

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.11
(146 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
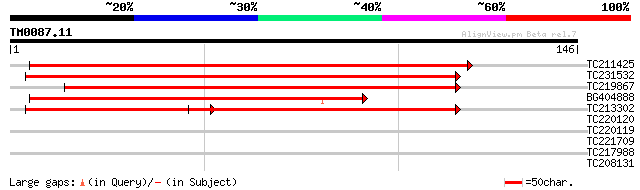
Score E
Sequences producing significant alignments: (bits) Value
TC211425 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transac... 216 3e-57
TC231532 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transac... 154 2e-38
TC219867 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transac... 150 2e-37
BG404888 140 3e-34
TC213302 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transac... 105 3e-32
TC220120 similar to GB|AAP80172.1|32362289|BT009671 At1g17330 {A... 27 3.5
TC220119 similar to GB|AAP80172.1|32362289|BT009671 At1g17330 {A... 27 3.5
TC221709 27 3.5
TC217988 similar to UP|Q9SSU7 (Q9SSU7) Endo-1,4-beta-glucanase, ... 26 7.8
TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein... 26 7.8
>TC211425 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transactivated
protein A-like {Oryza sativa (japonica cultivar-group);}
, partial (59%)
Length = 589
Score = 216 bits (550), Expect = 3e-57
Identities = 107/114 (93%), Positives = 109/114 (94%)
Frame = +2
Query: 6 SNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVAR 65
SNGF RDVSLQ+LSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVA+
Sbjct: 35 SNGFPRKRDVSLQELSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVAK 214
Query: 66 GLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTS 119
GLPNWS DDKEHLEEELSDVLLYLVRLADVCGLDLGQAAL KIVKNAQKYPVTS
Sbjct: 215 GLPNWSSDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALTKIVKNAQKYPVTS 376
>TC231532 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transactivated
protein A-like {Oryza sativa (japonica cultivar-group);}
, partial (52%)
Length = 732
Score = 154 bits (388), Expect = 2e-38
Identities = 76/112 (67%), Positives = 88/112 (77%)
Frame = +3
Query: 5 KSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVA 64
K G VSL L + L EFA+ R W+QYHSPRNLLLALVGEVGELSEIFQWKGEV
Sbjct: 219 KMTGVPQEAPVSLDQLKQILDEFAKERDWEQYHSPRNLLLALVGEVGELSEIFQWKGEVP 398
Query: 65 RGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
+GLP+W ++K HL EELSDVLLYLVRL+D+CG+DLG+AAL K+ NA KYP
Sbjct: 399 KGLPDWKEEEKVHLGEELSDVLLYLVRLSDICGVDLGKAALRKVQLNAIKYP 554
>TC219867 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transactivated
protein A-like {Oryza sativa (japonica cultivar-group);}
, partial (63%)
Length = 660
Score = 150 bits (379), Expect = 2e-37
Identities = 71/102 (69%), Positives = 87/102 (84%)
Frame = +3
Query: 15 VSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDD 74
V+L L + +A+FA+ R WD++HSPRNLLLALVGEVGELSEIFQWKGEV +GLP+W ++
Sbjct: 54 VTLDTLKQLMAQFAKERDWDRFHSPRNLLLALVGEVGELSEIFQWKGEVPKGLPDWKEEE 233
Query: 75 KEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
K HL EELSDVLLYLVRL+D+CG+DLG+AAL K+ NA KYP
Sbjct: 234 KVHLGEELSDVLLYLVRLSDICGVDLGKAALRKVQLNAIKYP 359
>BG404888
Length = 383
Score = 140 bits (352), Expect = 3e-34
Identities = 72/88 (81%), Positives = 75/88 (84%), Gaps = 1/88 (1%)
Frame = +2
Query: 6 SNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVAR 65
SNGF RDVSLQ+LSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVA+
Sbjct: 5 SNGFPRKRDVSLQELSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVAK 184
Query: 66 GLPNWSCDDKEHLE-EELSDVLLYLVRL 92
GLPNWS DDK L + V LYLVRL
Sbjct: 185GLPNWSSDDKGALRGKTFQMVWLYLVRL 268
>TC213302 similar to GB|BAD22064.1|47848239|AP005316 XTP3-transactivated
protein A-like {Oryza sativa (japonica cultivar-group);}
, partial (52%)
Length = 919
Score = 105 bits (262), Expect(2) = 3e-32
Identities = 50/70 (71%), Positives = 60/70 (85%)
Frame = +3
Query: 47 VGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALA 106
+GEVGELSEIFQWKGEV +GL +W ++K HL EELSDVLLYLVRL+D+CG+DLG+AAL
Sbjct: 417 LGEVGELSEIFQWKGEVPKGLLDWKEEEKVHLGEELSDVLLYLVRLSDMCGVDLGKAALR 596
Query: 107 KIVKNAQKYP 116
K+ NA KYP
Sbjct: 597 KVQLNAVKYP 626
Score = 49.3 bits (116), Expect(2) = 3e-32
Identities = 25/49 (51%), Positives = 31/49 (63%)
Frame = +2
Query: 5 KSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGEL 53
K G VSL L + + EFA+ R W+Q+HSPRNLLLALVG G +
Sbjct: 290 KMAGVPPEAHVSLDQLKQIMDEFAKERDWEQFHSPRNLLLALVG*SGRI 436
>TC220120 similar to GB|AAP80172.1|32362289|BT009671 At1g17330 {Arabidopsis
thaliana;} , partial (40%)
Length = 411
Score = 26.9 bits (58), Expect = 3.5
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -3
Query: 57 FQWKGEVARGLPNWSCDDKEH 77
F W G+ RGLP W +++H
Sbjct: 136 FPWSGDWKRGLPLWQGREQDH 74
>TC220119 similar to GB|AAP80172.1|32362289|BT009671 At1g17330 {Arabidopsis
thaliana;} , partial (50%)
Length = 479
Score = 26.9 bits (58), Expect = 3.5
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -2
Query: 57 FQWKGEVARGLPNWSCDDKEH 77
F W G+ RGLP W +++H
Sbjct: 256 FPWSGDWKRGLPLWQGREQDH 194
>TC221709
Length = 505
Score = 26.9 bits (58), Expect = 3.5
Identities = 12/37 (32%), Positives = 20/37 (53%)
Frame = -1
Query: 59 WKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADV 95
WK A+ P WS ++H++ + V L L+ L D+
Sbjct: 247 WKQTKAKNRPEWSESVEQHIKTKKMIVKLLLLWLCDI 137
>TC217988 similar to UP|Q9SSU7 (Q9SSU7) Endo-1,4-beta-glucanase, partial
(54%)
Length = 1156
Score = 25.8 bits (55), Expect = 7.8
Identities = 12/37 (32%), Positives = 18/37 (48%)
Frame = +2
Query: 25 AEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKG 61
AEF GWD H+ +LL+ V + + +KG
Sbjct: 155 AEFDNTFGWDNKHAGARILLSKEFLVQRVQSLHDYKG 265
>TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like,
partial (32%)
Length = 1371
Score = 25.8 bits (55), Expect = 7.8
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = +3
Query: 56 IFQWKGEVARGLPNWSCDDKEHLEEEL 82
IF KGEV NWS D EH +E+
Sbjct: 459 IFPGKGEVWALYRNWSPDWNEHTPDEV 539
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,320,309
Number of Sequences: 63676
Number of extensions: 63693
Number of successful extensions: 229
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 229
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 229
length of query: 146
length of database: 12,639,632
effective HSP length: 88
effective length of query: 58
effective length of database: 7,036,144
effective search space: 408096352
effective search space used: 408096352
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0087.11


