

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.15
(489 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
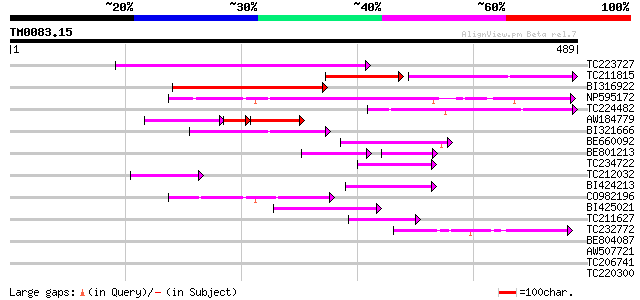
Score E
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 167 1e-41
TC211815 76 3e-27
BI316922 118 7e-27
NP595172 polyprotein [Glycine max] 112 5e-25
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 103 2e-22
AW184779 70 1e-21
BI321666 77 1e-14
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 76 4e-14
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 50 2e-11
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 59 4e-09
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 58 8e-09
BI424213 54 2e-07
CO982196 50 2e-06
BI425021 47 1e-05
TC211627 45 6e-05
TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial ... 42 8e-04
BE804087 39 0.005
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 38 0.009
TC206741 36 0.034
TC220300 similar to UP|Q9LZA5 (Q9LZA5) Maturase-like protein, pa... 33 0.38
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 167 bits (422), Expect = 1e-41
Identities = 84/222 (37%), Positives = 129/222 (57%), Gaps = 2/222 (0%)
Frame = +1
Query: 92 WLPEEKAEAKKLVRR-ASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHH 150
+LPE K+ +RR A+ + + L+KR L+C+ A +++ E+HEGS G H
Sbjct: 178 YLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTH 357
Query: 151 LGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQW 210
G ++ARK+LRAGYYW T+E D HV++C CQ AD +APP L+ + SPWPF W
Sbjct: 358 ANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMW 537
Query: 211 GMDLLGPFD-TAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGV 269
G+D++G + A ++++VA+DY+TKW+EA + V RF K +I R+G+P
Sbjct: 538 GIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRK 717
Query: 270 LITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESA 311
+ITDNGT +K ++ + I+ + P+ N E A
Sbjct: 718 IITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>TC211815
Length = 704
Score = 76.3 bits (186), Expect(2) = 3e-27
Identities = 36/67 (53%), Positives = 46/67 (67%)
Frame = +3
Query: 273 DNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAE 332
DNG QFT++ + GL+IK R TSV+HPQTN +AE+AN+VIL L+K L A G W E
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLLDGANGRWVE 182
Query: 333 QLDHVLW 339
L +LW
Sbjct: 183 DLVEILW 203
Score = 63.9 bits (154), Expect(2) = 3e-27
Identities = 44/145 (30%), Positives = 66/145 (45%)
Frame = +2
Query: 345 PHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGEDLDLLAERRAIANCR 404
P STT E+P+ L +G ++P E+ E F N + L DLDL+ + R
Sbjct: 227 PQSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIM 406
Query: 405 ELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTG 464
KQ +N K+ G ++ R K EGP+++ + G
Sbjct: 407 T*AFKQRMTRCFNSKLPSTV*GRGPS-MEGIQRSLEVLVRSKFTTN*EGPFKIRHNSKNG 583
Query: 465 AYKLETLAGKEIPRTWNATNLRRYY 489
AYKLE L+GK + R WN+ +L+ YY
Sbjct: 584 AYKLEELSGKVVLRIWNSMHLKVYY 658
>BI316922
Length = 405
Score = 118 bits (295), Expect = 7e-27
Identities = 54/134 (40%), Positives = 85/134 (63%)
Frame = +3
Query: 141 EIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSS 200
E+H G CG H + + +VLR GYY T+ K ++VK+C+ C + ++ H L +
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 201 LVSPWPFHQWGMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNV 260
+V+PWPF G+D+L PF + +KYL+V +D +TKWIE E +A I+ A V++F +N+
Sbjct: 183 IVAPWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNI 362
Query: 261 IRRFGVPGVLITDN 274
+ FG+P LI+DN
Sbjct: 363 VC*FGIPNTLISDN 404
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 112 bits (279), Expect = 5e-25
Identities = 101/370 (27%), Positives = 165/370 (44%), Gaps = 19/370 (5%)
Frame = +1
Query: 138 VLAEIHEGSCGHHLG-GRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPA 196
+L E H G H G R+LAR L+A +YWP +++D ++++C CQ+ + P
Sbjct: 3220 ILQEYHSSPIGGHAGITRTLAR--LKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAG 3393
Query: 197 RLSSLVSPWPFHQW---GMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPL-ATITSARV 252
L L P P W MD + + G L ++V +D TK+ PL A S V
Sbjct: 3394 LLQPL--PIPQQVWEDVAMDFITGLPNSFG-LSVIMVVIDRLTKYAHFIPLKADYNSKVV 3564
Query: 253 QRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESAN 312
F ++++ G+P +++D FTS ++ L +S HPQ++GQ+E N
Sbjct: 3565 AEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLN 3744
Query: 313 RVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVI-------- 364
+ + LR + W + L + Y T H + G +P+R +G E
Sbjct: 3745 KCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGREPPTLTRQACSI 3924
Query: 365 --PAEIREPSARTAGFDLNQNDQLLGEDLDLLAERRAIANCRELIAKQEAAIRYNKKVVL 422
PAE+RE QL D LLA+ + + + K++A +KK +
Sbjct: 3925 DDPAEVRE--------------QLTDRDA-LLAKLKINLTRAQQVMKRQA----DKKRLD 4047
Query: 423 RSFAAGDLVLQN----ASIGARSTERGKLAAKWEGPYRVVEATGTGAYKLETLAGKEIPR 478
SF GD VL A + KL+ ++ GP++V+ G AYKLE + I
Sbjct: 4048 VSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHP 4227
Query: 479 TWNATNLRRY 488
++ + L+ +
Sbjct: 4228 VFHVSQLKPF 4257
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 103 bits (257), Expect = 2e-22
Identities = 65/184 (35%), Positives = 99/184 (53%), Gaps = 3/184 (1%)
Frame = +1
Query: 309 ESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEI 368
E+AN+ I + ++K K DW E L L YRT+ ++TG +P+ L +G EAV+P E+
Sbjct: 1 EAANKNIKKIIQKMTVSYK-DWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 369 REPSAR---TAGFDLNQNDQLLGEDLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSF 425
PS R +G ++ Q + L+L+ +R A + +Q ++KKV LR F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 426 AAGDLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNL 485
GDLVL+ S + RGK A +EGP+ V A GA L + G+E+P N+ +
Sbjct: 358 HEGDLVLKKMSHAVKD-HRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVV 534
Query: 486 RRYY 489
+RYY
Sbjct: 535 KRYY 546
>AW184779
Length = 432
Score = 70.1 bits (170), Expect(3) = 1e-21
Identities = 31/69 (44%), Positives = 42/69 (59%)
Frame = +1
Query: 117 LFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAAD 176
L+KR LL+C+ A +L E+HEGS G H ++A+K+LR GYYW T+E D
Sbjct: 16 LYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDCCI 195
Query: 177 HVKQCDPCQ 185
HV +C CQ
Sbjct: 196 HVWKCHKCQ 222
Score = 43.9 bits (102), Expect(3) = 1e-21
Identities = 19/48 (39%), Positives = 32/48 (66%), Gaps = 1/48 (2%)
Frame = +3
Query: 208 HQWGMDLLGPFDT-APG*LKYLVVAVDYYTKWIEAEPLATITSARVQR 254
H WG+D++G + A +++VA+DY+TKW+EA A++T + V R
Sbjct: 288 HMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
Score = 27.3 bits (59), Expect(3) = 1e-21
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +2
Query: 185 QRHADLHHAPPARLSSLVSPWPF 207
Q AD +APP L+ L +PWP+
Sbjct: 224 QTFADNVNAPPIPLNVLAAPWPY 292
>BI321666
Length = 430
Score = 77.4 bits (189), Expect = 1e-14
Identities = 40/121 (33%), Positives = 66/121 (54%)
Frame = +2
Query: 156 LARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLL 215
++ VL++ +Y P++ KDA H + C+ CQR + L +++ F WG+D +
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFV 181
Query: 216 GPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNG 275
GPF + +Y++V VDY +KW+EA + V +F K + R GVP VLI + G
Sbjct: 182 GPFPPSFS-NEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGG 358
Query: 276 T 276
+
Sbjct: 359 S 361
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 75.9 bits (185), Expect = 4e-14
Identities = 40/100 (40%), Positives = 57/100 (57%), Gaps = 3/100 (3%)
Frame = -3
Query: 286 LLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTP 345
LL + R ++ HPQTNGQAE +NR I R L K + ++ DW+ +LD LWA+RT
Sbjct: 367 LLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAY 188
Query: 346 HSTTGESPYRLTFGTEAVIPAEIREP---SARTAGFDLNQ 382
+ G SPYR+ FG +P EI + +T F ++Q
Sbjct: 187 KAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQ 68
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 50.1 bits (118), Expect(2) = 2e-11
Identities = 23/61 (37%), Positives = 36/61 (58%)
Frame = +3
Query: 252 VQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESA 311
V +F KN+ RFG+P +LI+D G+ F +L ++ + + HPQTNGQA+ +
Sbjct: 18 VIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNGQAKVS 197
Query: 312 N 312
N
Sbjct: 198 N 200
Score = 36.6 bits (83), Expect(2) = 2e-11
Identities = 12/49 (24%), Positives = 28/49 (56%)
Frame = +2
Query: 321 KRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIR 369
K + ++ DW+ +L+ LWA +T + G +P+++ + +P E++
Sbjct: 227 KNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELK 373
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 59.3 bits (142), Expect = 4e-09
Identities = 28/68 (41%), Positives = 40/68 (58%)
Frame = -3
Query: 301 HPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGT 360
HPQTNGQAE +N+ I R L + ++ DWA +LD WAYR + G SP++L +G
Sbjct: 231 HPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYGK 52
Query: 361 EAVIPAEI 368
+ E+
Sbjct: 51 ACHLSVEL 28
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 58.2 bits (139), Expect = 8e-09
Identities = 27/63 (42%), Positives = 37/63 (57%)
Frame = +2
Query: 105 RRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAG 164
R A+ + + L+KR LL C+ +L E+HEGS G H G ++ARK+LRAG
Sbjct: 614 RLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFGTHSNGHAMARKILRAG 793
Query: 165 YYW 167
YYW
Sbjct: 794 YYW 802
>BI424213
Length = 426
Score = 53.9 bits (128), Expect = 2e-07
Identities = 28/79 (35%), Positives = 41/79 (51%)
Frame = +1
Query: 290 LHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTT 349
L K F++ HPQT+GQ + NR + LR L W E L HV +AY H TT
Sbjct: 34 LGTKLLFSTTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEFAYNRGVHRTT 213
Query: 350 GESPYRLTFGTEAVIPAEI 368
+SP+ + +G + P ++
Sbjct: 214 KQSPFEVVYGFNPLTPLDL 270
>CO982196
Length = 812
Score = 50.1 bits (118), Expect = 2e-06
Identities = 42/147 (28%), Positives = 64/147 (42%), Gaps = 4/147 (2%)
Frame = +1
Query: 138 VLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPAR 197
+L E+ + G H G ++V +W ++K D+V C+ C+R+ +P
Sbjct: 376 LLKELQDSPLGGHSGFFRTFKRVANV-VFWQGMKKTTRDYVAACEICRRNKTSTLSPAGL 552
Query: 198 LSSLVSPWPFHQW---GMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLA-TITSARVQ 253
L L P P W MD +G A G LVV VD TK+ L+ T+ V
Sbjct: 553 L*LL--PIPTKVWTDISMDFIGGLPKAQGKDNILVV-VDRLTKYAHFFALSHPYTAKEVA 723
Query: 254 RFFYKNVIRRFGVPGVLITDNGTQFTS 280
F K ++R G P +++D F S
Sbjct: 724 ELFIKELVRLHGFPASIVSDXXRLFMS 804
>BI425021
Length = 426
Score = 47.4 bits (111), Expect = 1e-05
Identities = 33/94 (35%), Positives = 47/94 (49%), Gaps = 1/94 (1%)
Frame = -1
Query: 228 LVVAVDYYTKWIEAEPLATITSAR-VQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDL 286
+ V V+ ++K I L T +A V F VI+ G P +++D F S ++DL
Sbjct: 336 IFVVVNRFSKGIHLGTLPTSHTAHMVASLFLNIVIKLHGFPRSIVSDRDPLFISHFWQDL 157
Query: 287 LDGLHIKQRFTSVEHPQTNGQAESANRVILRGLR 320
R +S HPQT+GQ E NRVI + LR
Sbjct: 156 FRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQYLR 55
>TC211627
Length = 1034
Score = 45.4 bits (106), Expect = 6e-05
Identities = 24/62 (38%), Positives = 34/62 (54%)
Frame = +2
Query: 293 KQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGES 352
K RF++ HPQT+GQ E NR++ + LR + D W + L Y T+ HS G S
Sbjct: 50 KLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSLAE*CYNTSVHSGIGFS 229
Query: 353 PY 354
P+
Sbjct: 230 PF 235
>TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (7%)
Length = 729
Score = 41.6 bits (96), Expect = 8e-04
Identities = 44/157 (28%), Positives = 70/157 (44%), Gaps = 3/157 (1%)
Frame = +1
Query: 332 EQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGEDL 391
E L HV +AY HSTT P+ + + + P + P + +GF +++ E +
Sbjct: 1 ECLPHVEFAYNRAVHSTTQHFPFEVVYDFNPLTPLDSL-PLSNISGFK-HKDAHAKVEYI 174
Query: 392 DLLAE--RRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTER-GKLA 448
L E + IA E KQ R KKVVL D V + +R KL
Sbjct: 175 KRLHEQAKTQIAKKNESYVKQTNKNR--KKVVLEP---SDWVWVHMRKERFPKQRMSKLQ 339
Query: 449 AKWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNL 485
+ +GP++V+E AYK++ E+ ++N +L
Sbjct: 340 PRGDGPFQVLERINYNAYKIDIPGEYEVSSSFNIADL 450
>BE804087
Length = 160
Score = 38.9 bits (89), Expect = 0.005
Identities = 21/54 (38%), Positives = 29/54 (52%)
Frame = +2
Query: 309 ESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEA 362
E+AN+ I + + K K DW E L YRT ++TG +PY L +G EA
Sbjct: 2 EAANKNIKKNI*KMTVSYK-DWHEMFSFALHMYRTLVRTSTGATPYSLVYGKEA 160
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 38.1 bits (87), Expect = 0.009
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Frame = +1
Query: 138 VLAEIHEGSCGHHLGG-RSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPA 196
++ E H G H G R++ R + A +YWP + +D V++C CQ+ H PA
Sbjct: 280 IMTESHASKVGGHAGTTRTIVR--INAQFYWPKMREDIMKFVQECVICQQAKVTHSLLPA 453
Query: 197 RL 198
L
Sbjct: 454 GL 459
>TC206741
Length = 1191
Score = 36.2 bits (82), Expect = 0.034
Identities = 13/25 (52%), Positives = 19/25 (76%)
Frame = -1
Query: 162 RAGYYWPTLEKDAADHVKQCDPCQR 186
++ +YWPTL K A ++V+ CD CQR
Sbjct: 303 QSSFYWPTLWKGAYEYVQTCDNCQR 229
>TC220300 similar to UP|Q9LZA5 (Q9LZA5) Maturase-like protein, partial (28%)
Length = 689
Score = 32.7 bits (73), Expect = 0.38
Identities = 20/67 (29%), Positives = 31/67 (45%), Gaps = 4/67 (5%)
Frame = -2
Query: 12 RARLRQLEEQETQSRPCPRFPRKRRRTGARSHPLTTRKH---AAGHPRPSGWPAIHRG-A 67
+A + QETQS P PR+ R ++P+ K + PRP+ W + +G +
Sbjct: 271 KAEQPAIPSQETQSSPTPRY-ESRYSPYQHANPIQVPKS*QPPSSTPRPTSWTLMWQGVS 95
Query: 68 HPGAGAH 74
HP H
Sbjct: 94 HPSTSPH 74
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.137 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,975,528
Number of Sequences: 63676
Number of extensions: 315582
Number of successful extensions: 1750
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1716
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1740
length of query: 489
length of database: 12,639,632
effective HSP length: 101
effective length of query: 388
effective length of database: 6,208,356
effective search space: 2408842128
effective search space used: 2408842128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0083.15


