

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.5
(140 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
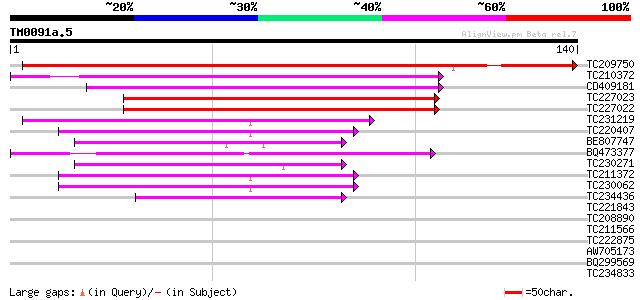
Score E
Sequences producing significant alignments: (bits) Value
TC209750 189 5e-49
TC210372 65 1e-11
CD409181 58 1e-09
TC227023 54 2e-08
TC227022 weakly similar to UP|Q8LQE3 (Q8LQE3) Cytokinesis regula... 50 5e-07
TC231219 48 2e-06
TC220407 similar to UP|Q6NLC8 (Q6NLC8) At1g66480, partial (31%) 44 3e-05
BE807747 similar to GP|20161304|dbj B1131G08.9 {Oryza sativa (ja... 43 4e-05
BQ473377 43 6e-05
TC230271 42 1e-04
TC211372 41 2e-04
TC230062 40 3e-04
TC234436 39 6e-04
TC221843 36 0.007
TC208890 similar to GB|AAO39927.1|28372890|BT003699 At3g10120 {A... 36 0.007
TC211566 weakly similar to UP|Q6NL13 (Q6NL13) At5g37840, partial... 35 0.009
TC222875 similar to GB|AAD14477.1|4249380|T2K10 ESTs gb|Z37637, ... 33 0.044
AW705173 32 0.097
BQ299569 32 0.097
TC234833 30 0.48
>TC209750
Length = 935
Score = 189 bits (479), Expect = 5e-49
Identities = 98/142 (69%), Positives = 111/142 (78%), Gaps = 5/142 (3%)
Frame = +3
Query: 4 CASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSP 63
CA TQYA KGGNH QSTVNIVH DGKLQQLKEP+KAW VLS+NPN ++C SESMYVGSP
Sbjct: 75 CAPTQYASKGGNHSWQSTVNIVHIDGKLQQLKEPIKAWKVLSENPNCYICCSESMYVGSP 254
Query: 64 MTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALA-----HSSSVFEPSS 118
M P+ P+EELQL IYFLVP KSR+PLSL+DL ALAIKANAALA +S + +P+S
Sbjct: 255 MPPLAPSEELQLGLIYFLVPIPKSRIPLSLQDLGALAIKANAALAAATPNYSYPILKPNS 434
Query: 119 SVPQINFKTHPVHSNVSLGYSH 140
Q F+THP SNVSL YSH
Sbjct: 435 ---QKKFRTHPTLSNVSLVYSH 491
>TC210372
Length = 631
Score = 65.1 bits (157), Expect = 1e-11
Identities = 36/107 (33%), Positives = 60/107 (55%)
Frame = +1
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG C+S+ +T +V DG+LQ+ PVK +L + P F+C+S+ M
Sbjct: 46 MGNCSSSDST-------QVATAKLVLQDGRLQEFSYPVKVSFLLQKYPACFICNSDEMEF 204
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
G ++ + ++ LQ +YF +P S+ R PL ++ ALA+KA++AL
Sbjct: 205 GDVVSAIDEDQVLQPGQLYFALPLSRLRHPLQPHEMAALAVKASSAL 345
>CD409181
Length = 600
Score = 58.2 bits (139), Expect = 1e-09
Identities = 30/88 (34%), Positives = 51/88 (57%)
Frame = -1
Query: 20 STVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIY 79
+T ++ DG+LQ+ PVK +L P F+C+S+ M ++P+ ++ LQ +Y
Sbjct: 558 ATAKLLVQDGRLQEFSYPVKVSFLLHNYPACFICNSDEMEFQDVVSPIHEDQVLQPGQLY 379
Query: 80 FLVPRSKSRLPLSLEDLCALAIKANAAL 107
F +P S R L ++ ALA+KA++AL
Sbjct: 378 FALPLSLLRHSLQPHEMAALAVKASSAL 295
>TC227023
Length = 875
Score = 53.9 bits (128), Expect = 2e-08
Identities = 24/78 (30%), Positives = 48/78 (60%)
Frame = +2
Query: 29 GKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSR 88
G+++Q KE VKA ++ ++PN+FL +S S+++G + + +EEL+ ++Y P +
Sbjct: 185 GEVKQFKEIVKAAELMLEHPNYFLVNSRSLHIGRRFSALGADEELEFGNVYIFFPMRRVN 364
Query: 89 LPLSLEDLCALAIKANAA 106
++ D+ L + AN+A
Sbjct: 365 SLVTAPDMAVLFLAANSA 418
>TC227022 weakly similar to UP|Q8LQE3 (Q8LQE3) Cytokinesis regulating
protein-like, partial (9%)
Length = 826
Score = 49.7 bits (117), Expect = 5e-07
Identities = 22/78 (28%), Positives = 48/78 (61%)
Frame = +3
Query: 29 GKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSR 88
G+++Q +E VKA ++ ++P++FL +S S+++G + + +EEL+ ++Y P +
Sbjct: 228 GEVKQFREIVKAAELMLEHPSYFLVNSRSLHIGRRFSALGADEELESGNVYIFFPMRRVN 407
Query: 89 LPLSLEDLCALAIKANAA 106
++ D+ L + AN+A
Sbjct: 408 SVVTPTDMAVLFLAANSA 461
>TC231219
Length = 1165
Score = 47.8 bits (112), Expect = 2e-06
Identities = 28/88 (31%), Positives = 43/88 (48%), Gaps = 1/88 (1%)
Frame = +1
Query: 4 CASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGS 62
C+ A G G + T ++ DG+ +LK PVK VL +P L SE++ + G
Sbjct: 133 CSIGVLAIMGNTFGAKKTTKVMKIDGETFKLKTPVKVGEVLKDHPGLVLLDSEAVKHYGV 312
Query: 63 PMTPVVPNEELQLNHIYFLVPRSKSRLP 90
P+ +++LQ +YFLV K P
Sbjct: 313 RAKPLEAHKDLQPKRLYFLVELPKETTP 396
>TC220407 similar to UP|Q6NLC8 (Q6NLC8) At1g66480, partial (31%)
Length = 846
Score = 43.5 bits (101), Expect = 3e-05
Identities = 25/75 (33%), Positives = 37/75 (49%), Gaps = 1/75 (1%)
Frame = +3
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G G ++ DG+ +LK P +A V+ P H L SE++ + G P+ P +
Sbjct: 87 GNTMGRSKKTKVMKVDGETFKLKTPARANDVVKDYPGHVLLDSEAVKHFGLRAKPLEPYQ 266
Query: 72 ELQLNHIYFLVPRSK 86
EL+ IYFLV K
Sbjct: 267 ELKPKKIYFLVELPK 311
>BE807747 similar to GP|20161304|dbj B1131G08.9 {Oryza sativa (japonica
cultivar-group)}, partial (48%)
Length = 407
Score = 43.1 bits (100), Expect = 4e-05
Identities = 26/75 (34%), Positives = 39/75 (51%), Gaps = 8/75 (10%)
Frame = +1
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFL---CSSESMYVG-----SPMTPVV 68
G + IVH +G+++++ VKA V+ +P H L CSS + G + V
Sbjct: 121 GALDVIRIVHSNGRVEEISGTVKASDVMKAHPKHVLKKPCSSPADAAGISGGVHKIVVVP 300
Query: 69 PNEELQLNHIYFLVP 83
P+ ELQ IYFL+P
Sbjct: 301 PDAELQRGKIYFLMP 345
>BQ473377
Length = 453
Score = 42.7 bits (99), Expect = 6e-05
Identities = 27/105 (25%), Positives = 47/105 (44%)
Frame = +1
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG C S + N + +VH +G ++ +P+ V+ HF+C+S + +
Sbjct: 4 MGACFSCNSSSTLKN------IRVVHLNGYVEDFDQPISVRQVIGYPQKHFVCTSTQL-L 162
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANA 105
T + + LQ +YF++P S +S DL LA + A
Sbjct: 163 SPCSTSMNGDTHLQPGQVYFMLPYSVLHADVSPVDLAGLAKRLTA 297
>TC230271
Length = 700
Score = 41.6 bits (96), Expect = 1e-04
Identities = 21/71 (29%), Positives = 36/71 (50%), Gaps = 4/71 (5%)
Frame = +2
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTP----VVPNEE 72
G + IVH +G+++++ +KA ++ +P H L S + P V P+ +
Sbjct: 161 GALDVIRIVHSNGRVEEISGTIKASEIMKAHPKHVLKKPSSPSTQDGVVPKIVVVPPDAD 340
Query: 73 LQLNHIYFLVP 83
LQ IYFL+P
Sbjct: 341 LQRGKIYFLMP 373
>TC211372
Length = 1053
Score = 41.2 bits (95), Expect = 2e-04
Identities = 24/75 (32%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Frame = +3
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G G + T ++ DG+ +LK P+K VL +P L SE++ + G P+ ++
Sbjct: 141 GNALGGKKTTKVMKIDGETFKLKTPIKVCDVLKDHPGLVLLESEAVKHYGIRAKPLEAHK 320
Query: 72 ELQLNHIYFLVPRSK 86
EL +YFLV K
Sbjct: 321 ELMPKRLYFLVELPK 365
>TC230062
Length = 1000
Score = 40.4 bits (93), Expect = 3e-04
Identities = 24/75 (32%), Positives = 37/75 (49%), Gaps = 1/75 (1%)
Frame = +3
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNE 71
G G + T ++ DG+ +LK P+K VL +P L SE++ + G P+ ++
Sbjct: 144 GNALGGKKTTKVMKIDGETFKLKTPIKVCDVLKNHPGLVLLESEAVKHYGIRAKPLEAHK 323
Query: 72 ELQLNHIYFLVPRSK 86
EL YFLV K
Sbjct: 324 ELMPKRFYFLVELPK 368
>TC234436
Length = 589
Score = 39.3 bits (90), Expect = 6e-04
Identities = 19/52 (36%), Positives = 28/52 (53%)
Frame = +2
Query: 32 QQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVP 83
+++ P+ A VL NPNH L S V + + P+ EL+ IYFL+P
Sbjct: 5 EEITRPITAGEVLKANPNHVLSKPTSQGVVRRILILSPDTELKRGSIYFLIP 160
>TC221843
Length = 555
Score = 35.8 bits (81), Expect = 0.007
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 2/84 (2%)
Frame = +3
Query: 12 KGGNHGMQST--VNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVP 69
KG +HG+ V +V +G + +L P+ + S+ P H + S P+
Sbjct: 234 KGLHHGVSENMMVKVVTSNGGIMELFSPITVECITSEFPGHGIFRSRRDMFSEPLPK--- 404
Query: 70 NEELQLNHIYFLVPRSKSRLPLSL 93
NEEL+ +Y+L+P + S SL
Sbjct: 405 NEELRGGKVYYLLPLNPSSSRKSL 476
>TC208890 similar to GB|AAO39927.1|28372890|BT003699 At3g10120 {Arabidopsis
thaliana;} , partial (8%)
Length = 787
Score = 35.8 bits (81), Expect = 0.007
Identities = 23/65 (35%), Positives = 36/65 (55%)
Frame = +2
Query: 19 QSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHI 78
++ V IV DGK+ + K P+K VL Q H + SES+ V + + P+ +L +
Sbjct: 77 ENVVKIVKTDGKVLEYKTPIKVEEVLIQFSGHAV--SESLTV---LRYLEPHTKLLRGQL 241
Query: 79 YFLVP 83
Y+LVP
Sbjct: 242 YYLVP 256
>TC211566 weakly similar to UP|Q6NL13 (Q6NL13) At5g37840, partial (41%)
Length = 632
Score = 35.4 bits (80), Expect = 0.009
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 1/74 (1%)
Frame = +1
Query: 14 GNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNEE 72
GN ++ DG+ +L P +A V+ P H L S ++ G P+ P+ +
Sbjct: 61 GNAMGSKKAKVMKIDGETFKLNTPARANDVVKDYPGHVLLDSHAVKNFGPRAKPLEPDYQ 240
Query: 73 LQLNHIYFLVPRSK 86
L+ IYFLV K
Sbjct: 241 LKPKKIYFLVELPK 282
>TC222875 similar to GB|AAD14477.1|4249380|T2K10 ESTs gb|Z37637, gb|AA042498
and gb|AA042269 come from this gene. {Arabidopsis
thaliana;} , partial (57%)
Length = 563
Score = 33.1 bits (74), Expect = 0.044
Identities = 23/76 (30%), Positives = 34/76 (44%), Gaps = 17/76 (22%)
Frame = +3
Query: 24 IVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTP----------------- 66
I H GK+++L PV A V+ NP H++ S+ + P+ P
Sbjct: 132 IQHPCGKIERLYWPVTASEVMRTNPGHYV----SLIIPLPVPPQEQNQEKKTVRFTRVKL 299
Query: 67 VVPNEELQLNHIYFLV 82
+ PNE L L H Y L+
Sbjct: 300 LRPNETLNLGHAYRLI 347
>AW705173
Length = 411
Score = 32.0 bits (71), Expect = 0.097
Identities = 12/33 (36%), Positives = 21/33 (63%)
Frame = +3
Query: 20 STVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFL 52
+T+ I H +GK+ + P+ A HV+ NP H++
Sbjct: 3 ATLVIQHTNGKVDKFYAPLSATHVMKTNPGHYV 101
>BQ299569
Length = 423
Score = 32.0 bits (71), Expect = 0.097
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 2/84 (2%)
Frame = +1
Query: 12 KGGNHGMQST--VNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVP 69
+G NHG+ V +V +G + +L P+ + ++ P + S P+
Sbjct: 52 RGLNHGVSEDMMVKVVTSNGGIMELFSPITVECITNEFPGQGIFRSRRDMFSEPLPK--- 222
Query: 70 NEELQLNHIYFLVPRSKSRLPLSL 93
NEEL +Y+L+P + S SL
Sbjct: 223 NEELHGGEVYYLLPLNPSSSRKSL 294
>TC234833
Length = 662
Score = 29.6 bits (65), Expect = 0.48
Identities = 9/41 (21%), Positives = 24/41 (57%)
Frame = +3
Query: 43 VLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVP 83
++ + P +C ++S ++G P+ + +++L YF++P
Sbjct: 6 IMXEFPEMMVCHADSFFIGQPIPVLSIDDKLMPGQTYFILP 128
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,544,454
Number of Sequences: 63676
Number of extensions: 120048
Number of successful extensions: 621
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 618
length of query: 140
length of database: 12,639,632
effective HSP length: 88
effective length of query: 52
effective length of database: 7,036,144
effective search space: 365879488
effective search space used: 365879488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0091a.5


