

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.5
(176 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
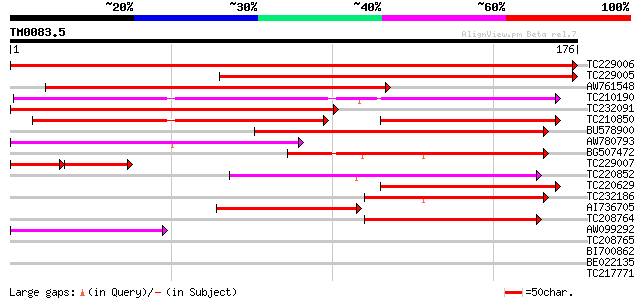
TC229006 327 2e-90
TC229005 212 8e-56
AW761548 137 2e-33
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 136 4e-33
TC232091 124 2e-29
TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 96 1e-20
BU578900 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidop... 77 3e-15
AW780793 71 3e-13
BG507472 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidop... 62 1e-10
TC229007 38 1e-08
TC220852 54 3e-08
TC220629 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 53 8e-08
TC232186 51 2e-07
AI736705 similar to GP|11994736|db far-red impaired response pro... 48 2e-06
TC208764 similar to UP|Q9ZVC9 (Q9ZVC9) Mutator-like transposase ... 45 2e-05
AW099292 42 2e-04
TC208765 39 0.002
BI700862 weakly similar to GP|16604382|gb| AT3g59470/T16L24_20 {... 36 0.011
BE022135 35 0.018
TC217771 27 0.48
>TC229006
Length = 991
Score = 327 bits (838), Expect = 2e-90
Identities = 156/176 (88%), Positives = 169/176 (95%)
Frame = +2
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFESEDAAK+FYDEYAR +GFVMRVMSCRR ERDGRILARRLGCNKEG+C++IRGK S
Sbjct: 113 MEFESEDAAKIFYDEYARRLGFVMRVMSCRRPERDGRILARRLGCNKEGYCVSIRGKFAS 292
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQEL 120
+R+PRASTREGCKAMIHIKY+KSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQEL
Sbjct: 293 VRKPRASTREGCKAMIHIKYNKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQEL 472
Query: 121 TAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEELLHQT 176
TAE+R KKRLCATYQEQL+SFMKIVEEH++KLS KIHH+VNNLKEFE IEELLHQT
Sbjct: 473 TAELRHKKRLCATYQEQLTSFMKIVEEHNEKLSAKIHHVVNNLKEFESIEELLHQT 640
>TC229005
Length = 679
Score = 212 bits (539), Expect = 8e-56
Identities = 102/111 (91%), Positives = 107/111 (95%)
Frame = +1
Query: 66 ASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQELTAEIR 125
ASTREGCKAMIHIKYDKSGKWVI KFVKDHNHPLVVSPREARQTMDEKDKKIQELTAE+R
Sbjct: 1 ASTREGCKAMIHIKYDKSGKWVIAKFVKDHNHPLVVSPREARQTMDEKDKKIQELTAELR 180
Query: 126 IKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEELLHQT 176
+KKRLCATYQEQLSSFMKIVEE ++KLS KIHH+VNNLKEFE IEELLHQT
Sbjct: 181 LKKRLCATYQEQLSSFMKIVEERNEKLSAKIHHVVNNLKEFESIEELLHQT 333
>AW761548
Length = 322
Score = 137 bits (346), Expect = 2e-33
Identities = 65/107 (60%), Positives = 80/107 (74%)
Frame = +2
Query: 12 FYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRASTREG 71
FYDEYAR VMRVMSC S+RDGRILAR LGCNK G+C+++R + S+R+P ASTRE
Sbjct: 2 FYDEYARR**SVMRVMSCGPSQRDGRILARTLGCNKHGYCVSLRRNISSVRKPMASTREC 181
Query: 72 CKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQ 118
CKA+IHI Y+ S +WV+ + DHNH VVSPRE TMD+KD I+
Sbjct: 182 CKALIHIIYNNSVRWVMPELANDHNHSSVVSPREVISTMDDKDNMIR 322
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 136 bits (343), Expect = 4e-33
Identities = 80/176 (45%), Positives = 105/176 (59%), Gaps = 6/176 (3%)
Frame = +1
Query: 2 EFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSI 61
EF SE AA FY+ YA VGF++RV RS RDG + R L CNKEG + K I
Sbjct: 4 EFGSEAAAHAFYNAYATEVGFIVRVSKLSRSRRDGTAIGRTLVCNKEG--FRMADKREKI 177
Query: 62 RRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREAR------QTMDEKDK 115
R RA TR GC+AMI ++ SGKWV+ KFVK+H HPL +P + R Q E+D
Sbjct: 178 VRQRAETRVGCRAMIMVRKLSSGKWVVAKFVKEHTHPL--TPGKGRRDFYYEQYPTEQD- 348
Query: 116 KIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEE 171
KI+EL+ ++ +K+ A Y+ L + +EE +D LS KI HIV+ +KE E EE
Sbjct: 349 KIRELSQQLATEKKRSAAYKRHLELIFEQIEEQNDNLSKKIQHIVDCVKEMEAKEE 516
>TC232091
Length = 530
Score = 124 bits (311), Expect = 2e-29
Identities = 56/102 (54%), Positives = 77/102 (74%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEF+SE+AAK FY+EYAR GFV+R+ C RSE D +I++RR CNK+G + +R K
Sbjct: 148 MEFQSEEAAKNFYEEYARREGFVVRLDRCHRSEVDKQIISRRFSCNKQGFHVRVRNKTKP 327
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVS 102
+ +PRAS REGC+AM+++K + GKWV+TKFVK+H+H L S
Sbjct: 328 VHKPRASIREGCEAMMYVKVNTCGKWVVTKFVKEHSHLLNAS 453
>TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (63%)
Length = 1326
Score = 95.5 bits (236), Expect = 1e-20
Identities = 49/92 (53%), Positives = 58/92 (62%)
Frame = +1
Query: 8 AAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRAS 67
AA FY+ YA VGF++RV RS RDG + R L CNKEG + K I R RA
Sbjct: 4 AAHAFYNAYATDVGFIVRVSKLSRSRRDGTAIGRTLVCNKEG--FRMADKREKIVRQRAE 177
Query: 68 TREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
TR GC+AMI ++ SGKWVI KFVK+H HPL
Sbjct: 178TRVGCRAMIMVRKLSSGKWVIAKFVKEHTHPL 273
Score = 46.6 bits (109), Expect = 6e-06
Identities = 22/56 (39%), Positives = 36/56 (64%)
Frame = +3
Query: 116 KIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEE 171
KI+EL+ ++ +K+ A Y+ L + +EE ++ LS KI HIV+++KE E EE
Sbjct: 798 KIRELSQQLATEKKRSAAYKRHLELIFEQIEEQNENLSKKIQHIVDSVKEMEAKEE 965
>BU578900 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (1%)
Length = 426
Score = 77.4 bits (189), Expect = 3e-15
Identities = 36/91 (39%), Positives = 57/91 (62%)
Frame = +1
Query: 77 HIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQELTAEIRIKKRLCATYQE 136
++K + GKWV+TKFVK+H+H L S ++ KD+ IQ+L E+ + RLC Y+
Sbjct: 1 YVKVNTCGKWVVTKFVKEHSHLLNASALPYNSLIESKDRIIQQLAKELEHQDRLCQHYRR 180
Query: 137 QLSSFMKIVEEHSDKLSVKIHHIVNNLKEFE 167
QL S ++ V+E + LS K+ +VN +K+ E
Sbjct: 181 QLFSLLETVDEQTKCLSTKVELVVNTVKKLE 273
>AW780793
Length = 419
Score = 70.9 bits (172), Expect = 3e-13
Identities = 35/92 (38%), Positives = 52/92 (56%), Gaps = 1/92 (1%)
Frame = +3
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG-HCLNIRGKLG 59
MEF S + A+ FY Y R +GF +R+ RRS + +++ + C+KEG +
Sbjct: 144 MEFNSREEAREFYIAYGRRIGFTVRIHHNRRSRVNNQVIGQDFVCSKEGFRAKKYLHRKD 323
Query: 60 SIRRPRASTREGCKAMIHIKYDKSGKWVITKF 91
+ P +TREGC+AMI + GKWV+TKF
Sbjct: 324 RVLPPPPATREGCQAMIRLALRDRGKWVVTKF 419
>BG507472 similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (1%)
Length = 410
Score = 62.4 bits (150), Expect = 1e-10
Identities = 35/93 (37%), Positives = 58/93 (61%), Gaps = 12/93 (12%)
Frame = +1
Query: 87 VITKFVKDHNHPLVVSPREARQ--------TMDEKDKKIQELTAEIRIK----KRLCATY 134
V+TKFVK+H H L+ SP + + DEKDK+I+EL+ E+ + KR CA Y
Sbjct: 1 VVTKFVKEHTHKLM-SPSKVPWRGSGKHLVSEDEKDKRIRELSLELYNERQKCKRRCAAY 177
Query: 135 QEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFE 167
+EQL+ + +E+H++ +S K+ +V +++E E
Sbjct: 178 EEQLNMILNDLEKHTEHISEKVAAVVRSIREIE 276
>TC229007
Length = 452
Score = 38.1 bits (87), Expect(2) = 1e-08
Identities = 16/17 (94%), Positives = 17/17 (99%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYA 17
MEFESEDAAK+FYDEYA
Sbjct: 289 MEFESEDAAKIFYDEYA 339
Score = 37.4 bits (85), Expect(2) = 1e-08
Identities = 17/21 (80%), Positives = 18/21 (84%)
Frame = +2
Query: 18 RHVGFVMRVMSCRRSERDGRI 38
R +GFVMRVMSCR SE DGRI
Sbjct: 341 RRLGFVMRVMSCRPSEGDGRI 403
>TC220852
Length = 989
Score = 54.3 bits (129), Expect = 3e-08
Identities = 34/99 (34%), Positives = 53/99 (53%), Gaps = 2/99 (2%)
Frame = +1
Query: 69 REGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREA--RQTMDEKDKKIQELTAEIRI 126
+E K + IK SG L+V+ +A Q++ EK+KKI+ELTAE+ +
Sbjct: 163 QEAAKKVCAIKNHSSGTAEGATVTNGSRGELLVADEDAPSNQSVAEKEKKIRELTAELEV 342
Query: 127 KKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
+ C Y+ L + +K +EE KLSVK+ + +LKE
Sbjct: 343 TNQRCEVYRANLLTVLKDMEEQKLKLSVKVQNARFSLKE 459
>TC220629 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (21%)
Length = 709
Score = 52.8 bits (125), Expect = 8e-08
Identities = 24/56 (42%), Positives = 39/56 (68%)
Frame = +3
Query: 116 KIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEE 171
KI+EL+ ++ I+++ ATY+ L + +EEH+D LS KI HIV+++KE E E+
Sbjct: 108 KIRELSQQLAIERKRSATYKRHLELIFEQIEEHNDSLSKKIQHIVDSVKEMETKEQ 275
>TC232186
Length = 860
Score = 51.2 bits (121), Expect = 2e-07
Identities = 24/61 (39%), Positives = 42/61 (68%), Gaps = 4/61 (6%)
Frame = +2
Query: 111 DEKDKKIQELTAEIRIK----KRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEF 166
DEKDK+I+EL+ E+ + KR CA Y+EQL+ + +E+H++ +S K+ +V +++E
Sbjct: 266 DEKDKRIRELSLELYNERQKCKRRCAAYEEQLNMILNDLEKHTEHISKKVAAVVRSIREI 445
Query: 167 E 167
E
Sbjct: 446 E 448
>AI736705 similar to GP|11994736|db far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (14%)
Length = 383
Score = 48.1 bits (113), Expect = 2e-06
Identities = 20/45 (44%), Positives = 29/45 (64%)
Frame = +3
Query: 65 RASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQT 109
R+ ++ CKA +H+K GKWVI FVK+HNH L+ + + QT
Sbjct: 66 RSCSKTDCKASMHVKRRSDGKWVIHSFVKEHNHELLPAQAVSEQT 200
>TC208764 similar to UP|Q9ZVC9 (Q9ZVC9) Mutator-like transposase
(At2g27110/T20P8.16), partial (4%)
Length = 774
Score = 44.7 bits (104), Expect = 2e-05
Identities = 21/55 (38%), Positives = 34/55 (61%)
Frame = +2
Query: 111 DEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
D+ DK I++L E+ R C Y+ L S +K VE+H +LSVK+ +I ++K+
Sbjct: 395 DDLDKNIRKLMNELECANRKCEIYRSNLLSVLKAVEDHKLELSVKVENIKISMKD 559
>AW099292
Length = 420
Score = 41.6 bits (96), Expect = 2e-04
Identities = 20/49 (40%), Positives = 26/49 (52%)
Frame = +2
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG 49
MEFES +AA FY EYA+ GF +S RRS + + C + G
Sbjct: 272 MEFESHEAAYAFYKEYAKSAGFGTAKLSSRRSRASKEFIDAKFSCIRYG 418
>TC208765
Length = 589
Score = 38.5 bits (88), Expect = 0.002
Identities = 21/59 (35%), Positives = 34/59 (57%), Gaps = 1/59 (1%)
Frame = +3
Query: 108 QTMDEKD-KKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
Q M + D I++L E+ R C Y+ L S +K VE+H +LSVK+ +I ++K+
Sbjct: 144 QHMSKDDLDNIRKLMNELECANRKCEIYRSNLLSVLKAVEDHKLELSVKVENIKISMKD 320
>BI700862 weakly similar to GP|16604382|gb| AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (14%)
Length = 422
Score = 35.8 bits (81), Expect = 0.011
Identities = 13/17 (76%), Positives = 15/17 (87%)
Frame = +2
Query: 83 SGKWVITKFVKDHNHPL 99
SGKWVITKF+ +H HPL
Sbjct: 14 SGKWVITKFIMEHTHPL 64
>BE022135
Length = 407
Score = 35.0 bits (79), Expect = 0.018
Identities = 15/20 (75%), Positives = 17/20 (85%)
Frame = +3
Query: 1 MEFESEDAAKVFYDEYARHV 20
MEFESE+AAK FY+ YAR V
Sbjct: 348 MEFESEEAAKAFYNSYARRV 407
>TC217771
Length = 757
Score = 27.3 bits (59), Expect(2) = 0.48
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = +2
Query: 98 PLVVSPREARQTMDEKDKKIQELTAEIRIKK 128
PL++S Q + E+D+KIQE + R+ K
Sbjct: 497 PLLLSSNNMDQMLKERDRKIQEYSRRARMYK 589
Score = 21.6 bits (44), Expect(2) = 0.48
Identities = 10/22 (45%), Positives = 11/22 (49%)
Frame = +1
Query: 74 AMIHIKYDKSGKWVITKFVKDH 95
A IH K W IT F+K H
Sbjct: 457 AWIHGKQYLPPLWSITAFIKQH 522
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,694,375
Number of Sequences: 63676
Number of extensions: 97974
Number of successful extensions: 515
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 506
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 507
length of query: 176
length of database: 12,639,632
effective HSP length: 91
effective length of query: 85
effective length of database: 6,845,116
effective search space: 581834860
effective search space used: 581834860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0083.5


