

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
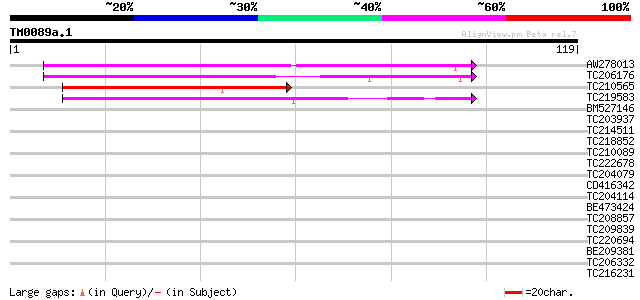
Score E
Sequences producing significant alignments: (bits) Value
AW278013 similar to SP|P27484|GRP2 Glycine-rich protein 2. [Wood... 58 7e-10
TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein... 57 9e-10
TC210565 similar to GB|AAO27142.1|27904309|AE014017 cold shock p... 41 7e-05
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 39 5e-04
BM527146 similar to GP|18491265|gb| At2g46160/T3F17.19 {Arabidop... 30 0.16
TC203937 thiol protease isoform A 28 0.47
TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protei... 28 0.47
TC218852 similar to GB|AAP37807.1|30725570|BT008448 At2g29660 {A... 28 0.61
TC210089 similar to UP|DBP1_YEAST (P24784) Probable ATP-dependen... 27 1.4
TC222678 weakly similar to UP|Q9SCR8 (Q9SCR8) Proline-rich prote... 27 1.4
TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding ... 23 2.2
CD416342 similar to SP|P42698|D111_ DNA-damage-repair/toleration... 26 2.3
TC204114 similar to UP|H2A_CICAR (O65759) Histone H2A, partial (... 26 2.3
BE473424 26 3.0
TC208857 weakly similar to UP|Q84KG8 (Q84KG8) Fatty acid delta-6... 26 3.0
TC209839 similar to UP|Q7Q155 (Q7Q155) AgCP8316 (Fragment), part... 26 3.0
TC220694 26 3.0
BE209381 similar to SP|O49885|R13A_ 60S ribosomal protein L13a. ... 26 3.0
TC206332 25 3.9
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 24 8.8
>AW278013 similar to SP|P27484|GRP2 Glycine-rich protein 2. [Wood tobacco]
{Nicotiana sylvestris}, partial (48%)
Length = 344
Score = 57.8 bits (138), Expect = 7e-10
Identities = 40/92 (43%), Positives = 49/92 (52%), Gaps = 1/92 (1%)
Frame = +1
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTILIPMAAL 67
V K+K FDDQKGF IT DD G + F+ S I+S GF LA E V+ L+ P
Sbjct: 37 VTGKVKWFDDQKGFGFITTDDSGEEFFVHQSSIRSDGFRSLALGESVEFLID-SDPEGRT 213
Query: 68 RR*RD*RPD*API*GTCCGGNDSSG-YGRGGG 98
+ PD +P+ GT GG G YG GGG
Sbjct: 214 KAVDVTGPDESPVQGTRGGGGGGRGSYGGGGG 309
>TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein-1, partial
(77%)
Length = 883
Score = 57.4 bits (137), Expect = 9e-10
Identities = 43/101 (42%), Positives = 53/101 (51%), Gaps = 10/101 (9%)
Frame = +3
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTILIPMAAL 67
V K+K F+DQKGF I+ DDG +DLF+ S+IKS GF LAE E V+ A+
Sbjct: 60 VSGKVKWFNDQKGFGFISPDDGSDDLFVHQSQIKSDGFRSLAEGESVEF---------AI 212
Query: 68 RR*RD*R--------PD*API*GTCCGGNDSSGY--GRGGG 98
D R PD A + GT GG+ Y GRGGG
Sbjct: 213 ESESDGRAKAVDVTGPDGASVQGTRRGGDGGRSYGGGRGGG 335
>TC210565 similar to GB|AAO27142.1|27904309|AE014017 cold shock protein CspA
{Buchnera aphidicola str. Bp (Baizongia pistaciae);} ,
partial (67%)
Length = 475
Score = 41.2 bits (95), Expect = 7e-05
Identities = 22/49 (44%), Positives = 35/49 (70%), Gaps = 1/49 (2%)
Frame = +3
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQ-GF*CLAEREFVDLLLT 59
+K F+++KGF IT +DGG+DLF+ S I++ GF L+E + V+ L+T
Sbjct: 81 VKWFNEEKGFGFITPEDGGSDLFVHYSAIQTDGGFRTLSEGQSVEFLVT 227
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 38.5 bits (88), Expect = 5e-04
Identities = 31/94 (32%), Positives = 47/94 (49%), Gaps = 7/94 (7%)
Frame = +2
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL-------TILIPM 64
+K F+ KGF IT DG +DLF+ + I+S G+ L++ + V+ LL T+ + +
Sbjct: 113 VKWFNAHKGFGFITPQDGTDDLFVHFTSIRSDGYRSLSDGQSVEFLLDYGDDGRTMAVDV 292
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
+ R R + G GG G GRGGG
Sbjct: 293 TSAVRSR--------LPGGFRGG--GGGRGRGGG 364
>BM527146 similar to GP|18491265|gb| At2g46160/T3F17.19 {Arabidopsis
thaliana}, partial (36%)
Length = 422
Score = 30.0 bits (66), Expect = 0.16
Identities = 15/37 (40%), Positives = 16/37 (42%)
Frame = -1
Query: 82 GTCCGGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRW 118
G CCGG GR G WR GR +R RW
Sbjct: 206 GCCCGGGGCGSRGRRRG*G-------WRGGRGRRGRW 117
>TC203937 thiol protease isoform A
Length = 1651
Score = 28.5 bits (62), Expect = 0.47
Identities = 15/38 (39%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Frame = -1
Query: 82 GTCCGGNDSSGYGRGGGYQEVVDIRIW-RNGRQKRWRW 118
G G + G G GGG + +D RIW R GR++ W
Sbjct: 397 GRAPGSGAA*GRGTGGGTRPELDRRIW*RRGRRRGRAW 284
>TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protein [imported]
- Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(88%)
Length = 1085
Score = 28.5 bits (62), Expect = 0.47
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = +1
Query: 105 IRIWRNGRQKRWRW 118
IRIW+ R+ RWRW
Sbjct: 688 IRIWKKSRKGRWRW 729
>TC218852 similar to GB|AAP37807.1|30725570|BT008448 At2g29660 {Arabidopsis
thaliana;} , partial (54%)
Length = 1201
Score = 28.1 bits (61), Expect = 0.61
Identities = 17/38 (44%), Positives = 19/38 (49%)
Frame = -1
Query: 61 LIPMAALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
LIP+ R* +*R TCCGG G G GGG
Sbjct: 202 LIPIRVGLR*GN*R--------TCCGGGGGGGSGGGGG 113
>TC210089 similar to UP|DBP1_YEAST (P24784) Probable ATP-dependent RNA
helicase DBP1 (Helicase CA1), partial (19%)
Length = 731
Score = 26.9 bits (58), Expect = 1.4
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = +1
Query: 87 GNDSSGYGRGGGY 99
GN+SSGYG GGY
Sbjct: 550 GNNSSGYGTSGGY 588
>TC222678 weakly similar to UP|Q9SCR8 (Q9SCR8) Proline-rich protein
(At3g50570), partial (14%)
Length = 508
Score = 26.9 bits (58), Expect = 1.4
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -3
Query: 87 GNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRW 118
G + YG G + V D+R G KRW+W
Sbjct: 287 GRNGWNYGV*GNVRNVGDLRNRNIGEGKRWQW 192
>TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein,
complete
Length = 748
Score = 23.1 bits (48), Expect(2) = 2.2
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = +1
Query: 86 GGNDSSGYGRGGGY 99
GG G GRGGGY
Sbjct: 349 GGGGGYGGGRGGGY 390
Score = 21.6 bits (44), Expect(2) = 2.2
Identities = 7/13 (53%), Positives = 8/13 (60%)
Frame = +3
Query: 106 RIWRNGRQKRWRW 118
R+WR RWRW
Sbjct: 444 RLWRRRWIWRWRW 482
>CD416342 similar to SP|P42698|D111_ DNA-damage-repair/toleration protein
DRT111 chloroplast precursor. [Mouse-ear cress],
partial (24%)
Length = 624
Score = 26.2 bits (56), Expect = 2.3
Identities = 13/33 (39%), Positives = 13/33 (39%)
Frame = +1
Query: 86 GGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRW 118
GG G GRG WR GR RW W
Sbjct: 172 GGGSLPGEGRGW----------WRRGRAWRWAW 240
>TC204114 similar to UP|H2A_CICAR (O65759) Histone H2A, partial (96%)
Length = 717
Score = 26.2 bits (56), Expect = 2.3
Identities = 7/13 (53%), Positives = 10/13 (76%)
Frame = -1
Query: 106 RIWRNGRQKRWRW 118
R+WRNG++ W W
Sbjct: 162 RLWRNGKRTLWSW 124
>BE473424
Length = 383
Score = 25.8 bits (55), Expect = 3.0
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = -3
Query: 93 YGRGGGYQEVVDIRIWRNGRQKRWRWF 119
+GRG G +R R R+++WRWF
Sbjct: 228 FGRGSG------LRGGRGRRRRQWRWF 166
>TC208857 weakly similar to UP|Q84KG8 (Q84KG8) Fatty acid delta-6 desaturase,
partial (21%)
Length = 648
Score = 25.8 bits (55), Expect = 3.0
Identities = 13/34 (38%), Positives = 15/34 (43%)
Frame = -3
Query: 85 CGGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWRW 118
CG D + R GG R W GR+ R RW
Sbjct: 295 CGRKDGGRWRRRGGGGR----RCWEGGRRGRRRW 206
>TC209839 similar to UP|Q7Q155 (Q7Q155) AgCP8316 (Fragment), partial (8%)
Length = 590
Score = 25.8 bits (55), Expect = 3.0
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +3
Query: 92 GYGRGGGYQEVVDIRIWR 109
G+ RGGG + V I++WR
Sbjct: 393 GFRRGGG*HQFVTIKVWR 446
>TC220694
Length = 548
Score = 25.8 bits (55), Expect = 3.0
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = -1
Query: 82 GTCCGGNDSSGYGRGGGYQEVVDIRIW 108
G+CCGG +G GG VV I IW
Sbjct: 257 GSCCGGGVE--WGADGGGVVVVGIVIW 183
>BE209381 similar to SP|O49885|R13A_ 60S ribosomal protein L13a. [Yellow
lupine] {Lupinus luteus}, partial (35%)
Length = 220
Score = 25.8 bits (55), Expect = 3.0
Identities = 7/18 (38%), Positives = 13/18 (71%)
Frame = +3
Query: 102 VVDIRIWRNGRQKRWRWF 119
++ +RIW+ ++R RWF
Sbjct: 66 IIPLRIWKKRERRRQRWF 119
>TC206332
Length = 519
Score = 25.4 bits (54), Expect = 3.9
Identities = 7/10 (70%), Positives = 10/10 (100%)
Frame = -1
Query: 108 WRNGRQKRWR 117
W+NGR+K+WR
Sbjct: 183 WKNGREKQWR 154
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 24.3 bits (51), Expect = 8.8
Identities = 9/14 (64%), Positives = 9/14 (64%)
Frame = +3
Query: 86 GGNDSSGYGRGGGY 99
GG GYG GGGY
Sbjct: 159 GGYGGGGYGGGGGY 200
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,113,106
Number of Sequences: 63676
Number of extensions: 61429
Number of successful extensions: 1270
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 1203
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1265
length of query: 119
length of database: 12,639,632
effective HSP length: 95
effective length of query: 24
effective length of database: 6,590,412
effective search space: 158169888
effective search space used: 158169888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0089a.1


