

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.21
(1215 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
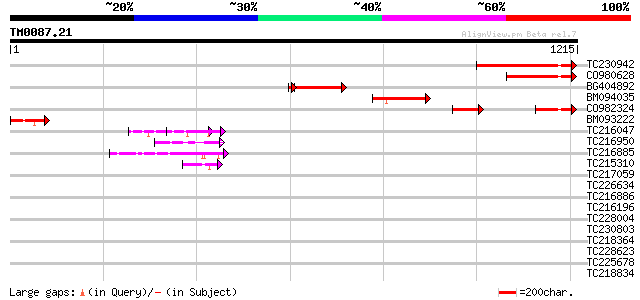
Score E
Sequences producing significant alignments: (bits) Value
TC230942 349 4e-96
CO980628 214 2e-55
BG404892 182 6e-49
BM094035 184 2e-46
CO982324 125 1e-28
BM093222 120 3e-27
TC216047 weakly similar to UP|IF3A_ARATH (Q9LD55) Eukaryotic tra... 49 1e-05
TC216950 49 1e-05
TC216885 45 2e-04
TC215310 similar to GB|AAP37804.1|30725564|BT008445 At2g15890 {A... 45 3e-04
TC217059 similar to UP|Q8L7W3 (Q8L7W3) At2g29210/F16P2.41, parti... 43 0.001
TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 40 0.006
TC216886 weakly similar to UP|BIR3_HUMAN (Q13489) Baculoviral IA... 40 0.008
TC216196 weakly similar to UP|BRAT_DROME (Q8MQJ9) Brain tumor pr... 40 0.008
TC228004 homologue to UP|IM30_PEA (Q03943) Membrane-associated 3... 39 0.011
TC230803 similar to UP|O45549 (O45549) C. elegans NUD-1 protein ... 38 0.031
TC218364 similar to UP|Q9P2S7 (Q9P2S7) Cisplatin resistance-asso... 37 0.040
TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (... 37 0.040
TC225678 similar to UP|Q6Y236 (Q6Y236) Kinase substrate HASPP28 ... 37 0.053
TC218834 UP|Q8I727 (Q8I727) TcC31.32, partial (8%) 37 0.069
>TC230942
Length = 1004
Score = 349 bits (896), Expect = 4e-96
Identities = 179/215 (83%), Positives = 192/215 (89%), Gaps = 1/215 (0%)
Frame = +3
Query: 1000 QVGSLMLESLSLLGHFALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTL 1059
QVGSL+LESLSLLGHFALFHPGNQAVLRWGKSPTILHKVCDLPF+FFSDPELMPILAGTL
Sbjct: 30 QVGSLVLESLSLLGHFALFHPGNQAVLRWGKSPTILHKVCDLPFVFFSDPELMPILAGTL 209
Query: 1060 VAACYGCEQNKFVVQQELSVDMLLSLLRSCKSAAPATQL-ITLDNSATDESSEPNQLGTE 1118
VAACYGCEQNKFVVQQELSVDMLLSLLRSC++AAPATQL TLDNS TDESSE NQLGTE
Sbjct: 210 VAACYGCEQNKFVVQQELSVDMLLSLLRSCRNAAPATQLNSTLDNSTTDESSECNQLGTE 389
Query: 1119 FRKTQVDIPVKHSRSNGKGIRAFLGKSGVLGNSLKNGRLKSLRDGKTTKSSEEVVPRQNL 1178
+K QVDIPVK+SRSNGKG RA GKSG GN++KNGR++S RDGKTTK+SEEV P+
Sbjct: 390 VKKPQVDIPVKNSRSNGKGPRASSGKSGASGNNIKNGRIRSQRDGKTTKNSEEVAPKH-- 563
Query: 1179 STSETSHSMLHCRFPQSFIDKVEQFFSAEIANGVD 1213
E S+ MLHCRFP SFIDKVEQFFSAEIAN VD
Sbjct: 564 --GEPSNLMLHCRFPPSFIDKVEQFFSAEIANRVD 662
>CO980628
Length = 800
Score = 214 bits (545), Expect = 2e-55
Identities = 113/150 (75%), Positives = 124/150 (82%), Gaps = 1/150 (0%)
Frame = -1
Query: 1065 GCEQNKFVVQQELSVDMLLSLLRSCKSAAPATQL-ITLDNSATDESSEPNQLGTEFRKTQ 1123
GCEQNKFVVQQELSVDMLLSLLRSC++AAPATQL TLDNS TDES E NQLGTE +K Q
Sbjct: 800 GCEQNKFVVQQELSVDMLLSLLRSCRNAAPATQLNSTLDNSTTDESGECNQLGTEIKKPQ 621
Query: 1124 VDIPVKHSRSNGKGIRAFLGKSGVLGNSLKNGRLKSLRDGKTTKSSEEVVPRQNLSTSET 1183
VD PVK+SRSNGKG RA GKSG GN++KN R++S RDGK TK+SEEV P+ E
Sbjct: 620 VDFPVKNSRSNGKGTRASSGKSGASGNNIKNCRIRSQRDGKITKNSEEVAPKH----GEP 453
Query: 1184 SHSMLHCRFPQSFIDKVEQFFSAEIANGVD 1213
S+ MLHCRFP SFIDKVEQFFSAEIANGVD
Sbjct: 452 SNLMLHCRFPPSFIDKVEQFFSAEIANGVD 363
>BG404892
Length = 371
Score = 182 bits (461), Expect(2) = 6e-49
Identities = 93/111 (83%), Positives = 98/111 (87%)
Frame = +3
Query: 610 FIASALVASHTSKPEACQVTLHLLKLLKVVLSAPANRSYFLAQNLLPPIIPMLSAALENY 669
F AL ASHTSKPEACQV LHLLKLL+VVLS PANRSYFLAQNLLPPIIPMLSAALENY
Sbjct: 39 FYCFALPASHTSKPEACQVMLHLLKLLRVVLSTPANRSYFLAQNLLPPIIPMLSAALENY 218
Query: 670 IKIAASLSSPGNFSLPSNKASLENFESVSEILNNFLWAVTAILGHISSEER 720
IKIAASLS PGN SLP +KAS+ENFES+SEILNN LW VTAI GHI+ EER
Sbjct: 219 IKIAASLSIPGNISLPPSKASVENFESISEILNNXLWTVTAIFGHINLEER 371
Score = 32.3 bits (72), Expect(2) = 6e-49
Identities = 15/16 (93%), Positives = 16/16 (99%)
Frame = +1
Query: 598 ELQASRQAGLIDFIAS 613
ELQASRQAGL+DFIAS
Sbjct: 4 ELQASRQAGLLDFIAS 51
>BM094035
Length = 415
Score = 184 bits (467), Expect = 2e-46
Identities = 94/136 (69%), Positives = 107/136 (78%), Gaps = 11/136 (8%)
Frame = +3
Query: 777 GKSSYIDWEYSPVAMELEIGGEGAKFADSV-----------GPLSVINGSSVMHLPDVPE 825
G+ SYIDWE S VAME EIG EGAKFAD+ PLSV GSSV+HLPDVPE
Sbjct: 6 GRLSYIDWESSLVAMEQEIGSEGAKFADAAHFVVNNSWENFNPLSVTTGSSVVHLPDVPE 185
Query: 826 DRPLDETIKVNRNEESISIGKDCKLEHDSSDKLKNDEMEKIDDLDEPQKNQGGDIANSFV 885
DRPL+E IKVN+++ESISIGKDC+LEHDSS KLKND+MEKIDDLDE +KNQ GDI N V
Sbjct: 186 DRPLEEMIKVNKSDESISIGKDCELEHDSSVKLKNDDMEKIDDLDESKKNQNGDITNLSV 365
Query: 886 SQKDEKHTMVNITTQK 901
QKDEKHT+V++T QK
Sbjct: 366 LQKDEKHTVVSVTVQK 413
>CO982324
Length = 776
Score = 125 bits (313), Expect = 1e-28
Identities = 60/66 (90%), Positives = 64/66 (96%)
Frame = -3
Query: 950 GVLKVLNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESL 1009
GVLKVLNNVALLDLVFLQ+MLARPDLKMEIFHLM F+LSHCAS+WK PNDQVGSL+LESL
Sbjct: 771 GVLKVLNNVALLDLVFLQQMLARPDLKMEIFHLMGFLLSHCASKWKAPNDQVGSLVLESL 592
Query: 1010 SLLGHF 1015
SLLGHF
Sbjct: 591 SLLGHF 574
Score = 119 bits (298), Expect = 8e-27
Identities = 61/87 (70%), Positives = 69/87 (79%)
Frame = -3
Query: 1127 PVKHSRSNGKGIRAFLGKSGVLGNSLKNGRLKSLRDGKTTKSSEEVVPRQNLSTSETSHS 1186
PVK+SRSNGKG RA GKSG GN++KN R++S RDGK TK+SEEV P+ E S+
Sbjct: 573 PVKNSRSNGKGTRASSGKSGASGNNIKNCRIRSQRDGKITKNSEEVAPKH----GEPSNL 406
Query: 1187 MLHCRFPQSFIDKVEQFFSAEIANGVD 1213
MLHCRFP SFIDKVEQFFSAEIANGVD
Sbjct: 405 MLHCRFPPSFIDKVEQFFSAEIANGVD 325
>BM093222
Length = 421
Score = 120 bits (302), Expect = 3e-27
Identities = 68/91 (74%), Positives = 73/91 (79%), Gaps = 6/91 (6%)
Frame = -1
Query: 1 MTTSPHRADILSSSLEAFRKIQQERASLKLSNNTEHAMSKCLTSESVGNMN------GTH 54
MTTSPHRADILSSSLEAFRKIQQERASL+ S TE+AMSKC+TSES+GN N GT
Sbjct: 271 MTTSPHRADILSSSLEAFRKIQQERASLQ-SGTTENAMSKCVTSESIGNTNKSRVNDGTD 95
Query: 55 NAKNSRTKSRKHIGSSDANQGNLNVKKHNIE 85
AK S TKSRK +GSSDA QGNLN KK NIE
Sbjct: 94 VAKYSVTKSRKQVGSSDAKQGNLNGKKRNIE 2
>TC216047 weakly similar to UP|IF3A_ARATH (Q9LD55) Eukaryotic translation
initiation factor 3 subunit 10 (eIF-3 theta) (Eukaryotic
translation initiation factor 3 large subunit) (eIF3a)
(p114), partial (22%)
Length = 664
Score = 49.3 bits (116), Expect = 1e-05
Identities = 50/195 (25%), Positives = 85/195 (42%), Gaps = 13/195 (6%)
Frame = +2
Query: 254 LRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYAR--------HQRSESRHE 305
LR R E+E +++QKL +T L R A L E Y + H R + +
Sbjct: 98 LRERQEME-KKLQKLAKTMDHLERAKREEAA---PLIEAAYQQRLVEERLLHDREQQQEV 265
Query: 306 AFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKED 365
Q + GD K VR + + +++ ++ + LRR + ++ + Q
Sbjct: 266 ELSKQ--RHEGDLKEKERLVRMMGNKEIYQARVVSHRQAEFNRLRREREERISRILQSRR 439
Query: 366 LAREEAVLERRKL-----IEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQL 420
RE+ RKL +E E+ Q+L E +EEA+ R + ERK R +E++
Sbjct: 440 QEREKM----RKLKYYLKLEEERQQKLHE----EEEARKREDAERKKKEEEERLRKLEEI 595
Query: 421 RRKEERAKAQQEEAE 435
K+ + + + EE E
Sbjct: 596 AEKQRQIERELEEKE 640
Score = 44.3 bits (103), Expect = 3e-04
Identities = 36/135 (26%), Positives = 65/135 (47%), Gaps = 8/135 (5%)
Frame = +2
Query: 336 KKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKE 395
++L+ + LH E + +++++ K + + DL +E ++ R + E Q R+ E
Sbjct: 212 QRLVEERLLHDRE--QQQEVELSKQRHEGDLKEKERLV--RMMGNKEIYQARVVSHRQAE 379
Query: 396 EAQIRREEERKASSAAREARAIEQLRRK--------EERAKAQQEEAELLAQKLAERLNE 447
++RRE E + S + R + RK EER + EE E ++ AER +
Sbjct: 380 FNRLRREREERISRILQSRRQEREKMRKLKYYLKLEEERQQKLHEEEEARKREDAERKKK 559
Query: 448 SEQRRKIYLEQIRER 462
E+ R LE+I E+
Sbjct: 560 EEEERLRKLEEIAEK 604
>TC216950
Length = 873
Score = 48.9 bits (115), Expect = 1e-05
Identities = 47/152 (30%), Positives = 76/152 (49%), Gaps = 1/152 (0%)
Frame = +2
Query: 310 QVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLARE 369
+ ++ +ES K EV+ E + L ++ ++L + ++ +I++KQKE+ AR+
Sbjct: 260 EAIRKRVEESLKSEEVQVEIERRLEEGRKRLNDEV-AAQLEKEKEAALIEAKQKEEQARK 436
Query: 370 EAVLERRKLIEAEKLQR-LAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAK 428
E E L+R L E +RK EEAQ RRE A+EQ RR+EER K
Sbjct: 437 EK----------EDLERMLEENRRKVEEAQ-RRE-------------ALEQQRREEERYK 544
Query: 429 AQQEEAELLAQKLAERLNESEQRRKIYLEQIR 460
+E + + ++ +E EQ R L QI+
Sbjct: 545 ELEEMQRQKEEAMRKKKHEEEQER---LNQIK 631
Score = 32.7 bits (73), Expect = 1.00
Identities = 42/179 (23%), Positives = 76/179 (41%)
Frame = +2
Query: 228 KLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHL 287
KL+ E K+ ++K EE E++ E ++L+ ++LN E A
Sbjct: 224 KLIEEETAKRVEEAIRKRVEESLKS-----EEVQVEIERRLEEGRKRLN--DEVAA---- 370
Query: 288 KLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGS 347
+ E EA L + AK+ +++ K E + + EEN++ +
Sbjct: 371 -----------QLEKEKEAALIE-AKQKEEQARKEKED--LERMLEENRRKV-------- 484
Query: 348 ELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERK 406
E+ R EA+ ++R+ E E+ + L EMQR+KEEA +++ E +
Sbjct: 485 ----------------EEAQRREALEQQRR--EEERYKELEEMQRQKEEAMRKKKHEEE 607
>TC216885
Length = 1122
Score = 45.1 bits (105), Expect = 2e-04
Identities = 67/275 (24%), Positives = 131/275 (47%), Gaps = 19/275 (6%)
Frame = +3
Query: 214 LGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQ 273
L ++ ER H+K + + KK +SDL + + K R + ET ++L +
Sbjct: 33 LEKQVKERKDWAHEKAI--QAAKKLSSDLIELKKFKMEREENKKLPKETGAAEELDNPT- 203
Query: 274 KLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITS--- 330
+ R+SE L+ G + + + EA A++ +A E+SK++ +TS
Sbjct: 204 -MMRLSEMENA--LRKTSGQMDQATAAVRKLEAEKAEI--KAELEASKLSASESVTSCLQ 368
Query: 331 LNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEM 390
+ + KK + +KL E ++ + Q I ++++ L +E + + ++ + ++ R E+
Sbjct: 369 VAKREKKCL--KKLLTWEKQKVKIHQDISDEKQKILEIQEELAQIKQCAKETEVTRKEEL 542
Query: 391 QRKKEEAQIRREEERKASSAA-----REARA------IEQLRRKEERAKAQQEEAELLAQ 439
+ KEEA EEER++ AA R +A I+ RRK++ + +QE + L A
Sbjct: 543 -KAKEEALALIEEERRSKEAAEANHKRNLKALRLKIEIDFQRRKDDLLRLEQEISRLKAP 719
Query: 440 KLAERL-----NESEQRRKIYLEQIRERANLRDQS 469
+ L ++E +R+ + + E N++D S
Sbjct: 720 ARSTTLPTSESEDAEPQRETLAKLLLELDNVKDFS 824
>TC215310 similar to GB|AAP37804.1|30725564|BT008445 At2g15890 {Arabidopsis
thaliana;} , partial (64%)
Length = 1381
Score = 44.7 bits (104), Expect = 3e-04
Identities = 31/92 (33%), Positives = 50/92 (53%), Gaps = 5/92 (5%)
Frame = -1
Query: 370 EAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERA-- 427
EA LER + ++ ++ QR+ E Q + E R EEER + +E E+ RR EE A
Sbjct: 265 EAELERVQRMQEQERQRIIEEQERALELARREEEERLRQAREQE----ERQRRLEEEARE 98
Query: 428 ---KAQQEEAELLAQKLAERLNESEQRRKIYL 456
+A+QE E L + +RL E+++++ L
Sbjct: 97 AAWRAEQERIEALRKAEEQRLAREEEKQRMVL 2
Score = 30.8 bits (68), Expect = 3.8
Identities = 29/102 (28%), Positives = 49/102 (47%)
Frame = -1
Query: 305 EAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKE 364
EA L +V + E ++ E E+ + L L ++ LR+A + Q + ++ E
Sbjct: 265 EAELERVQRMQEQERQRIIE--------EQERALELARREEEERLRQARE-QEERQRRLE 113
Query: 365 DLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERK 406
+ ARE A ++ IEA RK EE ++ REEE++
Sbjct: 112 EEAREAAWRAEQERIEA---------LRKAEEQRLAREEEKQ 14
>TC217059 similar to UP|Q8L7W3 (Q8L7W3) At2g29210/F16P2.41, partial (9%)
Length = 1565
Score = 42.7 bits (99), Expect = 0.001
Identities = 40/180 (22%), Positives = 88/180 (48%)
Frame = +1
Query: 284 VRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQK 343
++ +++ ++ + +S RH ++ + + +VN S++E++ K +
Sbjct: 778 IKKSDVKDQSHSNYAKSSYRHHK--SETTQDLVGKGDRVNHNASHDSVSEDSGK----HR 939
Query: 344 LHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREE 403
G + +R ++ + + ED + +++ LE RK K +R E + +KEE + RREE
Sbjct: 940 REGKDRKRHKRSEKKYTSSDEDYS-DDSELEDRK---EAKRRRKEEKKLQKEEKRRRREE 1107
Query: 404 ERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+R+ RE R E+L+ K + +E A+++ ++SE +K ++R +A
Sbjct: 1108KRR----RREERRAEKLKMKSKTDDISDDEE---AKQMGYHQSDSEDEQKKLEIELRNKA 1266
>TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (51%)
Length = 1024
Score = 40.0 bits (92), Expect = 0.006
Identities = 38/144 (26%), Positives = 73/144 (50%), Gaps = 12/144 (8%)
Frame = +2
Query: 339 ILRQKLHGSELRRAEKLQVIKS---------KQKEDLAREEAVLE---RRKLIEAEKLQR 386
++ + + E+ EK+++ K ++ D ARE L+ ++K ++ E+L+R
Sbjct: 263 VVHEAIRKMEIVADEKMRMFKKARLAFDACERELADKAREAGELKMDRQKKKLQIEELER 442
Query: 387 LAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLN 446
+ + K EA + + KA+ A REA EQL+R AK+ + E E + L ++L+
Sbjct: 443 IVRL--KNAEADMF---QLKANEAKREA---EQLQR-IALAKSDKSEEEYTSNYLKQKLS 595
Query: 447 ESEQRRKIYLEQIRERANLRDQSS 470
E+E ++ E+I+ + R S
Sbjct: 596 EAEAEKQYLYEKIKLQETSRVSQS 667
>TC216886 weakly similar to UP|BIR3_HUMAN (Q13489) Baculoviral IAP
repeat-containing protein 3 (Inhibitor of apoptosis
protein 1) (HIAP1) (HIAP-1) (C-IAP2) (TNFR2-TRAF
signaling complex protein 1) (IAP homolog C) (Apoptosis
inhibitor 2) (API2), partial (4%)
Length = 1137
Score = 39.7 bits (91), Expect = 0.008
Identities = 42/147 (28%), Positives = 71/147 (47%), Gaps = 5/147 (3%)
Frame = +1
Query: 314 RAGDESSKVNEVRFITSLNE---ENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREE 370
+A E+SK++ +T+ E KK + +KL E ++A KLQ S +KE + + +
Sbjct: 121 KAEMEASKLSASESVTACLEVAKREKKCL--KKLLAWEKQKA-KLQQEISDEKEKILKTK 291
Query: 371 AVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAA--REARAIEQLRRKEERAK 428
+L + + + E + E + KEEA EEER AA R +E LR K E
Sbjct: 292 EILVQIRQCQKEAEVKWKEELKAKEEALALVEEERHCKEAAESNNKRKLEALRLKIEIDF 471
Query: 429 AQQEEAELLAQKLAERLNESEQRRKIY 455
+ ++ L ++ RL S Q +++
Sbjct: 472 QRHKDDLLRLEQELSRLKASAQSAELH 552
>TC216196 weakly similar to UP|BRAT_DROME (Q8MQJ9) Brain tumor protein,
partial (3%)
Length = 1679
Score = 39.7 bits (91), Expect = 0.008
Identities = 24/70 (34%), Positives = 46/70 (65%), Gaps = 3/70 (4%)
Frame = +3
Query: 379 IEAEKLQRLAEMQRKKEE--AQIRREEERKASSAAREARAIEQLRRKEER-AKAQQEEAE 435
I+ E+ + L ++ ++EE A+I++EEE + + +E + ++++++ER AKAQ EE E
Sbjct: 36 IDDEEDEHLVKVHLEEEERLAKIQQEEEERLAKIQQEDERLAKIQQEDERLAKAQLEEDE 215
Query: 436 LLAQKLAERL 445
LA+ + E L
Sbjct: 216 QLARAIQESL 245
>TC228004 homologue to UP|IM30_PEA (Q03943) Membrane-associated 30 kDa
protein, chloroplast precursor (M30), partial (64%)
Length = 1088
Score = 39.3 bits (90), Expect = 0.011
Identities = 44/176 (25%), Positives = 77/176 (43%), Gaps = 4/176 (2%)
Frame = +1
Query: 340 LRQKLHGSELRRAE-KLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQ 398
+ KLH L+ K Q+ K +EDLARE L+RRK A++ ++K +
Sbjct: 241 INTKLHNKLLKSGTXKAQLALQKGEEDLARE--ALKRRKSYADNASSLKAQLDQQKTVVE 414
Query: 399 IRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERL-NESEQRRKIYLE 457
R S +EAR+ K++ KA+ + A+ A K++E L N + E
Sbjct: 415 NLVSNTRLLESKIQEARS------KKDTLKARAQSAK-TATKVSEMLGNVNTSSALSAFE 573
Query: 458 QIRERANLRDQSSPLLRRSLNKDGQGRST--PNNSSDDSQTNIASGLGSSLGIGSI 511
++ E+ + + L + + D +G+ +S DD N+ L + G +
Sbjct: 574 KMEEKVLTMESQAEALGQLTSDDLEGKFALLEGSSVDDDLANLKKELSGASKKGDL 741
>TC230803 similar to UP|O45549 (O45549) C. elegans NUD-1 protein
(Corresponding sequence F53A2.4), partial (6%)
Length = 949
Score = 37.7 bits (86), Expect = 0.031
Identities = 23/66 (34%), Positives = 35/66 (52%)
Frame = +3
Query: 388 AEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNE 447
A+ +KKE + REE R+A ARE+R +Q R E R + + EE E +KL E
Sbjct: 288 AKASKKKEIKRQEREERRQAEEVARESRLAKQDRYSEMR-RLKDEEREAQERKLEEEARA 464
Query: 448 SEQRRK 453
+ + +
Sbjct: 465 QKAKEE 482
Score = 36.6 bits (83), Expect = 0.069
Identities = 24/75 (32%), Positives = 42/75 (56%)
Frame = +3
Query: 359 KSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIE 418
K+ +K+++ R+E ERR+ E + RLA+ R E ++ ++EER+A E A
Sbjct: 291 KASKKKEIKRQERE-ERRQAEEVARESRLAKQDRYSEMRRL-KDEEREAQERKLEEEARA 464
Query: 419 QLRRKEERAKAQQEE 433
Q ++EE A + E+
Sbjct: 465 QKAKEEEAAALEFEK 509
>TC218364 similar to UP|Q9P2S7 (Q9P2S7) Cisplatin resistance-associated
overexpressed protein, partial (6%)
Length = 868
Score = 37.4 bits (85), Expect = 0.040
Identities = 43/154 (27%), Positives = 70/154 (44%), Gaps = 14/154 (9%)
Frame = +2
Query: 331 LNEENKKLILRQKLHGSELRRAEKLQVIKSKQKE------DLAREEAVLERRKLIEAEKL 384
L E + R+ E +R E L+V KQ E + REE ER KL +
Sbjct: 56 LREMETRFEQREVERERERQRRENLRVELDKQWERKLEEREKEREEREKERDKL----RR 223
Query: 385 QRLAE---MQRKKEEAQIRREEERKASSAARE----ARAIEQLRRKEER-AKAQQEEAEL 436
QR+AE M+++ EE + +R EE E R +E +R +E ++ + E ++
Sbjct: 224 QRMAEWEAMEKENEEMERKRREEELIHEREWEERMNCRRLEWKKRVDEMLSQHRAEMGQM 403
Query: 437 LAQKLAERLNESEQRRKIYLEQIRERANLRDQSS 470
+ L E+ N + Q I+ + + A L D +S
Sbjct: 404 QTRFLHEQQNLTNQLLGIFSQWPAQPAGLSDHTS 505
>TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (85%)
Length = 1082
Score = 37.4 bits (85), Expect = 0.040
Identities = 27/109 (24%), Positives = 53/109 (47%), Gaps = 2/109 (1%)
Frame = +3
Query: 350 RRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEER--KA 407
R AE + +K+KE+ A + A RK + +RLAE + K+ E +++ +++ +
Sbjct: 225 REAEGSKSRAAKKKEEEAEKRAEAAARKA----EARRLAEQEEKELEKSMKKVDKKATRV 392
Query: 408 SSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYL 456
S + +E RR+EE + +AE ++ + E E R + +
Sbjct: 393 SIPVPKVTEVELRRRREEEQAEAERKAEEAKKQQSRTAAEEEYERMVLI 539
>TC225678 similar to UP|Q6Y236 (Q6Y236) Kinase substrate HASPP28 (Fragment),
partial (54%)
Length = 912
Score = 37.0 bits (84), Expect = 0.053
Identities = 37/131 (28%), Positives = 68/131 (51%), Gaps = 13/131 (9%)
Frame = +1
Query: 298 QRSESRHEAFLAQVA-KRAGDESSKVNEVRFITS--LNEENKKLILRQKLHG-------- 346
++ E+ HE +V+ +G+ES + + T + EN L+ + L
Sbjct: 274 RQKEAEHEEEPEEVSGDESGEESEEETSKKKGTQGVIEIENPNLVKPKSLKARDVDVGKT 453
Query: 347 SELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEK-LQRLAEMQRKKEEAQIRREEER 405
+EL R E+ ++ K + E R + E+ K +A+K L+RLA +++++E+A +REEE+
Sbjct: 454 TELSRREREEIEKQRAHERYMRLQ---EQGKTEQAKKDLERLALIRQQREDAAKKREEEK 624
Query: 406 KASSAAR-EAR 415
A + EAR
Sbjct: 625 AAKEQKKAEAR 657
>TC218834 UP|Q8I727 (Q8I727) TcC31.32, partial (8%)
Length = 1405
Score = 36.6 bits (83), Expect = 0.069
Identities = 38/145 (26%), Positives = 64/145 (43%)
Frame = +3
Query: 350 RRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASS 409
RR +KL + ++QK EEA L L++ +R AEM+++ E ++ E ER +
Sbjct: 336 RRQKKLLMRHARQKH---LEEATLREADLLQELDRERTAEMEKELERQRL-LEIERAKTK 503
Query: 410 AAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERANLRDQS 469
R +E+ R+ + + + E+AE + S + R ++ RER N R +
Sbjct: 504 ELRHNLDMEKERQTQRELQREIEQAESGLRPSRRDFPSSSRPR----DRFRERENGRSGN 671
Query: 470 SPLLRRSLNKDGQGRSTPNNSSDDS 494
+R G G P S S
Sbjct: 672 EGSIRA-----GSGSLQPEIPSTSS 731
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.129 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,597,874
Number of Sequences: 63676
Number of extensions: 552121
Number of successful extensions: 2690
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 2619
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2669
length of query: 1215
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1107
effective length of database: 5,762,624
effective search space: 6379224768
effective search space used: 6379224768
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0087.21


