

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.16
(294 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
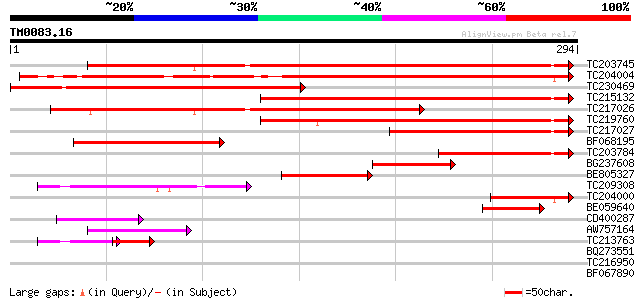
Score E
Sequences producing significant alignments: (bits) Value
TC203745 weakly similar to UP|O88972 (O88972) KH domain RNA bind... 283 5e-77
TC204004 similar to UP|Q8W574 (Q8W574) At1g09660/F21M12_5, parti... 265 2e-71
TC230469 similar to GB|BAB09296.1|9758634|AB011476 RNA-binding p... 258 2e-69
TC215132 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partia... 223 1e-58
TC217026 weakly similar to UP|O88972 (O88972) KH domain RNA bind... 196 1e-50
TC219760 weakly similar to UP|O88972 (O88972) KH domain RNA bind... 196 1e-50
TC217027 weakly similar to UP|O88972 (O88972) KH domain RNA bind... 141 3e-34
BF068195 134 5e-32
TC203784 95 3e-20
BG237608 homologue to PIR|T01782|T017 GDP dissociation inhibitor... 67 1e-11
BE805327 similar to GP|20161243|dbj P0408G07.16 {Oryza sativa (j... 64 1e-10
TC209308 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partia... 59 3e-09
TC204000 homologue to UP|Q84XC3 (Q84XC3) Ubiquitin/ribosomal fus... 57 8e-09
BE059640 46 2e-05
CD400287 similar to PIR|S04125|S041 chlorophyll a/b-binding prot... 45 5e-05
AW757164 44 9e-05
TC213763 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partia... 38 5e-04
BQ273551 38 0.006
TC216950 32 0.26
BF067890 similar to GP|9651915|gb|A 4-coumarate:coA ligase 2 {Ru... 31 0.76
>TC203745 weakly similar to UP|O88972 (O88972) KH domain RNA binding protein
QKI-7B, partial (20%)
Length = 1338
Score = 283 bits (725), Expect = 5e-77
Identities = 150/255 (58%), Positives = 180/255 (69%), Gaps = 3/255 (1%)
Frame = +3
Query: 41 KYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTLLGNASVLGQSRLEHASP--LATG 98
+YLTELL E LGPFM LP C RL+NQEILRV+ +L N RL H SP +A+
Sbjct: 156 QYLTELLAEHQKLGPFMQALPICSRLLNQEILRVSGMLSNQGFGDFDRLRHRSPSPMASS 335
Query: 99 GIFSN-GGADVNGWASRFQSEMPSLLQSSATPSWLSPQGSSSGLVVKKTMRVDIPVEVYP 157
+ S+ G + GW S Q + W S S VK+ +R++IPV+ YP
Sbjct: 336 NLMSSVTGTGLGGWNSLQQERLRGT--PGMAMDWQVAPASPSSYTVKRILRLEIPVDTYP 509
Query: 158 NFNFVGRLLGPRGNSLKRVEASTDCRVLIRGRGSIKDPAKEEMMRGKPGYEHLNEPLHVL 217
NFNFVGRLLGPRGNSLKRVEAST CRV IRG+GSIKDP KEE +RG+PGYEHLNE LH+L
Sbjct: 510 NFNFVGRLLGPRGNSLKRVEASTGCRVYIRGKGSIKDPDKEEKLRGRPGYEHLNEQLHIL 689
Query: 218 VEAEFPAEIIDARLMQAREILEDLLKPVDESQDYYKKQQLRELALLNGTLREEGSPMSGS 277
+EA+ PA ++D RL QA+EI+E+LLKPV+ES+DY K+QQLRELA+LN REE SGS
Sbjct: 690 IEADLPANVVDLRLRQAQEIIEELLKPVEESEDYIKRQQLRELAMLNSNFREESPGPSGS 869
Query: 278 VSPFHNSLGMKRAKT 292
VSPF NS GMKRAKT
Sbjct: 870 VSPF-NSSGMKRAKT 911
>TC204004 similar to UP|Q8W574 (Q8W574) At1g09660/F21M12_5, partial (62%)
Length = 1255
Score = 265 bits (676), Expect = 2e-71
Identities = 155/290 (53%), Positives = 192/290 (65%), Gaps = 3/290 (1%)
Frame = +1
Query: 6 GAGRYMAFPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYR 65
G+G Y FPP PPSP +R ++SS+ ++D+YL ELL ER L PF+ VLP +
Sbjct: 109 GSGMYFPFPP----PPSP----IRPSSSSS--DRDRYLAELLAERQKLVPFLQVLPQSTK 258
Query: 66 LINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMPSLLQS 125
L+ QEI R++ G G H T + D+ GWA Q + P+ +
Sbjct: 259 LLTQEIRRMSVGGGG----GGGGFNHEPAADTPPPYFRP-MDLEGWAIEVQQDKPNPQRM 423
Query: 126 SATPSWLSPQGSSSGLVVKKTMRVDIPVEVYPNFNFVGRLLGPRGNSLKRVEASTDCRVL 185
A W +P VVK+ +R+D+PV+ +PN+NFVGR+LGPRGNSLKRVEA T+CRV
Sbjct: 424 MA---WPAP-------VVKRVIRLDVPVDKFPNYNFVGRILGPRGNSLKRVEAMTECRVY 573
Query: 186 IRGRGSIKDPAKEEMMRGKPGYEHLNEPLHVLVEAEFPAEIIDARLMQAREILEDLLKPV 245
IRG GS+KD KEE ++ KPGYEHL EPLHVLVEAEFP +II+ARL A ILE+LLKPV
Sbjct: 574 IRGCGSVKDSIKEEKLKEKPGYEHLKEPLHVLVEAEFPEDIINARLDHAVAILENLLKPV 753
Query: 246 DESQDYYKKQQLRELALLNGTLREEGSPMSGSVSPF---HNSLGMKRAKT 292
DES D+YKKQQLRELA+LNGTLREE MS S+SP NS GMKRAKT
Sbjct: 754 DESLDHYKKQQLRELAMLNGTLREESPSMSPSMSPSMSPFNSTGMKRAKT 903
>TC230469 similar to GB|BAB09296.1|9758634|AB011476 RNA-binding protein-like
{Arabidopsis thaliana;} , partial (32%)
Length = 476
Score = 258 bits (660), Expect = 2e-69
Identities = 129/153 (84%), Positives = 141/153 (91%)
Frame = +3
Query: 1 MMSSSGAGRYMAFPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVL 60
MMSSSGA RYMAFPPSPSAPPSPHLSG + S+ L++ DKYLTELLGER+ L PFMAVL
Sbjct: 24 MMSSSGAARYMAFPPSPSAPPSPHLSG--ALRSTPLSDPDKYLTELLGERNKLSPFMAVL 197
Query: 61 PHCYRLINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMP 120
PHC+RL+NQEILRVTTL+GNASVLGQS LEHASPLATGGIFSNGGADVNGWASRFQSE P
Sbjct: 198 PHCFRLLNQEILRVTTLMGNASVLGQSGLEHASPLATGGIFSNGGADVNGWASRFQSERP 377
Query: 121 SLLQSSATPSWLSPQGSSSGLVVKKTMRVDIPV 153
SLLQSS+T SWLSPQGSSSG++VKKT+RVDIPV
Sbjct: 378 SLLQSSSTQSWLSPQGSSSGIIVKKTVRVDIPV 476
>TC215132 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partial (55%)
Length = 753
Score = 223 bits (567), Expect = 1e-58
Identities = 108/162 (66%), Positives = 130/162 (79%)
Frame = +3
Query: 131 WLSPQGSSSGLVVKKTMRVDIPVEVYPNFNFVGRLLGPRGNSLKRVEASTDCRVLIRGRG 190
W + S +VKK +R+DIP + YPNFNFVGRLLGPRGNSLKRVEA+T CRV IRG+G
Sbjct: 51 WQTSPNVPSSPIVKKILRLDIPKDSYPNFNFVGRLLGPRGNSLKRVEATTGCRVFIRGKG 230
Query: 191 SIKDPAKEEMMRGKPGYEHLNEPLHVLVEAEFPAEIIDARLMQAREILEDLLKPVDESQD 250
SIKD KEEM+RG+PGYEHLN+PLH+++EAE P + D RLMQA+EI+++LLKPVDESQD
Sbjct: 231 SIKDLDKEEMLRGRPGYEHLNDPLHIIIEAELPTSVADVRLMQAQEIIQELLKPVDESQD 410
Query: 251 YYKKQQLRELALLNGTLREEGSPMSGSVSPFHNSLGMKRAKT 292
YK+QQLRELA+LN REE +SGSVSPF S +KR KT
Sbjct: 411 LYKRQQLRELAMLNSNFREESPQLSGSVSPF-TSNEIKRVKT 533
>TC217026 weakly similar to UP|O88972 (O88972) KH domain RNA binding protein
QKI-7B, partial (19%)
Length = 696
Score = 196 bits (498), Expect = 1e-50
Identities = 106/200 (53%), Positives = 131/200 (65%), Gaps = 6/200 (3%)
Frame = +2
Query: 22 SPHLSGLRSAASSALAEQD---KYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTLL 78
+P+ S R+A+ + + +YL+ELL E LGPFM VLP C RL+NQEILRV+ +L
Sbjct: 101 NPNFSPARAASPQIRSNPEVDSQYLSELLAEHQKLGPFMQVLPICSRLLNQEILRVSGML 280
Query: 79 GNASVLGQSRLEHASP--LATGGIFSN-GGADVNGWASRFQSEMPSLLQSSATPSWLSPQ 135
N RL H SP +A+ + SN G + GW S Q + T W S
Sbjct: 281 SNQGFGDFDRLRHRSPSPMASSNLMSNVSGTGLGGWNSLQQERLCGA--PGMTMDWQSAP 454
Query: 136 GSSSGLVVKKTMRVDIPVEVYPNFNFVGRLLGPRGNSLKRVEASTDCRVLIRGRGSIKDP 195
S S VK+ +R++IPV+ YPNFNFVGRLLGPRGNSLKRVEA+T CRV IRG+GSIKDP
Sbjct: 455 ASPSSFTVKRILRLEIPVDTYPNFNFVGRLLGPRGNSLKRVEATTGCRVYIRGKGSIKDP 634
Query: 196 AKEEMMRGKPGYEHLNEPLH 215
KEE +RG+PGYEHLNEPLH
Sbjct: 635 DKEEKLRGRPGYEHLNEPLH 694
>TC219760 weakly similar to UP|O88972 (O88972) KH domain RNA binding protein
QKI-7B, partial (18%)
Length = 957
Score = 196 bits (497), Expect = 1e-50
Identities = 107/205 (52%), Positives = 129/205 (62%), Gaps = 43/205 (20%)
Frame = +1
Query: 131 WLSPQGSSSGLVVKKTMRVDIPVEVYPN-------------------------------- 158
W + G S +VK+T+R+DI + YPN
Sbjct: 28 WQTSPGVPSSHIVKRTLRLDIANDSYPNVRRYVEHFNAFCCILMKIYTMHCSNSFMVLY* 207
Query: 159 -----------FNFVGRLLGPRGNSLKRVEASTDCRVLIRGRGSIKDPAKEEMMRGKPGY 207
FN VGRLLGPRGNSLKRVEA+T CRV IRG+GSIK+ KEE++RG+PGY
Sbjct: 208 ALRH*A*LKFQFNLVGRLLGPRGNSLKRVEATTGCRVFIRGKGSIKELDKEELLRGRPGY 387
Query: 208 EHLNEPLHVLVEAEFPAEIIDARLMQAREILEDLLKPVDESQDYYKKQQLRELALLNGTL 267
EHLNEPLHVL+EAE P ++D RL QA+EI+E+LLKP+DESQD YK+QQLRELA+LN
Sbjct: 388 EHLNEPLHVLIEAELPVNVVDIRLRQAQEIIEELLKPMDESQDLYKRQQLRELAMLNSNF 567
Query: 268 REEGSPMSGSVSPFHNSLGMKRAKT 292
REE +S S S F NS MKRAKT
Sbjct: 568 REESPQLSASPSTF-NSNEMKRAKT 639
>TC217027 weakly similar to UP|O88972 (O88972) KH domain RNA binding protein
QKI-7B, partial (6%)
Length = 921
Score = 141 bits (356), Expect = 3e-34
Identities = 69/95 (72%), Positives = 80/95 (83%)
Frame = +2
Query: 198 EEMMRGKPGYEHLNEPLHVLVEAEFPAEIIDARLMQAREILEDLLKPVDESQDYYKKQQL 257
EE +RG+PGYEHLNEPLH+L+EAE PA ++D RL QA+EI+E+LLKPVDESQDY K+QQL
Sbjct: 2 EEKLRGRPGYEHLNEPLHILIEAELPANVVDIRLRQAQEIIEELLKPVDESQDYIKRQQL 181
Query: 258 RELALLNGTLREEGSPMSGSVSPFHNSLGMKRAKT 292
RELA+LN REE SGSVSPF NS GMKRAKT
Sbjct: 182 RELAMLNSNFREESPGPSGSVSPF-NSSGMKRAKT 283
>BF068195
Length = 365
Score = 134 bits (337), Expect = 5e-32
Identities = 65/78 (83%), Positives = 71/78 (90%)
Frame = +1
Query: 34 SALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTLLGNASVLGQSRLEHAS 93
+AL++ DKYL ELLGER+ L FMAVLPHC+RL NQEILRVTTL+GNASVLGQS LEHAS
Sbjct: 130 NALSDPDKYLAELLGERNKLSRFMAVLPHCFRLFNQEILRVTTLMGNASVLGQSGLEHAS 309
Query: 94 PLATGGIFSNGGADVNGW 111
PLATGGIFSNGGADVNGW
Sbjct: 310 PLATGGIFSNGGADVNGW 363
>TC203784
Length = 787
Score = 95.1 bits (235), Expect = 3e-20
Identities = 49/70 (70%), Positives = 57/70 (81%)
Frame = +2
Query: 223 PAEIIDARLMQAREILEDLLKPVDESQDYYKKQQLRELALLNGTLREEGSPMSGSVSPFH 282
PA I+D RL QA+EI+E+LLKPV+ES+DY K+QQLRELA+LN REE SGSVSPF
Sbjct: 2 PANIVDIRLRQAQEIIEELLKPVEESEDYIKRQQLRELAMLNSNFREESPGPSGSVSPF- 178
Query: 283 NSLGMKRAKT 292
NS GMKRAKT
Sbjct: 179 NSSGMKRAKT 208
>BG237608 homologue to PIR|T01782|T017 GDP dissociation inhibitor - common
tobacco, partial (13%)
Length = 432
Score = 66.6 bits (161), Expect = 1e-11
Identities = 28/43 (65%), Positives = 35/43 (81%)
Frame = +2
Query: 189 RGSIKDPAKEEMMRGKPGYEHLNEPLHVLVEAEFPAEIIDARL 231
+GSIKD KE M+RGKPGYEHLN+PLH+++EAE P + D RL
Sbjct: 302 KGSIKDLDKE*MLRGKPGYEHLNDPLHIIIEAELPTSVADVRL 430
>BE805327 similar to GP|20161243|dbj P0408G07.16 {Oryza sativa (japonica
cultivar-group)}, partial (17%)
Length = 145
Score = 63.5 bits (153), Expect = 1e-10
Identities = 27/47 (57%), Positives = 37/47 (78%)
Frame = +2
Query: 142 VVKKTMRVDIPVEVYPNFNFVGRLLGPRGNSLKRVEASTDCRVLIRG 188
VVK+ + +D+PV+ +PN+N+VGR+LGP GN LKRVEA T+C I G
Sbjct: 5 VVKRVIMLDVPVDKFPNYNYVGRILGPGGNFLKRVEAMTECMAHITG 145
>TC209308 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partial (14%)
Length = 943
Score = 58.9 bits (141), Expect = 3e-09
Identities = 47/116 (40%), Positives = 57/116 (48%), Gaps = 5/116 (4%)
Frame = +2
Query: 15 PSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQEILRV 74
PSP SP+++ +YLTELL ER LGPFM VLP C RLINQEILRV
Sbjct: 281 PSPQRANSPNIN-----MRGNFDVDSQYLTELLAERQKLGPFMQVLPLCTRLINQEILRV 445
Query: 75 T---TLLGNA--SVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMPSLLQS 125
T LL N S + R + S +A+ N + GW S S+L S
Sbjct: 446 TGKNELLQNQGFSDFDRMRFINLSHMAS----PNSTPNFTGWNSLSHEVRTSVLCS 601
>TC204000 homologue to UP|Q84XC3 (Q84XC3) Ubiquitin/ribosomal fusion protein,
partial (56%)
Length = 1061
Score = 57.4 bits (137), Expect = 8e-09
Identities = 31/46 (67%), Positives = 35/46 (75%), Gaps = 3/46 (6%)
Frame = +2
Query: 250 DYYKKQQLRELALLNGTLREEGSPMSGSVSPF---HNSLGMKRAKT 292
D+YKKQ LRELA+LNGTLREE MS S+SP NS GMKRA+T
Sbjct: 596 DHYKKQPLRELAMLNGTLREESPSMSPSMSPSMSPFNSTGMKRAQT 733
>BE059640
Length = 165
Score = 45.8 bits (107), Expect = 2e-05
Identities = 22/32 (68%), Positives = 24/32 (74%)
Frame = +1
Query: 246 DESQDYYKKQQLRELALLNGTLREEGSPMSGS 277
DESQDY K+QQLRELA+LN REE SGS
Sbjct: 70 DESQDYIKRQQLRELAMLNSNFREESPGPSGS 165
>CD400287 similar to PIR|S04125|S041 chlorophyll a/b-binding protein type III
precursor - tomato, partial (21%)
Length = 599
Score = 44.7 bits (104), Expect = 5e-05
Identities = 22/45 (48%), Positives = 26/45 (56%)
Frame = +1
Query: 25 LSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQ 69
L GL ++ +YLTELL E LGPFM LP C RL+NQ
Sbjct: 463 LQGLLTSCQFGTRVDSQYLTELLAEHQKLGPFMQALPICSRLLNQ 597
>AW757164
Length = 388
Score = 43.9 bits (102), Expect = 9e-05
Identities = 22/54 (40%), Positives = 32/54 (58%)
Frame = +2
Query: 41 KYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTLLGNASVLGQSRLEHASP 94
+YL+ +L E LGPFM VL C RL+ Q++ RV+ +L + +L H SP
Sbjct: 80 QYLSHVLAEHRNLGPFMQVLSICMRLLRQKMWRVSGMLDHQRCGDFDKLRHRSP 241
>TC213763 similar to UP|Q75GR5 (Q75GR5) Expressed protein, partial (13%)
Length = 616
Score = 38.1 bits (87), Expect(2) = 5e-04
Identities = 17/22 (77%), Positives = 19/22 (86%)
Frame = +3
Query: 54 GPFMAVLPHCYRLINQEILRVT 75
GPFM VLP C RL+NQEILRV+
Sbjct: 498 GPFMQVLPLCTRLLNQEILRVS 563
Score = 22.3 bits (46), Expect(2) = 5e-04
Identities = 15/44 (34%), Positives = 21/44 (47%)
Frame = +2
Query: 15 PSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMA 58
PS + SP+++ S + +YLTELL E L F A
Sbjct: 395 PSTARANSPNIN-----MRSNFEGESQYLTELLAEHQKLWTFHA 511
>BQ273551
Length = 424
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/46 (41%), Positives = 25/46 (54%)
Frame = +2
Query: 18 SAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHC 63
S+P + ++ S + +YLTELL E LGPFM VLP C
Sbjct: 287 SSPSTARVNSPNINMRSNFEAESQYLTELLAEHQKLGPFMQVLPLC 424
>TC216950
Length = 873
Score = 32.3 bits (72), Expect = 0.26
Identities = 28/95 (29%), Positives = 49/95 (51%), Gaps = 8/95 (8%)
Frame = +2
Query: 173 LKRVEASTDCRVLIRGRGSIKDPAKEEMM------RGKPGYEHLNEPLHVLVEAEFPAEI 226
LK +E T RV R +++ K E + R + G + LN+ + +E E A +
Sbjct: 221 LKLIEEETAKRVEEAIRKRVEESLKSEEVQVEIERRLEEGRKRLNDEVAAQLEKEKEAAL 400
Query: 227 IDARLM--QAREILEDLLKPVDESQDYYKKQQLRE 259
I+A+ QAR+ EDL + ++E++ ++ Q RE
Sbjct: 401 IEAKQKEEQARKEKEDLERMLEENRRKVEEAQRRE 505
>BF067890 similar to GP|9651915|gb|A 4-coumarate:coA ligase 2 {Rubus idaeus},
partial (8%)
Length = 353
Score = 30.8 bits (68), Expect = 0.76
Identities = 22/83 (26%), Positives = 34/83 (40%)
Frame = +1
Query: 101 FSNGGADVNGWASRFQSEMPSLLQSSATPSWLSPQGSSSGLVVKKTMRVDIPVEVYPNFN 160
F++ A VN + + F S L S TP W S S L ++P ++PN
Sbjct: 61 FNSAKAYVNNFPTFFSSISHETLTFSTTPKWKSNNLHSLSLTTTSFSVPNLPGHLHPNAP 240
Query: 161 FVGRLLGPRGNSLKRVEASTDCR 183
L P+ +++R CR
Sbjct: 241 PSPHLPFPKPLTIQRPSMPHQCR 309
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,218,164
Number of Sequences: 63676
Number of extensions: 170231
Number of successful extensions: 2119
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 1815
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2097
length of query: 294
length of database: 12,639,632
effective HSP length: 97
effective length of query: 197
effective length of database: 6,463,060
effective search space: 1273222820
effective search space used: 1273222820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0083.16


