

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.6
(242 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
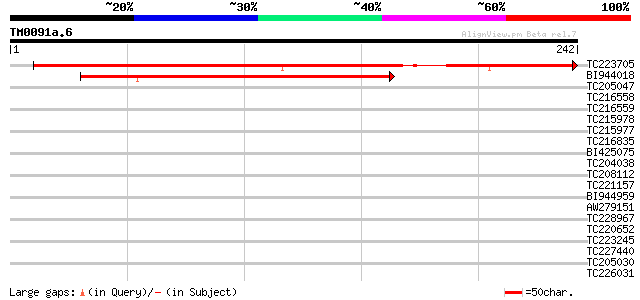
Score E
Sequences producing significant alignments: (bits) Value
TC223705 330 5e-91
BI944018 268 1e-72
TC205047 weakly similar to UP|Q6QUQ2 (Q6QUQ2) Protein disulfide ... 40 0.001
TC216558 weakly similar to UP|O49574 (O49574) Predicted protein,... 39 0.003
TC216559 similar to UP|O49574 (O49574) Predicted protein, partia... 39 0.003
TC215978 weakly similar to UP|Q8S1K2 (Q8S1K2) PDI-like protein, ... 38 0.004
TC215977 weakly similar to UP|Q6QUQ2 (Q6QUQ2) Protein disulfide ... 38 0.004
TC216835 weakly similar to UP|Q9LPJ0 (Q9LPJ0) F6N18.12, partial ... 30 0.98
BI425075 30 1.3
TC204038 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydrat... 30 1.3
TC208112 weakly similar to GB|AAM98099.1|22655016|AY139793 At1g0... 24 1.6
TC221157 similar to UP|Q94EI9 (Q94EI9) AT3g14410/MLN21_19, parti... 28 2.9
BI944959 28 3.7
AW279151 28 4.9
TC228967 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-li... 27 6.4
TC220652 weakly similar to UP|Q93VV0 (Q93VV0) AT3g09320/F3L24_19... 27 6.4
TC223245 weakly similar to UP|Q7VPL4 (Q7VPL4) SufI protein, part... 27 6.4
TC227440 UP|Q8LKV1 (Q8LKV1) GAGA-binding protein, complete 27 6.4
TC205030 UP|ACCD_SOYBN (P49158) Acetyl-coenzyme A carboxylase ca... 27 6.4
TC226031 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, ... 27 6.4
>TC223705
Length = 918
Score = 330 bits (845), Expect = 5e-91
Identities = 155/238 (65%), Positives = 180/238 (75%), Gaps = 6/238 (2%)
Frame = +3
Query: 11 HPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQ 70
H QHKL F+++ PFKCDGCKE+GIGSRYKCSICD+DLH C+IPSP+L HPFY KCSF
Sbjct: 3 HKQHKLAFDYSESPFKCDGCKELGIGSRYKCSICDFDLHTHCSIPSPSLFHPFYPKCSFH 182
Query: 71 FMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL--DDGEVKLYLYRKV 128
F+S PPG+ RYCNACEK V GF+YHC SCGFDLHPCCAKLP VL D+ V+L+LYRKV
Sbjct: 183 FLSQPPGDTSRYCNACEKPVKGFLYHCFSCGFDLHPCCAKLPTVLGSDEHGVRLFLYRKV 362
Query: 129 SSPCHRCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDHVYVAGGKGRNKVVETSVPGL 188
SS CHRCGRKG WSYRS CK YNLHVACVREMVVE+W GG GL
Sbjct: 363 SSACHRCGRKGSGWSYRSSCKRYNLHVACVREMVVESWHE----GG------------GL 494
Query: 189 KNTLYAAHNSRRSKG----KVKKCCEIAGLAVQFVISAVLGDPTALIAGVVGSLMSRA 242
N + + + R KG +V++CCE+AG+AVQ +SAVLGDPT LIAG++GSLMSRA
Sbjct: 495 MNVVESGCHKGRGKGSGGSRVRRCCEVAGMAVQVAVSAVLGDPTVLIAGIMGSLMSRA 668
>BI944018
Length = 409
Score = 268 bits (686), Expect = 1e-72
Identities = 116/135 (85%), Positives = 129/135 (94%), Gaps = 1/135 (0%)
Frame = +3
Query: 31 KEIGIGSRYKCSICDYDLHMQCA-IPSPTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKD 89
+E+GIGSRYKCSICD+DLHM CA I SPTL HPFYTKC+FQFMS PPG+ PRYCNACEKD
Sbjct: 3 REVGIGSRYKCSICDFDLHMHCAMITSPTLHHPFYTKCNFQFMSKPPGDTPRYCNACEKD 182
Query: 90 VNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCK 149
V+GF+YHCK+CGFDLHPCCAKLP++LDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCK
Sbjct: 183 VSGFLYHCKACGFDLHPCCAKLPLLLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCK 362
Query: 150 SYNLHVACVREMVVE 164
+YNLHVACVR+M+VE
Sbjct: 363 NYNLHVACVRDMLVE 407
Score = 39.7 bits (91), Expect = 0.001
Identities = 25/88 (28%), Positives = 39/88 (43%), Gaps = 1/88 (1%)
Frame = +3
Query: 24 PFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKC-SFQFMSSPPGNIPRY 82
P C+ C++ G Y C C +DLH CA P L+ K ++ +SSP
Sbjct: 153 PRYCNACEKDVSGFLYHCKACGFDLHPCCA-KLPLLLDDGEVKLYLYRKVSSP------- 308
Query: 83 CNACEKDVNGFVYHCKSCGFDLHPCCAK 110
C+ C + + Y K ++LH C +
Sbjct: 309 CHRCGRKGRSWSYRSKCKNYNLHVACVR 392
>TC205047 weakly similar to UP|Q6QUQ2 (Q6QUQ2) Protein disulfide isomerase,
partial (91%)
Length = 2014
Score = 39.7 bits (91), Expect = 0.001
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +3
Query: 82 YCNACEKDVNGFVYHCKSCGFDLHPCCA 109
YC+AC ++ + + Y+C C FDLHP CA
Sbjct: 1590 YCDACNEEGHIWSYYCGDCDFDLHPKCA 1673
Score = 36.6 bits (83), Expect = 0.010
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +3
Query: 25 FKCDGCKEIGIGSRYKCSICDYDLHMQCAI 54
+ CD C E G Y C CD+DLH +CA+
Sbjct: 1587 YYCDACNEEGHIWSYYCGDCDFDLHPKCAL 1676
Score = 28.5 bits (62), Expect = 2.9
Identities = 14/42 (33%), Positives = 19/42 (44%)
Frame = +3
Query: 119 EVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACVRE 160
E +L L R+ C C +G WSY ++LH C E
Sbjct: 1554 EHELVLTRRRVYYCDACNEEGHIWSYYCGDCDFDLHPKCALE 1679
>TC216558 weakly similar to UP|O49574 (O49574) Predicted protein, partial
(64%)
Length = 743
Score = 38.5 bits (88), Expect = 0.003
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = +3
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQC 52
+H L+ + A + CD CK+ G + C +CDYDLH C
Sbjct: 330 EHVLKLDM-AKAYVCDSCKKQGKFWTFSCDVCDYDLHPSC 446
Score = 32.7 bits (73), Expect = 0.15
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +3
Query: 83 CNACEKDVNGFVYHCKSCGFDLHPCC 108
C++C+K + + C C +DLHP C
Sbjct: 369 CDSCKKQGKFWTFSCDVCDYDLHPSC 446
Score = 28.5 bits (62), Expect(2) = 0.13
Identities = 8/30 (26%), Positives = 18/30 (59%)
Frame = +3
Query: 132 CHRCGRKGRSWSYRSKCKSYNLHVACVREM 161
C C ++G+ W++ Y+LH +C+ ++
Sbjct: 369 CDSCKKQGKFWTFSCDVCDYDLHPSCLEKV 458
Score = 23.1 bits (48), Expect(2) = 0.13
Identities = 12/29 (41%), Positives = 15/29 (51%)
Frame = +2
Query: 84 NACEKDVNGFVYHCKSCGFDLHPCCAKLP 112
N CE VNG + C+S F +H K P
Sbjct: 188 N*CEWKVNGLLIWCRS--FPIHRIADKRP 268
>TC216559 similar to UP|O49574 (O49574) Predicted protein, partial (85%)
Length = 866
Score = 38.5 bits (88), Expect = 0.003
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = +2
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQC 52
+H L+ + A + CD CK+ G + C +CDYDLH C
Sbjct: 587 EHVLKLDM-AKAYVCDSCKKQGKFWTFSCDVCDYDLHPSC 703
Score = 32.7 bits (73), Expect = 0.15
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +2
Query: 83 CNACEKDVNGFVYHCKSCGFDLHPCC 108
C++C+K + + C C +DLHP C
Sbjct: 626 CDSCKKQGKFWTFSCDVCDYDLHPSC 703
Score = 28.5 bits (62), Expect(2) = 0.10
Identities = 8/30 (26%), Positives = 18/30 (59%)
Frame = +2
Query: 132 CHRCGRKGRSWSYRSKCKSYNLHVACVREM 161
C C ++G+ W++ Y+LH +C+ ++
Sbjct: 626 CDSCKKQGKFWTFSCDVCDYDLHPSCLEKV 715
Score = 23.5 bits (49), Expect(2) = 0.10
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = +1
Query: 86 CEKDVNGFVYHCKSCGFDLHPCCAKLP 112
CE +VNG + C+S F +H K P
Sbjct: 451 CEWEVNGLLIRCRS--FPVHRITDKRP 525
>TC215978 weakly similar to UP|Q8S1K2 (Q8S1K2) PDI-like protein, partial (34%)
Length = 1554
Score = 38.1 bits (87), Expect = 0.004
Identities = 15/54 (27%), Positives = 26/54 (47%)
Frame = +3
Query: 57 PTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAK 110
P LV+ + +S G P C C++ + + Y C CG+++HP C +
Sbjct: 1095 PALVYHQGHRHDLNLVSDGNGGGPFICCVCDEQGSSWAYQCLQCGYEVHPKCVR 1256
Score = 31.2 bits (69), Expect = 0.44
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = +3
Query: 132 CHRCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDHVYV 171
C C +G SW+Y+ Y +H CVR VE D+V V
Sbjct: 1173 CCVCDEQGSSWAYQCLQCGYEVHPKCVR--TVERDDNVLV 1286
Score = 31.2 bits (69), Expect = 0.44
Identities = 12/29 (41%), Positives = 16/29 (54%)
Frame = +3
Query: 24 PFKCDGCKEIGIGSRYKCSICDYDLHMQC 52
PF C C E G Y+C C Y++H +C
Sbjct: 1164 PFICCVCDEQGSSWAYQCLQCGYEVHPKC 1250
>TC215977 weakly similar to UP|Q6QUQ2 (Q6QUQ2) Protein disulfide isomerase,
partial (8%)
Length = 1237
Score = 38.1 bits (87), Expect = 0.004
Identities = 15/54 (27%), Positives = 26/54 (47%)
Frame = +1
Query: 57 PTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAK 110
P LV+ + +S G P C C++ + + Y C CG+++HP C +
Sbjct: 691 PALVYHEGHRHDLNLVSDGNGGGPFICCVCDEQGSSWAYQCLQCGYEVHPKCVR 852
Score = 31.2 bits (69), Expect = 0.44
Identities = 12/29 (41%), Positives = 16/29 (54%)
Frame = +1
Query: 24 PFKCDGCKEIGIGSRYKCSICDYDLHMQC 52
PF C C E G Y+C C Y++H +C
Sbjct: 760 PFICCVCDEQGSSWAYQCLQCGYEVHPKC 846
Score = 29.3 bits (64), Expect = 1.7
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +1
Query: 132 CHRCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDH 168
C C +G SW+Y+ Y +H CVR VE D+
Sbjct: 769 CCVCDEQGSSWAYQCLQCGYEVHPKCVR--TVERHDN 873
>TC216835 weakly similar to UP|Q9LPJ0 (Q9LPJ0) F6N18.12, partial (47%)
Length = 776
Score = 30.0 bits (66), Expect = 0.98
Identities = 20/61 (32%), Positives = 30/61 (48%), Gaps = 9/61 (14%)
Frame = +2
Query: 31 KEIGIGSRYKCSICDYDLHMQCAIPS-PTLVHP-FYTKCSFQFMSS-------PPGNIPR 81
K G+G R+ S Y M+CA PS P L HP + + SF +++ PP +P
Sbjct: 542 KARGLGGRFAVSALVYLYRMRCAFPSVPGLSHPKEFNR*SFSLLTTLQNLLNYPPSFLPS 721
Query: 82 Y 82
+
Sbjct: 722 F 724
>BI425075
Length = 408
Score = 29.6 bits (65), Expect = 1.3
Identities = 18/69 (26%), Positives = 28/69 (40%), Gaps = 2/69 (2%)
Frame = +1
Query: 44 CDYD-LHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKS-CG 101
C YD +H+QC C S IP+YC+ C++ G +H G
Sbjct: 61 CKYDTVHLQCQA------------CGGMMPSRTGFGIPQYCSGCDRSFCGAYWHALGVTG 204
Query: 102 FDLHPCCAK 110
+P C++
Sbjct: 205 NGSYPVCSQ 231
>TC204038 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratase-like
protein, partial (37%)
Length = 1102
Score = 29.6 bits (65), Expect = 1.3
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +1
Query: 30 CKEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
C+++ G ++C +C DL+ C P P L
Sbjct: 715 CEDLYTGGTWRCQVC*RDLYRSCQRPRPGL 804
>TC208112 weakly similar to GB|AAM98099.1|22655016|AY139793 At1g05070/T7A14_6
{Arabidopsis thaliana;} , partial (29%)
Length = 827
Score = 23.9 bits (50), Expect(2) = 1.6
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = -1
Query: 54 IPSPTLVHPFYTKCSFQFMSSPPGNIPRY 82
+PSP L+H F +C + P GN R+
Sbjct: 548 LPSPLLLHKFPPQCCTFLL--PSGNAKRF 468
Score = 23.9 bits (50), Expect(2) = 1.6
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = -3
Query: 8 HFSHPQHKLRFEHNA 22
H +HP + RF HNA
Sbjct: 624 HSTHPYSRARFSHNA 580
>TC221157 similar to UP|Q94EI9 (Q94EI9) AT3g14410/MLN21_19, partial (60%)
Length = 661
Score = 28.5 bits (62), Expect = 2.9
Identities = 10/37 (27%), Positives = 19/37 (51%)
Frame = -3
Query: 1 MKYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGS 37
+ + ++ S+P+HK + HN C + I +GS
Sbjct: 428 LPFGKLKSCSYPKHKNSYRHNCLQHLCKSNRNIQVGS 318
>BI944959
Length = 304
Score = 28.1 bits (61), Expect = 3.7
Identities = 15/44 (34%), Positives = 22/44 (49%), Gaps = 2/44 (4%)
Frame = -2
Query: 92 GFVY--HCKSCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSPCH 133
GF Y + +SC A P+ + Y++RK+SSPCH
Sbjct: 273 GFSYTANTESCSL------ASQPLT*TSSHIIFYVFRKMSSPCH 160
>AW279151
Length = 263
Score = 27.7 bits (60), Expect = 4.9
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = -3
Query: 60 VHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFV 94
++PFY C F F N+ +YC A + G V
Sbjct: 153 MYPFYLHCFFSFYPYN*KNVNQYCMALPPQLTGTV 49
>TC228967 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-like protease
{Arabidopsis thaliana;} , partial (21%)
Length = 1106
Score = 27.3 bits (59), Expect = 6.4
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = -2
Query: 132 CHRCGRKGRSWSYRSKCKSY 151
CH C +GRS+S + C+ Y
Sbjct: 358 CHGCSHQGRSYSSSAPCRVY 299
>TC220652 weakly similar to UP|Q93VV0 (Q93VV0) AT3g09320/F3L24_19, partial
(13%)
Length = 1156
Score = 27.3 bits (59), Expect = 6.4
Identities = 14/52 (26%), Positives = 24/52 (45%), Gaps = 14/52 (26%)
Frame = +1
Query: 63 FYTKCSFQFMSSPP----GNIPR----------YCNACEKDVNGFVYHCKSC 100
F++ +F+ +PP G+ P YC+ C K + +HC+SC
Sbjct: 457 FFSFAAFRCAGTPPNILWGSYPAVGKDDLENYTYCHYCSKPKSPRAHHCRSC 612
>TC223245 weakly similar to UP|Q7VPL4 (Q7VPL4) SufI protein, partial (4%)
Length = 404
Score = 27.3 bits (59), Expect = 6.4
Identities = 24/89 (26%), Positives = 35/89 (38%), Gaps = 3/89 (3%)
Frame = +1
Query: 99 SCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSPCHRCGR---KGRSWSYRSKCKSYNLHV 155
S G +H ++P +D V+L+ V S C G K +S + N
Sbjct: 133 SDGLQIHHQSHQIPFNVDPRTVQLFKVSPVLSVCLVEGSDVGKKTLYSRGVTIQVRNDEE 312
Query: 156 ACVREMVVENWDHVYVAGGKGRNKVVETS 184
+ VV+ W A G GRN + TS
Sbjct: 313 SAAFHCVVQQWKKEVNAQGNGRNGTITTS 399
>TC227440 UP|Q8LKV1 (Q8LKV1) GAGA-binding protein, complete
Length = 1305
Score = 27.3 bits (59), Expect = 6.4
Identities = 9/34 (26%), Positives = 19/34 (55%)
Frame = -1
Query: 43 ICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPP 76
+C Y +H+ ++ +P + H F+ CS ++ P
Sbjct: 507 VCRYHIHIFVSLTNPRIPHVFHRVCSLGHTTNVP 406
>TC205030 UP|ACCD_SOYBN (P49158) Acetyl-coenzyme A carboxylase carboxyl
transferase subunit beta (ACCASE beta chain) , complete
Length = 4503
Score = 27.3 bits (59), Expect = 6.4
Identities = 18/63 (28%), Positives = 25/63 (39%)
Frame = +1
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEV 120
+PFY K SF F SS +I CN GF + C K + + +V
Sbjct: 3703 YPFYDKLSFHFCSSSRPSISGNCN----------------GFFISSCSKKQDFLDTENKV 3834
Query: 121 KLY 123
K +
Sbjct: 3835 KFW 3843
>TC226031 similar to UP|YIPL_SOLTU (P59234) Yippee-like protein, partial
(85%)
Length = 725
Score = 27.3 bits (59), Expect = 6.4
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = -3
Query: 49 HMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRY 82
H Q I +P L+H + KC Q+ S P P +
Sbjct: 423 HEQSHISNPQLIHSPHRKCQPQYASQWPSVCPHF 322
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.138 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,836,701
Number of Sequences: 63676
Number of extensions: 314539
Number of successful extensions: 2108
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 2073
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2101
length of query: 242
length of database: 12,639,632
effective HSP length: 95
effective length of query: 147
effective length of database: 6,590,412
effective search space: 968790564
effective search space used: 968790564
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0091a.6


