

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079c.14
(171 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
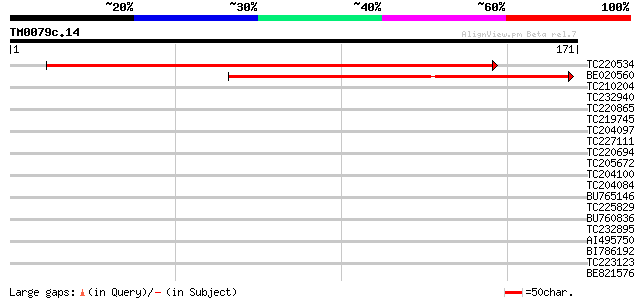
Score E
Sequences producing significant alignments: (bits) Value
TC220534 238 8e-64
BE020560 109 6e-25
TC210204 similar to UP|Q9SY84 (Q9SY84) F14N23.30, partial (21%) 30 0.43
TC232940 homologue to UP|Q7XVJ5 (Q7XVJ5) OJ000126_13.10 protein,... 30 0.57
TC220865 weakly similar to UP|Q6K271 (Q6K271) Flowering time con... 30 0.57
TC219745 30 0.74
TC204097 homologue to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-bindin... 30 0.74
TC227111 homologue to UP|Q8L5G3 (Q8L5G3) Proline rich protein 3 ... 30 0.74
TC220694 29 0.97
TC205672 similar to UP|Q9SCL3 (Q9SCL3) PRE-MRNA SPLICING FACTOR ... 29 1.3
TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, ... 29 1.3
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 29 1.3
BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvo... 28 1.6
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 28 1.6
BU760836 28 1.6
TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfat... 28 1.6
AI495750 28 2.2
BI786192 similar to GP|1403333|emb| mucin {Homo sapiens}, partia... 28 2.2
TC223123 similar to UP|Q9VY72 (Q9VY72) CG32611-PB, partial (3%) 28 2.2
BE821576 27 6.3
>TC220534
Length = 421
Score = 238 bits (608), Expect = 8e-64
Identities = 113/136 (83%), Positives = 123/136 (90%)
Frame = +3
Query: 12 GGGCAVRGGNIYWGRKHATDFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYL 71
GGG AVRGGNIYWGRK TDFRGIVVIFAWVSVPQ LLQ+FV+L SSL WNSLVC+AHYL
Sbjct: 12 GGGFAVRGGNIYWGRKEETDFRGIVVIFAWVSVPQNLLQEFVDLCSSLRWNSLVCFAHYL 191
Query: 72 SAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHCL 131
SAF DES +PLAFCV+DELIEEL+T+SCPVVFAAFSAGSKACL++V QLIDGRCETP L
Sbjct: 192 SAFHDESAMPLAFCVLDELIEELRTRSCPVVFAAFSAGSKACLFRVFQLIDGRCETPLNL 371
Query: 132 HNYHLLRNCVSGHIYD 147
NY LLRNC+SGHIYD
Sbjct: 372 PNYQLLRNCLSGHIYD 419
>BE020560
Length = 335
Score = 109 bits (273), Expect = 6e-25
Identities = 51/104 (49%), Positives = 74/104 (71%)
Frame = +2
Query: 67 YAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCE 126
++ +L+ F E LA +++EL+E LK + CP+VFA+FS G+KAC+ KVLQ+I G E
Sbjct: 8 HSQFLNMFFPEKATILAVDILNELVEVLKIRPCPIVFASFSGGAKACMQKVLQIISGNSE 187
Query: 127 TPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
H + +Y ++R+C+SG+IYDS P+D TSD G RF L PS+ KV
Sbjct: 188 A-HNMDDYQIVRDCISGYIYDSSPVDFTSDXGVRFLLQPSVLKV 316
>TC210204 similar to UP|Q9SY84 (Q9SY84) F14N23.30, partial (21%)
Length = 782
Score = 30.4 bits (67), Expect = 0.43
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = +2
Query: 4 AESEGGGGGGGCAVRGGNIYW 24
+E E GGGGG V GGN+ W
Sbjct: 359 SECENGGGGGEGGVEGGNV*W 421
>TC232940 homologue to UP|Q7XVJ5 (Q7XVJ5) OJ000126_13.10 protein, partial
(13%)
Length = 796
Score = 30.0 bits (66), Expect = 0.57
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = -3
Query: 3 GAESEGGGGGGGCAVRGG 20
GAE GGGGGG AV GG
Sbjct: 143 GAEEAVGGGGGGSAVEGG 90
>TC220865 weakly similar to UP|Q6K271 (Q6K271) Flowering time control protein
isoform rFCA-1, partial (6%)
Length = 709
Score = 30.0 bits (66), Expect = 0.57
Identities = 14/33 (42%), Positives = 22/33 (66%)
Frame = +2
Query: 113 CLYKVLQLIDGRCETPHCLHNYHLLRNCVSGHI 145
C+++ LQ + G+ + P+CL +Y LLR C HI
Sbjct: 38 CIHRWLQCLWGQFKLPNCLLSYLLLR-CKENHI 133
>TC219745
Length = 769
Score = 29.6 bits (65), Expect = 0.74
Identities = 14/41 (34%), Positives = 21/41 (51%), Gaps = 3/41 (7%)
Frame = -3
Query: 127 TPHCLHNYHLLRNCVSGHIYDSGP---LDVTSDFGFRFALH 164
TP C H+ L +C H +D+G L ++ GF F +H
Sbjct: 257 TPTCSHDQW*LLHCSKSHFFDNGSQSHLGLSQQKGFGFGVH 135
>TC204097 homologue to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein,
partial (82%)
Length = 764
Score = 29.6 bits (65), Expect = 0.74
Identities = 15/27 (55%), Positives = 15/27 (55%)
Frame = +1
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDFRG 34
GGGGGGG RGG Y GR RG
Sbjct: 487 GGGGGGGGYNRGGGGYGGRSGGGRDRG 567
>TC227111 homologue to UP|Q8L5G3 (Q8L5G3) Proline rich protein 3 (Fragment),
partial (60%)
Length = 683
Score = 29.6 bits (65), Expect = 0.74
Identities = 15/52 (28%), Positives = 21/52 (39%)
Frame = -2
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATDFRGIVVIFAWVSVPQTLLQDFVN 54
G + GGGGGG A G + +W + W+ VP L+ N
Sbjct: 142 GPPAAAGGGGGGAAAAGAHCWW--------------YGWMPVPAIALRSNPN 29
>TC220694
Length = 548
Score = 29.3 bits (64), Expect = 0.97
Identities = 16/46 (34%), Positives = 22/46 (47%), Gaps = 2/46 (4%)
Frame = -1
Query: 2 RGAESEGGGGGGGCAVRGGNIYWGRKHATDFRGIVV--IFAWVSVP 45
+G G GGG + GG + WG D G+VV I W+ +P
Sbjct: 296 KGHRFSNGFDGGGGSCCGGGVEWG----ADGGGVVVVGIVIWIEIP 171
>TC205672 similar to UP|Q9SCL3 (Q9SCL3) PRE-MRNA SPLICING FACTOR SF2-like
protein, partial (73%)
Length = 1155
Score = 28.9 bits (63), Expect = 1.3
Identities = 16/35 (45%), Positives = 21/35 (59%)
Frame = +2
Query: 2 RGAESEGGGGGGGCAVRGGNIYWGRKHATDFRGIV 36
RG GGGGGGG + GG + +H ++FR IV
Sbjct: 401 RGYGGGGGGGGGGGSGAGGGRFGISRH-SEFRVIV 502
>TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, complete
Length = 1145
Score = 28.9 bits (63), Expect = 1.3
Identities = 13/19 (68%), Positives = 13/19 (68%)
Frame = -3
Query: 8 GGGGGGGCAVRGGNIYWGR 26
GGGGGGG RGG Y GR
Sbjct: 384 GGGGGGGGYNRGGGGYGGR 328
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 28.9 bits (63), Expect = 1.3
Identities = 13/19 (68%), Positives = 13/19 (68%)
Frame = +2
Query: 8 GGGGGGGCAVRGGNIYWGR 26
GGGGGGG RGG Y GR
Sbjct: 1493 GGGGGGGGYNRGGGGYGGR 1549
>BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (25%)
Length = 407
Score = 28.5 bits (62), Expect = 1.6
Identities = 13/26 (50%), Positives = 14/26 (53%)
Frame = +3
Query: 2 RGAESEGGGGGGGCAVRGGNIYWGRK 27
RG GGGGGGG GG GR+
Sbjct: 156 RGGGGGGGGGGGGGGGGGGGFLGGRE 233
Score = 27.7 bits (60), Expect = 2.8
Identities = 13/25 (52%), Positives = 14/25 (56%)
Frame = +2
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRK 27
G E GGGGGGG GG GR+
Sbjct: 314 GGEGGGGGGGGGGGGGGGGGRGGRR 388
Score = 27.7 bits (60), Expect = 2.8
Identities = 12/19 (63%), Positives = 12/19 (63%)
Frame = +1
Query: 2 RGAESEGGGGGGGCAVRGG 20
RG E GGGGGGG GG
Sbjct: 91 RGGEGGGGGGGGGGGGGGG 147
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 28.5 bits (62), Expect = 1.6
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = -3
Query: 3 GAESEGGGGGGGCAVRGG 20
G + +GGGGGGGC GG
Sbjct: 579 GDQVDGGGGGGGCLGVGG 526
>BU760836
Length = 444
Score = 28.5 bits (62), Expect = 1.6
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -1
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDFRGIVVIF 39
GGGGGGG + + + HA R I+V+F
Sbjct: 294 GGGGGGGPGIHRLGLVFPHPHAD*VREILVVF 199
>TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (14%)
Length = 907
Score = 28.5 bits (62), Expect = 1.6
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = -1
Query: 3 GAESEGGGGGGGCAVRGG 20
G E EG GGGGG RGG
Sbjct: 313 GVEEEGDGGGGGVFRRGG 260
>AI495750
Length = 309
Score = 28.1 bits (61), Expect = 2.2
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = +1
Query: 77 ESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDG 123
E+ V + +EL+E L KSC + + G + K+LQ IDG
Sbjct: 64 ETRVEIKEPPTEELLEPLMCKSCVLKVSIHCQGCTRKVKKILQSIDG 204
>BI786192 similar to GP|1403333|emb| mucin {Homo sapiens}, partial (4%)
Length = 427
Score = 28.1 bits (61), Expect = 2.2
Identities = 11/15 (73%), Positives = 11/15 (73%)
Frame = -2
Query: 6 SEGGGGGGGCAVRGG 20
S GGGGGG C RGG
Sbjct: 366 SGGGGGGGSCRQRGG 322
>TC223123 similar to UP|Q9VY72 (Q9VY72) CG32611-PB, partial (3%)
Length = 680
Score = 28.1 bits (61), Expect = 2.2
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = -1
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDF 32
GGGGGGGCA G + GR + F
Sbjct: 320 GGGGGGGCA---GTVTRGRLRSVSF 255
>BE821576
Length = 677
Score = 26.6 bits (57), Expect = 6.3
Identities = 25/96 (26%), Positives = 39/96 (40%), Gaps = 6/96 (6%)
Frame = +2
Query: 46 QTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPL------AFCVVDELIEELKTKSC 99
+ LL F ++ SS+G +L HY+ ++ E L FC++ +E +K
Sbjct: 368 KVLLGRFSDVSSSMG--ALHRLFHYIFSYALEIQNCLNGCSG*IFCIIFNTVEWIKX-XL 538
Query: 100 PVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYH 135
PVV K K+ Q R + C H H
Sbjct: 539 PVVICKLHVTIKIIKIKISQXXXXRHTSIFCTHRXH 646
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.141 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,439,720
Number of Sequences: 63676
Number of extensions: 149931
Number of successful extensions: 3514
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 2230
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3022
length of query: 171
length of database: 12,639,632
effective HSP length: 90
effective length of query: 81
effective length of database: 6,908,792
effective search space: 559612152
effective search space used: 559612152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0079c.14


