

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089b.6
(647 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
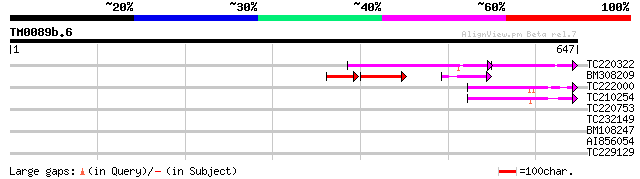
Score E
Sequences producing significant alignments: (bits) Value
TC220322 83 3e-32
BM308209 66 5e-27
TC222000 weakly similar to PIR|H84710|H84710 Mutator-like transp... 60 2e-09
TC210254 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F1... 57 3e-08
TC220753 41 0.001
TC232149 32 0.003
BM108247 weakly similar to PIR|H84710|H847 Mutator-like transpos... 39 0.009
AI856054 35 0.14
TC229129 similar to UP|Q6NM52 (Q6NM52) At4g15770, complete 29 7.5
>TC220322
Length = 1264
Score = 82.8 bits (203), Expect(2) = 3e-32
Identities = 56/169 (33%), Positives = 85/169 (50%), Gaps = 3/169 (1%)
Frame = +2
Query: 386 AKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNAT 445
AK+T + M L+ ++ +A+ +L + P AW K F+ K D + N E FN
Sbjct: 8 AKSTTVAEFEGHMAHLKTINCQAWEYLNKWPKQAWTKAHFSTIPKVDNICNNTCEVFNFR 187
Query: 446 ILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQC 505
IL R KP++TM++ IRSYIM A K+ + +RL+ E + P
Sbjct: 188 ILQYRCKPIITMLEEIRSYIMRTMAARKVKLSGKPGPLCLVQYKRLEKEFHFANQWTPIW 367
Query: 506 CG*DM---FEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAIS 551
CG +M +EV + ++ V+L TC C W+L G+P RH IA I+
Sbjct: 368 CGDNMGLRYEVH--MWGNKVEVNLAGWTCTCGVWQLTGMPCRHAIATIT 508
Score = 74.7 bits (182), Expect(2) = 3e-32
Identities = 38/97 (39%), Positives = 48/97 (49%)
Frame = +1
Query: 551 SHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEYILSPLYRRALGRPKKL 610
SH G KPED H + AY TY+H + P+ G + W T + + P R GRPKK
Sbjct: 508 SHKGGKPEDMCHEWLSIEAYNKTYQHFIEPVQGPQYWAQTQYTHPVPPYKRVQRGRPKKN 687
Query: 611 RMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHNTRS 647
R R ED H +L ++ RCGQ HN RS
Sbjct: 688 RKRSVDEDNVTGH-KLKRKLAEFTCGRCGQTNHNIRS 795
>BM308209
Length = 435
Score = 65.9 bits (159), Expect(3) = 5e-27
Identities = 27/52 (51%), Positives = 40/52 (76%)
Frame = +2
Query: 401 LQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDK 452
++ ++ EAY WL+++ T W K F++YTKCDVLMN L ++FN+TIL AR+K
Sbjct: 122 IKEVNNEAYDWLIKVGTKLWRKHTFSYYTKCDVLMNNLCDAFNSTILCAREK 277
Score = 49.3 bits (116), Expect(3) = 5e-27
Identities = 21/37 (56%), Positives = 28/37 (74%)
Frame = +3
Query: 362 RHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKM 398
RH+Y NFKN F G ++RNLMM AAKA++ + W +KM
Sbjct: 6 RHIYNNFKNKFVGGTLLRNLMMRAAKASFEEAWDEKM 116
Score = 45.1 bits (105), Expect(3) = 5e-27
Identities = 23/58 (39%), Positives = 31/58 (52%)
Frame = +3
Query: 493 WEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAI 550
W ++ HL+ FEV H + D+FIV+L +C N WE+ GI RH AAI
Sbjct: 285 WFATLSGHLV--------FEVSHSMLSDKFIVNLNSWSCS*NLWEITGILCRHACAAI 434
>TC222000 weakly similar to PIR|H84710|H84710 Mutator-like transposase
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (12%)
Length = 617
Score = 60.5 bits (145), Expect = 2e-09
Identities = 43/132 (32%), Positives = 58/132 (43%), Gaps = 7/132 (5%)
Frame = +2
Query: 523 IVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPIS 582
IV++ H+C C W+L GIP H AA+ + F + A++R TY + PI
Sbjct: 17 IVNIGSHSCSCRDWQLNGIPCSHAAAALISCRKDVYAFSQKCFTAASFRDTYAETIHPIP 196
Query: 583 GQKLWPTTN----DEYIL---SPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKF 635
G+ W T D+ IL P RR GRP+K D D +H
Sbjct: 197 GKLEWSKTGNSSMDDNILVVRPPKLRRPPGRPEKRMCVD---DLNREKHTVHC------- 346
Query: 636 SRCGQFGHNTRS 647
SRC Q GH R+
Sbjct: 347 SRCNQTGHYKRT 382
>TC210254 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17A9_9
{Arabidopsis thaliana;} , partial (6%)
Length = 814
Score = 56.6 bits (135), Expect = 3e-08
Identities = 34/129 (26%), Positives = 54/129 (41%), Gaps = 4/129 (3%)
Frame = +1
Query: 523 IVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPIS 582
IVD+ C C W+L G+P H IA G+ P D+ Y+ YR TY + P+
Sbjct: 64 IVDIDNWDCSCKGWQLTGVPCCHAIAVFECVGRSPYDYCSRYFTVENYRLTYAESIHPVP 243
Query: 583 GQKLWPTTNDE----YILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRC 638
P + ++ P +R GRPK ++ + I + + S+C
Sbjct: 244 NVDKPPVQGESTALVMVIPPPTKRPPGRPKMKQVES-----------IDIIKRQLQCSKC 390
Query: 639 GQFGHNTRS 647
GHN ++
Sbjct: 391 KGLGHNRKT 417
>TC220753
Length = 1312
Score = 41.2 bits (95), Expect = 0.001
Identities = 31/123 (25%), Positives = 51/123 (41%), Gaps = 2/123 (1%)
Frame = +2
Query: 524 VDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
V L + C ++ + +P H IAA + +DFV Y+ Y+H +
Sbjct: 215 VKLNEWLCDYGQFQALRLPCSHAIAACAFCNLNYDDFVDPVYKLENIFKVYQHHFHSLGS 394
Query: 584 QKLWPTTNDEYILS-PLYRRAL-GRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQF 641
+ WP + +S P RR + GRP R+ + +++ N + K S C
Sbjct: 395 EDTWPQYLGPHFMSDPSKRRQISGRPATTRIHNEMDESIPNRLK--------KCSLCKTE 550
Query: 642 GHN 644
GHN
Sbjct: 551 GHN 559
>TC232149
Length = 1005
Score = 32.0 bits (71), Expect(2) = 0.003
Identities = 27/93 (29%), Positives = 48/93 (51%), Gaps = 4/93 (4%)
Frame = -3
Query: 454 VLTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERL-KWEIQM---TSHLIPQCCG*D 509
+L M + IR+YIM + + KM R ++P R+ K +++ T++ + G
Sbjct: 427 ILNMCEDIRTYIMCKMKSDKLKMATRLRPLVPMQQSRIEKGKVESNK*TANWVGDPDG-T 251
Query: 510 MFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIP 542
F+V H + + VDL+ + C W+L+GIP
Sbjct: 250 RFKVCHYDT--RLNVDLQN*SFTCRMWQLIGIP 158
Score = 27.3 bits (59), Expect(2) = 0.003
Identities = 17/55 (30%), Positives = 30/55 (53%)
Frame = -1
Query: 398 MMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDK 452
M ++ ++ +A+ +L + P +W K F+ C K YES+NA IL ++K
Sbjct: 573 MKVVKRMNEKAWAYLDKGPRESWTKAYFS--ENC-----KDYESYNAKILKLKNK 430
>BM108247 weakly similar to PIR|H84710|H847 Mutator-like transposase
[imported] - Arabidopsis thaliana, partial (4%)
Length = 265
Score = 38.5 bits (88), Expect = 0.009
Identities = 25/79 (31%), Positives = 41/79 (51%)
Frame = +2
Query: 380 NLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLY 439
+L+ A+ AT + +KM E++ + PEA WL Q S W F T+ L + +
Sbjct: 20 HLLWKASHATTTIAFKEKMGEIEEVSPEAAKWLQQFXPSQWALVHFK-GTRFGHLSSNI- 193
Query: 440 ESFNATILLARDKPVLTMV 458
E FN IL R+ P++ ++
Sbjct: 194 EEFNKWILDTRELPIIXVI 250
>AI856054
Length = 173
Score = 34.7 bits (78), Expect = 0.14
Identities = 16/37 (43%), Positives = 28/37 (75%)
Frame = -1
Query: 440 ESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKM 476
ESFN+ IL AR+K +++M++ I Y+M R+ T ++K+
Sbjct: 137 ESFNSVILEAREKLIVSMLEDIHIYLMERWDTNSKKV 27
>TC229129 similar to UP|Q6NM52 (Q6NM52) At4g15770, complete
Length = 661
Score = 28.9 bits (63), Expect = 7.5
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = -1
Query: 394 WTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTF 427
W K + + +H + + L PTSAWC+ F
Sbjct: 628 WIKTNYKTRKMHQSSSSILKYSPTSAWCRTTIPF 527
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.148 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,880,253
Number of Sequences: 63676
Number of extensions: 477316
Number of successful extensions: 3200
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 3173
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3198
length of query: 647
length of database: 12,639,632
effective HSP length: 103
effective length of query: 544
effective length of database: 6,081,004
effective search space: 3308066176
effective search space used: 3308066176
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0089b.6


