P A P SP P P L P + PP +SP+ A + AP
Sbjct: 427 SPPPASPPPATPPPASPPPFSPPPATPPPATPPPALTPTPLSSPPATSPAPAPAKVVAPA 606
Query: 184 --PNTAP 188
P+ AP
Sbjct: 607 LSPSLAP 627
Score = 30.4 bits (67), Expect = 0.87
Identities = 25/104 (24%), Positives = 40/104 (38%), Gaps = 5/104 (4%)
Frame = +1
Query: 115 LPFGVLCTPITMIRYNPPFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVLPKADPPKSS 172
L ++ T + + P P+P P + P S PP P P A P +
Sbjct: 133 LLLALIFTVVAGVGAQGPSTSPAPPTPQSSPPPIQSSPPPVPSSPPPAQSPPPASTPPPA 312
Query: 173 PSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTP---SKRPATQP 213
P ++ PP++ P S ++P +P S PA+ P
Sbjct: 313 PLSSPPPASPPPSSPP---PASPPPASPPPASPPPASPPPASPP 435
>BI315914 similar to PIR|T09854|T098 proline-rich cell wall protein - upland
cotton, partial (37%)
Length = 246
Score = 40.4 bits (93), Expect = 8e-04
Identities = 29/83 (34%), Positives = 35/83 (41%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P PS + P SP P P + P A PP SSP A+ +PP + P
Sbjct: 13 PPASTPPPSPLSSPPPSSPPPSS----PPPSSPPPASPPPSSPPPASPPPSSPPPSTPPP 180
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
S +P TP PAT P
Sbjct: 181 S--SPPPFSPPPATPP--PATPP 237
Score = 35.4 bits (80), Expect = 0.027
Identities = 18/59 (30%), Positives = 25/59 (41%)
Frame = +1
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+PP P PS P P + P P P + P + PP + P ++ PP T P
Sbjct: 49 SPPPSSPPPSSP-PPSSPPPASPPPSSPPPASPPPSSPPPSTPPPSSPPPFSPPPATPP 222
Score = 34.3 bits (77), Expect = 0.060
Identities = 19/59 (32%), Positives = 23/59 (38%)
Frame = +1
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+PP P PS P P + P P P + P PP S P + PP T P
Sbjct: 64 SPPPSSPPPSSPP-PASPPPSSPPPASPPPSSPPPSTPPPSSPPPFSPPPATPPPATPP 237
Score = 32.0 bits (71), Expect = 0.30
Identities = 18/68 (26%), Positives = 27/68 (39%)
Frame = +1
Query: 146 TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTP 205
+ SP P P + P + PP SSP ++ +PP ++P + P P
Sbjct: 1 SQSPPPASTPPPSPLSSPPPSSPPPSSPPPSSPPPASPPPSSPPPASPPPSSPPPSTPPP 180
Query: 206 SKRPATQP 213
S P P
Sbjct: 181 SSPPPFSP 204
>CA939026
Length = 426
Score = 40.0 bits (92), Expect = 0.001
Identities = 26/88 (29%), Positives = 40/88 (44%), Gaps = 1/88 (1%)
Frame = +3
Query: 134 KKPSPSMPAQPKTD-SPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKA 192
K+ S + A P + SP PQI+ P N P SPS+ + PP ++
Sbjct: 39 KRKSMAETASPSSKPSPMDPQIQNPAPSNSNSNPPIPSPSPSHNMASPSLPPLPQDQQQQ 218
Query: 193 LSQHVSAPKKVTPSKRPATQPLRRSNRL 220
L Q +S P++ ++ QPL SN +
Sbjct: 219 LQQQLSPPQQQQQQQQQQQQPLVSSNSM 302
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 40.0 bits (92), Expect = 0.001
Identities = 33/89 (37%), Positives = 39/89 (43%), Gaps = 10/89 (11%)
Frame = +3
Query: 135 KPSPSMPAQPKT---DSPK--PPQIKGCQPKNV----LPKADPPKSSPSNAAEAIRAPPN 185
KP+P P PK SPK PPQ QP V P + PP P + PP
Sbjct: 216 KPAPVTPPAPKVAPASSPKAPPPQTPQPQPPKVSPVVTPTSPPPIPPPPVSTPPPLPPPK 395
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPA-TQP 213
AP A + AP K TP+ PA T+P
Sbjct: 396 IAPTP-AKTPPAPAPAKATPAPAPAPTKP 479
Score = 37.7 bits (86), Expect = 0.005
Identities = 26/83 (31%), Positives = 33/83 (39%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP + P P P +P P P + P PPK +P+ A + PP AP
Sbjct: 276 PPPQTPQPQPPKVSPVVTPTSPPPIPPPPVSTPPPLPPPKIAPTPA----KTPPAPAP-A 440
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
KA AP K P+ P P
Sbjct: 441 KATPAPAPAPTKPAPTPAPVPPP 509
Score = 30.8 bits (68), Expect = 0.66
Identities = 21/81 (25%), Positives = 30/81 (36%)
Frame = +3
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P P T P PP P P P K++P+ A + P AP+ +
Sbjct: 339 PPPIPPPPVSTPPPLPPPKIAPTPAKTPPAPAPAKATPAPAPAPTKPAPTPAPVPPPPTL 518
Query: 196 HVSAPKKVTPSKRPATQPLRR 216
P P+ P++ RR
Sbjct: 519 -APTPVVEVPAPAPSSHKHRR 578
>TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall protein,
partial (48%)
Length = 1076
Score = 39.7 bits (91), Expect = 0.001
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 2/85 (2%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP + P +P+ P + SP P P + P + PP SSP ++ +PP ++P
Sbjct: 318 PPVQSSPPPVPSSPPPSQSPPPASTPPPSPLSSPPPSSPPPSSPPPSSPPPASPPPSSPP 497
Query: 190 KKALSQHVSAPKKVTP-SKRPATQP 213
+ P P S PAT P
Sbjct: 498 PASPPPSSPPPSSPPPFSPPPATPP 572
Score = 39.3 bits (90), Expect = 0.002
Identities = 27/90 (30%), Positives = 35/90 (38%), Gaps = 7/90 (7%)
Frame = +3
Query: 131 PPFKKPSPSM-----PAQPKTDSPKP--PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP 183
PP P PS P+ P SP P P P + P + PP S P ++ P
Sbjct: 378 PPASTPPPSPLSSPPPSSPPPSSPPPSSPPPASPPPSSPPPASPPPSSPPPSSPPPFSPP 557
Query: 184 PNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
P T P A P +TP+ P + P
Sbjct: 558 PATPP--PATPPPAVPPPSLTPTVTPLSSP 641
Score = 36.6 bits (83), Expect = 0.012
Identities = 28/90 (31%), Positives = 35/90 (38%), Gaps = 6/90 (6%)
Frame = +3
Query: 130 NPPFKKPSPSMP--AQPKTDSPKP--PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
+PP P PS P A P SP P P P + P + PP + P PP+
Sbjct: 429 SPPPSSPPPSSPPPASPPPSSPPPASPPPSSPPPSSPPPFSPPPATPPPATPPPAVPPPS 608
Query: 186 TAPMKKALSQHVSAPKKVTPSK--RPATQP 213
P LS + +PSK PA P
Sbjct: 609 LTPTVTPLSSPPVSSPAPSPSKVFAPALSP 698
Score = 32.7 bits (73), Expect = 0.17
Identities = 21/61 (34%), Positives = 27/61 (43%), Gaps = 3/61 (4%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP---PNTA 187
PPF P PA P +P P V P + PP SSP+ + + AP P+ A
Sbjct: 540 PPFSPP----PATPPPATPPPAVPPPSLTPTVTPLSSPPVSSPAPSPSKVFAPALSPSLA 707
Query: 188 P 188
P
Sbjct: 708 P 710
Score = 31.6 bits (70), Expect = 0.39
Identities = 22/84 (26%), Positives = 29/84 (34%), Gaps = 2/84 (2%)
Frame = +3
Query: 132 PFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
P P+P P + P S PP P P A P SP + +PP ++P
Sbjct: 276 PSTSPAPPTPQASPPPVQSSPPPVPSSPPPSQSPPPASTPPPSP------LSSPPPSSPP 437
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ P PS P P
Sbjct: 438 PSSPPPSSPPPASPPPSSPPPASP 509
>CO979245
Length = 730
Score = 39.7 bits (91), Expect = 0.001
Identities = 31/93 (33%), Positives = 38/93 (40%), Gaps = 9/93 (9%)
Frame = +3
Query: 130 NPPFK----KPSP--SMPAQPKTD---SPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
NPP + KP+P + PA+P +PKPP PK + A PPK P AA
Sbjct: 150 NPPVEEAAPKPTPGAAAPAKPPPGVAAAPKPPPGAAAAPKPLPGAAAPPKP-PPGAAPPP 326
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP P L + PK PA P
Sbjct: 327 NPPPEAIPENPPLPVDATLPKPRPVDAMPANPP 425
Score = 35.4 bits (80), Expect = 0.027
Identities = 27/91 (29%), Positives = 35/91 (37%), Gaps = 8/91 (8%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKP--------PQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
PP + +P P ++PKP P ++ PK P A P P A A +
Sbjct: 66 PPPEPKAPKPPLPWLLNAPKPPPEPAPANPPVEEAAPKPT-PGAAAPAKPPPGVAAAPKP 242
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP A K L +AP K P P P
Sbjct: 243 PPGAAAAPKPL-PGAAAPPKPPPGAAPPPNP 332
Score = 32.3 bits (72), Expect = 0.23
Identities = 24/77 (31%), Positives = 35/77 (45%), Gaps = 4/77 (5%)
Frame = +3
Query: 116 PFGVLCTPI-TMIRYNPPFKKPSPSMPAQPKTDSPKPPQI--KGCQPKNVLPKA-DPPKS 171
P V P+ ++ NPP K + P D PKPP+ + C P + +PP +
Sbjct: 426 PIAVFEKPLGELLPANPPPKPLGAAKPLPSCLDCPKPPRTDPRTCCPDPICAGCPNPPGA 605
Query: 172 SPSNAAEAIRAPPNTAP 188
AA A++ PP AP
Sbjct: 606 CAGTAATALK-PPXNAP 653
>TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Syndrome
protein homolog, partial (8%)
Length = 1045
Score = 39.7 bits (91), Expect = 0.001
Identities = 31/109 (28%), Positives = 41/109 (37%), Gaps = 23/109 (21%)
Frame = +3
Query: 131 PPFKKPS--PSMPAQPKTDSPKPPQIKGC------------------QPKNVLPKADPPK 170
PP PS PS P + SP PP + +P + P PP
Sbjct: 225 PPSAPPSTSPSTPLTAPSPSPSPPSLPPAPAGSPGASTSSAMACAAPKPSSPSPPPPPPP 404
Query: 171 SSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVT---PSKRPATQPLRR 216
SPS A + P+T ++ S PK T PS P+ P+RR
Sbjct: 405 PSPSAATTSPPTKPSTKSKPLPSTRGTSRPKNPTAP*PSTAPSRSPIRR 551
>TC224934 similar to UP|Q9SII5 (Q9SII5) Expressed protein
(At2g17230/T23A1.9), partial (70%)
Length = 953
Score = 39.3 bits (90), Expect = 0.002
Identities = 22/74 (29%), Positives = 31/74 (41%)
Frame = +3
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQH 196
SPS P +P SP P P + P A +SP + + P+ +P A
Sbjct: 390 SPSTPTRPAPTSPAPSPSPASTPTSATPTAHTSPASPCRTSSPPPSRPSPSPSTTATGST 569
Query: 197 VSAPKKVTPSKRPA 210
S+P K +P K A
Sbjct: 570 *SSPPKTSPWKTSA 611
Score = 37.0 bits (84), Expect = 0.009
Identities = 30/114 (26%), Positives = 45/114 (39%), Gaps = 2/114 (1%)
Frame = +3
Query: 116 PFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN 175
P G +P T R P PSPS + P + +P + P P SPS
Sbjct: 372 PTGGALSPSTPTRPAPTSPAPSPSPASTPTSATPTAHTSPASPCRTSSPPPSRPSPSPST 551
Query: 176 AAEAI--RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKN 227
A +PP T+P K + +AP V+ + + + RS W ++N
Sbjct: 552 TATGST*SSPPKTSPWKTS-----AAPSAVSTTSPSHPRWVTRSLTPGWGTQEN 698
>TC215091 UP|CUT2_CAEEL (P34682) Cuticlin 2 precursor, partial (10%)
Length = 913
Score = 39.3 bits (90), Expect = 0.002
Identities = 24/88 (27%), Positives = 39/88 (44%)
Frame = +1
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
TP+ + PP + S P PK P+ P + +P++ + + PPK+ P A
Sbjct: 469 TPLGPPKAGPPRAPITTSGP--PKAGPPRAP-VTASEPESPVTPSGPPKAGPPRAPITTS 639
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRP 209
PP P + ++ S P K P + P
Sbjct: 640 GPPKAGPPRAPIT--TSGPPKAGPPRAP 717
Score = 38.9 bits (89), Expect = 0.002
Identities = 25/81 (30%), Positives = 33/81 (39%), Gaps = 1/81 (1%)
Frame = +1
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P P T S P I +P+ + PPK+ P A PP P + ++
Sbjct: 385 PKAGPPRAPVTASEPEPPITASEPEPPVTPLGPPKAGPPRAPITTSGPPKAGPPRAPVT- 561
Query: 196 HVSAPKK-VTPSKRPATQPLR 215
S P+ VTPS P P R
Sbjct: 562 -ASEPESPVTPSGPPKAGPPR 621
Score = 33.1 bits (74), Expect = 0.13
Identities = 20/80 (25%), Positives = 28/80 (35%), Gaps = 1/80 (1%)
Frame = +1
Query: 136 PSPSMPAQPKTDS-PKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
P P P T S P+PP P+ + + P K+ P A PP P + ++
Sbjct: 238 PKAGPPKAPVTASEPEPPVTPSGPPRAPISTSGPLKAGPPRALITTSGPPKAGPPRAPVT 417
Query: 195 QHVSAPKKVTPSKRPATQPL 214
P P PL
Sbjct: 418 ASEPEPPITASEPEPPVTPL 477
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/56 (28%), Positives = 24/56 (42%)
Frame = +1
Query: 154 IKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRP 209
+ +P+ + + PPK+ P A PP P K ++ P VTPS P
Sbjct: 148 VTASEPEPPVTPSGPPKAGPPRALITTSGPPKAGPPKAPVTASEPEP-PVTPSGPP 312
>TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {Arabidopsis
thaliana;} , partial (22%)
Length = 978
Score = 39.3 bits (90), Expect = 0.002
Identities = 34/89 (38%), Positives = 38/89 (42%), Gaps = 10/89 (11%)
Frame = +1
Query: 135 KPSPSMPAQPKT---DSPKPP--QIKGCQPKNVLPKA----DPPKSSPSNAAEAIRAPPN 185
KP+P P PK SPK P Q QP V P A PP P A PP
Sbjct: 214 KPAPVTPPAPKVAPASSPKAPTPQTPPPQPPKVSPVAAPTLPPPLPPPPVATPPPLPPPK 393
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPA-TQP 213
AP A + AP K TP+ PA T+P
Sbjct: 394 IAPTP-AKTSPAPAPAKATPAPAPAPTKP 477
Score = 33.9 bits (76), Expect = 0.078
Identities = 27/83 (32%), Positives = 32/83 (38%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP + P S A P P PP P P PPK +P+ A + P AP
Sbjct: 289 PPPQPPKVSPVAAPTLPPPLPPP-----PVATPPPLPPPKIAPTPA----KTSPAPAP-A 438
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
KA AP K P+ P P
Sbjct: 439 KATPAPAPAPTKPAPTPAPIPPP 507
Score = 30.4 bits (67), Expect = 0.87
Identities = 21/81 (25%), Positives = 30/81 (36%)
Frame = +1
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P P T P PP P P P K++P+ A + P AP+ +
Sbjct: 337 PPPLPPPPVATPPPLPPPKIAPTPAKTSPAPAPAKATPAPAPAPTKPAPTPAPIPPPPTL 516
Query: 196 HVSAPKKVTPSKRPATQPLRR 216
P P+ P++ RR
Sbjct: 517 -APTPIVEVPAPAPSSHKHRR 576
>TC204653 similar to UP|Q9AT20 (Q9AT20) Histone H1 (Fragment), partial (68%)
Length = 1501
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/108 (29%), Positives = 45/108 (41%), Gaps = 27/108 (25%)
Frame = +1
Query: 135 KPSPSMPAQPKTDSPKP---------PQIKGCQPKNVLPKADP----------------- 168
KP +PA+PK +PKP P+ +PK P A P
Sbjct: 643 KPKAKVPAKPKP-APKPKAAAAAASKPKPAAAKPKGTKPAAKPKPKPAAKTKAAAAKPAA 819
Query: 169 -PKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLR 215
PK+ P A+A R T+P KKA A KKV P+K+ + ++
Sbjct: 820 KPKAKP---AKASRTSTRTSPGKKAAPAAKPAAKKVAPAKKAPVKSVK 954
Score = 37.4 bits (85), Expect = 0.007
Identities = 25/84 (29%), Positives = 33/84 (38%), Gaps = 4/84 (4%)
Frame = +1
Query: 134 KKPSPSMPAQPKTDSPKP-PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKA 192
KKP+P+ P PKP P K K KA + + + A P AP KA
Sbjct: 517 KKPAPAKPQPKPKPKPKPKPAAKAKDTKATKSKAPAKPKAAAKPKAKVPAKPKPAPKPKA 696
Query: 193 LSQHVSAPKKVT---PSKRPATQP 213
+ S PK +PA +P
Sbjct: 697 AAAAASKPKPAAAKPKGTKPAAKP 768
Score = 30.4 bits (67), Expect = 0.87
Identities = 22/80 (27%), Positives = 33/80 (40%), Gaps = 3/80 (3%)
Frame = +1
Query: 132 PFKKPSPSMPAQPKTDSPKP---PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
P KP P A+ K + KP P+ K + + P K + A A + AP
Sbjct: 754 PAAKPKPKPAAKTKAAAAKPAAKPKAKPAKASRTSTRTSPGKKAAPAAKPAAK---KVAP 924
Query: 189 MKKALSQHVSAPKKVTPSKR 208
KKA + V +P+K+
Sbjct: 925 AKKAPVKSVKPKSVKSPAKK 984
>TC211175 weakly similar to UP|Q9LIE8 (Q9LIE8) Similarity to cell wall-plasma
membrane linker protein, partial (3%)
Length = 609
Score = 39.3 bits (90), Expect = 0.002
Identities = 26/81 (32%), Positives = 39/81 (48%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P+ PA+ T SP P Q P P PP ++P + A + PP++ P
Sbjct: 193 PPSPATHPAPPAKSPTLSP-PSQSPAVSPSGSTP---PPATNPPAKSPATQ-PPSSVPPA 357
Query: 191 KALSQHVSAPKKVTPSKRPAT 211
+ S +VS+P +P+ P T
Sbjct: 358 SSPSNNVSSPPAPSPATPPPT 420
Score = 36.2 bits (82), Expect = 0.016
Identities = 29/98 (29%), Positives = 35/98 (35%), Gaps = 1/98 (1%)
Frame = +1
Query: 117 FGVLCTPITMIRY-NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN 175
F + T + +R N P PSP P T P P P P PP SP+
Sbjct: 100 FALFFTSLCSVRADNAP--APSPKSSPSPPTAKPPSPATHPAPPAKS-PTLSPPSQSPAV 270
Query: 176 AAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ PP T P P+K PATQP
Sbjct: 271 SPSGSTPPPATNP----------------PAKSPATQP 336
>TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, partial (91%)
Length = 865
Score = 39.3 bits (90), Expect = 0.002
Identities = 27/86 (31%), Positives = 32/86 (36%), Gaps = 4/86 (4%)
Frame = +2
Query: 132 PFKKPSPSMPAQPK----TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
P KP+P P PK T P PP K P PK PP +P+ PP +
Sbjct: 149 PSPKPTPPTPPTPKPTPPTPKPTPPTPKPTPPT---PKPTPPTPTPTPPTPKAGCPPPSP 319
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P K PK P+ P P
Sbjct: 320 P--KPTPPTPPTPKPTPPTPTPTPTP 391
Score = 38.5 bits (88), Expect = 0.003
Identities = 31/86 (36%), Positives = 33/86 (38%), Gaps = 3/86 (3%)
Frame = +2
Query: 131 PPFKKPSPS---MPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
PP PSPS P P T P PP K P PK PP P+ P T
Sbjct: 128 PPTPTPSPSPKPTPPTPPTPKPTPPTPKPTPPT---PKPTPPTPKPTPP-----TPTPTP 283
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P KA S PK TP P +P
Sbjct: 284 PTPKAGCPPPSPPKP-TPPTPPTPKP 358
Score = 37.7 bits (86), Expect = 0.005
Identities = 26/84 (30%), Positives = 31/84 (35%), Gaps = 1/84 (1%)
Frame = +2
Query: 131 PPFKKPSPSMPAQ-PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP KP+P P P T +P PP PK P PPK +P PP P
Sbjct: 218 PPTPKPTPPTPKPTPPTPTPTPP-----TPKAGCPPPSPPKPTPPTPPTPKPTPPTPTPT 382
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ PK P P + P
Sbjct: 383 PTPPTP--PTPKAGCPPPSPPSPP 448
>BU927072
Length = 314
Score = 38.9 bits (89), Expect = 0.002
Identities = 28/74 (37%), Positives = 34/74 (45%)
Frame = +2
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
PS S A PK+ SP PPQ PKA PP +S +A PP AP + +
Sbjct: 56 PSASPAATPKS-SPSPPQAAS-------PKASPPTTS---SAPPPTTPPPPAPAPVSKTA 202
Query: 196 HVSAPKKVTPSKRP 209
S+P TPS P
Sbjct: 203 PASSPPSPTPSSGP 244
Score = 27.3 bits (59), Expect = 7.3
Identities = 20/66 (30%), Positives = 26/66 (39%), Gaps = 9/66 (13%)
Frame = +2
Query: 132 PFKKPSPSMPAQPK----TDSPKPPQIKGCQP-----KNVLPKADPPKSSPSNAAEAIRA 182
P PSP A PK T S PP P P + PP +PS+ A +
Sbjct: 80 PKSSPSPPQAASPKASPPTTSSAPPPTTPPPPAPAPVSKTAPASSPPSPTPSSGPTA-SS 256
Query: 183 PPNTAP 188
P + +P
Sbjct: 257 PSSDSP 274
>TC228693 similar to UP|Q8S9A6 (Q8S9A6) Glucosyltransferase-3, partial (59%)
Length = 907
Score = 38.5 bits (88), Expect = 0.003
Identities = 27/86 (31%), Positives = 35/86 (40%), Gaps = 4/86 (4%)
Frame = +1
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT--- 186
+PP K P P P P T PP P P PP SPS A+ +PP +
Sbjct: 193 HPPTKTPPPPPPPSPAT----PP------PNTSPPSPPPPPPSPSTASPRSPSPPFSLPW 342
Query: 187 -APMKKALSQHVSAPKKVTPSKRPAT 211
+P A ++ TPS +P T
Sbjct: 343 PSPSSSAAPPATTSVASSTPSPKPQT 420
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.133 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,358,797
Number of Sequences: 63676
Number of extensions: 249335
Number of successful extensions: 5562
Number of sequences better than 10.0: 720
Number of HSP's better than 10.0 without gapping: 2995
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4143
length of query: 267
length of database: 12,639,632
effective HSP length: 96
effective length of query: 171
effective length of database: 6,526,736
effective search space: 1116071856
effective search space used: 1116071856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0091b.13


