

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
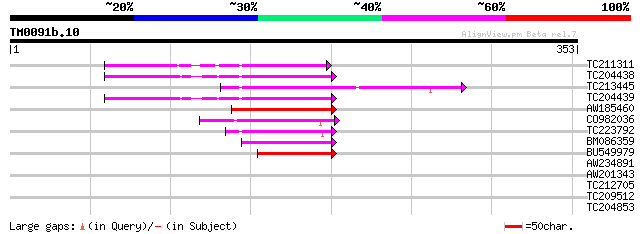
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 60 1e-09
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 60 2e-09
TC213445 58 7e-09
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 57 1e-08
AW185460 49 3e-06
CO982036 46 3e-05
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 44 1e-04
BM086359 42 4e-04
BU549979 42 5e-04
AW234891 39 0.005
AW201343 29 3.7
TC212705 28 6.3
TC209512 similar to UP|Q6SF22 (Q6SF22) Tetracycline resistance p... 28 6.3
TC204853 28 8.2
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 60.5 bits (145), Expect = 1e-09
Identities = 50/142 (35%), Positives = 70/142 (49%), Gaps = 1/142 (0%)
Frame = +3
Query: 60 E*NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*E 119
E F L L I +F++Q Y LK F MD++ P+ T + RS ++KD E
Sbjct: 600 ELKFLLGLQIIQKVYGIFIHQEKYTKSHLKRFRMDEAKPMATPMH-RSTIIDKD-----E 761
Query: 120 KNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLK 178
K E Y I +L YL S +R DI F + L AR+ S P H K++ RYL
Sbjct: 762 KGNHTS*KE--YSGMIDSLSYLTS-SRPDIVFVVCLCARFQSYPKISHVTAVKRILRYLV 932
Query: 179 GALDMSLFFPFMSKLNLIGYAD 200
G + L+F S+ +L+GY D
Sbjct: 933 GTTNHCLWFKKRSEFDLLGYCD 998
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 59.7 bits (143), Expect = 2e-09
Identities = 42/145 (28%), Positives = 75/145 (50%), Gaps = 1/145 (0%)
Frame = +1
Query: 60 E*NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*E 119
E + L L ++ ++D +F+ Q Y ++K F M+ + T L ++KD
Sbjct: 3880 ELTYFLGLQVKQMEDSIFLSQSKYAKNIVKKFGMENASHKRTPAPTH-LKLSKD------ 4038
Query: 120 KNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLK 178
+ +D + Y S I +LLYL + +R DI++A+ + ARY ++P H N K++ +Y+
Sbjct: 4039 EAGTSVDQSL-YRSMIGSLLYLTA-SRPDITYAVGVCARYQANPKISHLNQVKRILKYVN 4212
Query: 179 GALDMSLFFPFMSKLNLIGYADAGY 203
G D + + S L+GY DA +
Sbjct: 4213 GTSDYGIMYCHCSDSMLVGYCDADW 4287
>TC213445
Length = 705
Score = 57.8 bits (138), Expect = 7e-09
Identities = 49/157 (31%), Positives = 82/157 (52%), Gaps = 4/157 (2%)
Frame = +2
Query: 132 LSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFM 190
L+ I++ LYL++ +R I F++ + RY ++P + H + K++ RYL G +++ L++P
Sbjct: 191 LAMIESFLYLST-SRPHIMFSVCMCVRYQANPKESHLSVIKRIMRYLLGIINLGLWYPKN 367
Query: 191 SKLNLIGYADAGYLSKYLKQVTCLHVEEQPFHGDL*KKL**PPQLTLQEY*HYMRQDENT 250
S NL+GY+DA + K L + L V +HG + K+ QL Q +
Sbjct: 368 SSYNLVGYSDAKLIEKVLVTLVILLV-LL*YHGIVKSKIVLSYQLQKQNIFLLEVIMHKS 544
Query: 251 FG*DS*FNAY---LILVICPLERCRQQLYMKIILHAL 284
FG*D+ F LI+ + + Q +Y KII L
Sbjct: 545 FG*DNNFLTMV*NLIIYLSDVTIQVQLIYPKIIFCTL 655
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 57.4 bits (137), Expect = 1e-08
Identities = 41/145 (28%), Positives = 74/145 (50%), Gaps = 1/145 (0%)
Frame = +1
Query: 60 E*NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*E 119
E + L L ++ ++D +F+ Q Y ++K F M+ + T L ++KD
Sbjct: 3877 ELTYFLGLQVKQMEDSIFLSQSRYAKNIVKKFGMENASHKRTPAPTH-LKLSKD------ 4035
Query: 120 KNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLK 178
+ +D + Y S I +LLYL + +R DI++A+ + ARY ++P H K++ +Y+
Sbjct: 4036 EAGTSVDQSL-YRSMIGSLLYLTA-SRPDITYAVGVCARYQANPKISHLTQVKRILKYVN 4209
Query: 179 GALDMSLFFPFMSKLNLIGYADAGY 203
G D + + S L+GY DA +
Sbjct: 4210 GTSDYGIMYCHCSNPMLVGYCDADW 4284
>AW185460
Length = 411
Score = 48.9 bits (115), Expect = 3e-06
Identities = 25/67 (37%), Positives = 42/67 (62%), Gaps = 2/67 (2%)
Frame = +2
Query: 139 LYLASH-TRLDISFAINLLARYSSSPTQRHWN-GKQVFRYLKGALDMSLFFPFMSKLNLI 196
LY+ S TR DI +A +LL+R+ SP+Q H+ GK++ RYL+G +++ + L+
Sbjct: 71 LYIKSQATRPDIMYATSLLSRFMQSPSQIHFGAGKRILRYLQGTKAFGIWYTTETNSELL 250
Query: 197 GYADAGY 203
GY D+ +
Sbjct: 251 GYTDSDW 271
>CO982036
Length = 674
Score = 45.8 bits (107), Expect = 3e-05
Identities = 31/91 (34%), Positives = 49/91 (53%), Gaps = 4/91 (4%)
Frame = -2
Query: 119 EKNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYL 177
+ + +L Y S + AL Y + R +ISFA+N + ++ S+P HW K++ RYL
Sbjct: 394 KSDSDLFSGPTFYRSVVGALQY-TTVIRPEISFAVNKVCQFMSNPLDSHWTEVKRILRYL 218
Query: 178 KGALDMSL-FFPFMSK--LNLIGYADAGYLS 205
KG+L L P +S L + G+ DA + S
Sbjct: 217 KGSLSYGL*LKPAISSQPLPIRGFCDADWAS 125
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 43.5 bits (101), Expect = 1e-04
Identities = 25/73 (34%), Positives = 43/73 (58%), Gaps = 4/73 (5%)
Frame = +1
Query: 135 IKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKL 193
I +L YL + +R +I FA++L++R+ P H K+V R +KG + + FPF +K
Sbjct: 34 IGSLRYLCN-SRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVLFPFKAKS 210
Query: 194 ---NLIGYADAGY 203
+L+GY D+ +
Sbjct: 211 GKPDLLGYTDSDW 249
>BM086359
Length = 427
Score = 42.0 bits (97), Expect = 4e-04
Identities = 20/60 (33%), Positives = 33/60 (54%), Gaps = 1/60 (1%)
Frame = +1
Query: 145 TRLDISFAINLLARYSSSPTQRHWNGK-QVFRYLKGALDMSLFFPFMSKLNLIGYADAGY 203
TRLDI+FA+ +L+++ PT WN ++ RY+K A L + ++ Y DA +
Sbjct: 214 TRLDITFAVGVLSQFLKDPTDSQWNATIRILRYIKNAPGPGLLYEDKGNGKVVCYFDADW 393
>BU549979
Length = 615
Score = 41.6 bits (96), Expect = 5e-04
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Frame = -1
Query: 155 LLARYSSSPTQRHWN-GKQVFRYLKGALDMSLFFPFMSKLNLIGYADAGY 203
+L RY S+P HW K+V RYL+G D L + + L +IGY+D+ +
Sbjct: 615 VLGRYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYKQTNCLEVIGYSDSDF 466
>AW234891
Length = 408
Score = 38.5 bits (88), Expect = 0.005
Identities = 25/70 (35%), Positives = 35/70 (49%)
Frame = +3
Query: 128 EVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNGKQVFRYLKGALDMSLFF 187
E+ L ++ LLYL + TR DI FA L Y S K+ +YLKG + F
Sbjct: 57 EISKLDLVRHLLYLTT-TRPDICFATQLSHSYHFSTCCITKQPKKFLKYLKGNSGKRILF 233
Query: 188 PFMSKLNLIG 197
P S L+++G
Sbjct: 234 PHTSVLHILG 263
>AW201343
Length = 396
Score = 28.9 bits (63), Expect = 3.7
Identities = 22/70 (31%), Positives = 32/70 (45%), Gaps = 17/70 (24%)
Frame = -3
Query: 168 WNGKQVFRYLKGALDMSLFFPFMSKL-----NLIGYAD------------AGYLSKYLKQ 210
W K V Y+KG L+ + FP SKL LI Y+D +GY+ K+L+
Sbjct: 298 WQPKTVLTYIKGILEYGVLFP--SKLLQQGNELIVYSDFDSCGKSNKNNTSGYILKFLEA 125
Query: 211 VTCLHVEEQP 220
++QP
Sbjct: 124 PISWCSKKQP 95
>TC212705
Length = 616
Score = 28.1 bits (61), Expect = 6.3
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = -2
Query: 55 R*KS*E*NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYM 93
R +*E +FCL+ SI + D Y+ +YI L F+M
Sbjct: 486 RLSN*EKHFCLSFSIHFCSDLNKKYKTSYILPKLSFFFM 370
>TC209512 similar to UP|Q6SF22 (Q6SF22) Tetracycline resistance protein,
partial (5%)
Length = 1375
Score = 28.1 bits (61), Expect = 6.3
Identities = 16/53 (30%), Positives = 26/53 (48%), Gaps = 5/53 (9%)
Frame = -3
Query: 173 VFRYLKGALDMSLFFPFMSKLNLIGYADA-----GYLSKYLKQVTCLHVEEQP 220
VF A+ +SL PF+ L + Y+ GYL + L + CL++ + P
Sbjct: 1364 VFNIADEAIALSLSHPFLILLLIFSYSQFYKEKDGYLEQQLLRQGCLYIHDWP 1206
>TC204853
Length = 1087
Score = 27.7 bits (60), Expect = 8.2
Identities = 11/35 (31%), Positives = 19/35 (53%)
Frame = -1
Query: 185 LFFPFMSKLNLIGYADAGYLSKYLKQVTCLHVEEQ 219
+ +PF+ ++L GY SK++ + C H EQ
Sbjct: 700 IIYPFIIVIDLPHIIHHGYFSKFITKSNCSHFTEQ 596
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,705,184
Number of Sequences: 63676
Number of extensions: 186673
Number of successful extensions: 2104
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2095
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2103
length of query: 353
length of database: 12,639,632
effective HSP length: 98
effective length of query: 255
effective length of database: 6,399,384
effective search space: 1631842920
effective search space used: 1631842920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0091b.10


