

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
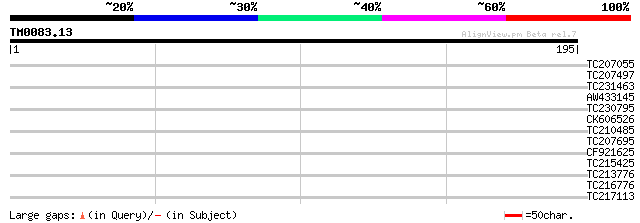
Query= TM0083.13
(195 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207055 homologue to UP|Q9LSD6 (Q9LSD6) ARP2 (Actin-related pro... 28 2.1
TC207497 similar to UP|Q9LN49 (Q9LN49) F18O14.21, partial (80%) 28 2.7
TC231463 28 2.7
AW433145 similar to GP|1061040|emb| sterol-C-methyltransferase {... 28 3.5
TC230795 weakly similar to UP|O23299 (O23299) Carnitine racemase... 27 6.0
CK606526 27 6.0
TC210485 27 6.0
TC207695 weakly similar to UP|Q945N7 (Q945N7) AT5g04420/T32M21_2... 27 6.0
CF921625 27 7.8
TC215425 similar to GB|AAR99502.1|41080591|AY513232 3,5-epimeras... 27 7.8
TC213776 similar to GB|BAC20173.1|23495359|AB073713 BOR1 {Arabid... 27 7.8
TC216776 similar to UP|Q75WV0 (Q75WV0) ALG2-interacting protein ... 27 7.8
TC217113 similar to UP|Q708X5 (Q708X5) Leucine rich repeat prote... 27 7.8
>TC207055 homologue to UP|Q9LSD6 (Q9LSD6) ARP2 (Actin-related protein 2),
partial (98%)
Length = 1724
Score = 28.5 bits (62), Expect = 2.1
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = -2
Query: 39 KQRDPPGNDTYHPKSQASHPPQDKMTSPEA 68
K RDP +DTYHP H QDK+ +A
Sbjct: 385 KTRDPK-HDTYHPSFAQCHCSQDKIYQADA 299
>TC207497 similar to UP|Q9LN49 (Q9LN49) F18O14.21, partial (80%)
Length = 1665
Score = 28.1 bits (61), Expect = 2.7
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = +2
Query: 40 QRDPPGNDTYHPKSQASHPPQDKMTSPEAYLKALRAEALD 79
+R G +TY P++ S PPQ M + A + + ALD
Sbjct: 173 ERSGLGEETYVPEAMHSIPPQPSMAAARAEAEQVMFGALD 292
>TC231463
Length = 730
Score = 28.1 bits (61), Expect = 2.7
Identities = 15/36 (41%), Positives = 18/36 (49%)
Frame = -2
Query: 83 YYSYQTQVTAPGRIRSLVSGLLTEDLDPQLKAGLLT 118
YY +Q + S S LT LDPQLK LL+
Sbjct: 564 YYQHQIMTLIAESVLSFASLSLTTQLDPQLKLVLLS 457
>AW433145 similar to GP|1061040|emb| sterol-C-methyltransferase {Arabidopsis
thaliana}, partial (19%)
Length = 226
Score = 27.7 bits (60), Expect = 3.5
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 4/48 (8%)
Frame = -3
Query: 14 PCHVRKEAVTAKRHTPPKARG----RGLAKQRDPPGNDTYHPKSQASH 57
P H R V +RH PP+ G RG A+Q++ PG + +S A H
Sbjct: 218 PIHPR--IVPPRRHEPPRVDGRRSHRGQARQQN-PGCGLWRGRSHAGH 84
>TC230795 weakly similar to UP|O23299 (O23299) Carnitine racemase like
protein (AT4g14430/dl3255c), partial (47%)
Length = 749
Score = 26.9 bits (58), Expect = 6.0
Identities = 17/62 (27%), Positives = 22/62 (35%)
Frame = +1
Query: 5 ETKGHRPIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMT 64
+ HR +RP H R + +R P GR R PP P PP
Sbjct: 100 QVPSHRRLRPPHRRARQILLQRLRLPLGPGRRRPALRPPP------PPPHVRRPPPRPRR 261
Query: 65 SP 66
+P
Sbjct: 262 AP 267
>CK606526
Length = 513
Score = 26.9 bits (58), Expect = 6.0
Identities = 15/36 (41%), Positives = 18/36 (49%), Gaps = 3/36 (8%)
Frame = +2
Query: 28 TPPKARGRG---LAKQRDPPGNDTYHPKSQASHPPQ 60
TPP GRG K PPGN+ P+S PP+
Sbjct: 155 TPP*KPGRGGPPPQKNSQPPGNNPPPPRSPQRIPPK 262
>TC210485
Length = 746
Score = 26.9 bits (58), Expect = 6.0
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +3
Query: 33 RGRGLAKQRDPPGNDTYHPKSQASHP 58
R + A+ P ++TY PK+Q SHP
Sbjct: 576 RTKDCARAATPQ*SNTYQPKNQISHP 653
>TC207695 weakly similar to UP|Q945N7 (Q945N7) AT5g04420/T32M21_20, partial
(87%)
Length = 2070
Score = 26.9 bits (58), Expect = 6.0
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = -2
Query: 43 PPGNDTYHPKSQASHPPQDKMTSPEAYL 70
PP N T P S+A P + TSPE +
Sbjct: 602 PPNNITLEPNSEADCPTRATGTSPEVLI 519
>CF921625
Length = 523
Score = 26.6 bits (57), Expect = 7.8
Identities = 14/46 (30%), Positives = 21/46 (45%)
Frame = +1
Query: 17 VRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDK 62
++ V K+ P K R +++ PP HPK + PPQ K
Sbjct: 40 IKPPPVGGKKPPPKKRRAPPKGEKKKPPPQKKPHPKKK---PPQKK 168
>TC215425 similar to GB|AAR99502.1|41080591|AY513232
3,5-epimerase/4-reductase {Arabidopsis thaliana;} ,
partial (74%)
Length = 800
Score = 26.6 bits (57), Expect = 7.8
Identities = 20/89 (22%), Positives = 38/89 (42%), Gaps = 2/89 (2%)
Frame = +2
Query: 11 PIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAYL 70
P+R R+ + RH+PP+A R ++R P Q+ P + +P +
Sbjct: 194 PLRLGPPREPGLPRNRHSPPQALPRLQRRRRHGPPQRRLVRVPQSRDHPNQRRRNPHSRR 373
Query: 71 KALRAEALDCNLYY--SYQTQVTAPGRIR 97
+ R L + ++ ++ +P RIR
Sbjct: 374 RLPRPRPHSHQLRHRLHFRVRLRSPPRIR 460
>TC213776 similar to GB|BAC20173.1|23495359|AB073713 BOR1 {Arabidopsis
thaliana;} , partial (4%)
Length = 510
Score = 26.6 bits (57), Expect = 7.8
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = -3
Query: 133 RLEVNMLIWGLEVSATEH 150
+LE+N+L WG E T H
Sbjct: 136 KLELNLLFWGFECGETHH 83
>TC216776 similar to UP|Q75WV0 (Q75WV0) ALG2-interacting protein X, partial
(11%)
Length = 855
Score = 26.6 bits (57), Expect = 7.8
Identities = 21/67 (31%), Positives = 26/67 (38%), Gaps = 3/67 (4%)
Frame = +3
Query: 5 ETKGHRPIRPCH---VRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
+T RP P + V + V H PP G G A+ PP P S A +PP
Sbjct: 180 QTDPARPQAPYYQQPVEQPPVPPYGHHPPPPYG-GPAQHHQPPPPYHIPPSSTAPYPPPQ 356
Query: 62 KMTSPEA 68
P A
Sbjct: 357 VHQQPPA 377
>TC217113 similar to UP|Q708X5 (Q708X5) Leucine rich repeat protein
precursor, partial (96%)
Length = 1553
Score = 26.6 bits (57), Expect = 7.8
Identities = 17/58 (29%), Positives = 22/58 (37%)
Frame = +2
Query: 9 HRPIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSP 66
HR P H R A R P + RG L R PG ++ + H P +P
Sbjct: 383 HRASLPPHPRPHREQALRRDPGR-RGETLTSNRPEPGRQRPKRQNSSVHHPAGLTQTP 553
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,874,339
Number of Sequences: 63676
Number of extensions: 126049
Number of successful extensions: 841
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 815
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 836
length of query: 195
length of database: 12,639,632
effective HSP length: 92
effective length of query: 103
effective length of database: 6,781,440
effective search space: 698488320
effective search space used: 698488320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0083.13


