

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
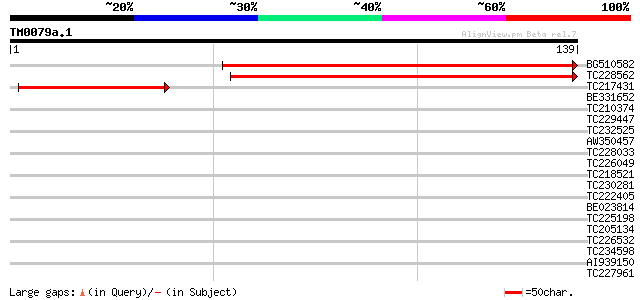
Score E
Sequences producing significant alignments: (bits) Value
BG510582 similar to GP|15010772|gb| At1g49590/F14J22_9 {Arabidop... 134 1e-32
TC228562 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (18%) 134 1e-32
TC217431 similar to GB|AAN18131.1|23308323|BT000562 At5g09390/T5... 47 4e-06
BE331652 similar to GP|15528704|dbj P0583G08.4 {Oryza sativa (ja... 38 0.001
TC210374 31 0.22
TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, parti... 29 0.64
TC232525 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type,... 29 0.64
AW350457 29 0.83
TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (... 29 0.83
TC226049 similar to UP|O24078 (O24078) Protein phosphatase 2C, p... 28 1.1
TC218521 similar to UP|Q8W596 (Q8W596) Fe-superoxide dismutase ... 28 1.1
TC230281 similar to UP|Q9FNY4 (Q9FNY4) DNA polymerase lambda, pa... 28 1.1
TC222405 similar to UP|Q6NQ64 (Q6NQ64) At4g27340, partial (8%) 28 1.4
BE023814 28 1.4
TC225198 similar to UP|Q6Z6A4 (Q6Z6A4) Oxidation protection prot... 28 1.4
TC205134 similar to UP|C862_ARATH (O23066) Cytochrome P450 86A2 ... 28 1.9
TC226532 similar to GB|AAM51367.1|21436395|AY117292 IAA-amino ac... 28 1.9
TC234598 similar to UP|Q9LMI5 (Q9LMI5) T2D23.4 protein, partial ... 28 1.9
AI939150 27 2.4
TC227961 similar to UP|Q9SAF0 (Q9SAF0) F3F19.19 protein (At1g131... 27 3.2
>BG510582 similar to GP|15010772|gb| At1g49590/F14J22_9 {Arabidopsis
thaliana}, partial (39%)
Length = 306
Score = 134 bits (338), Expect = 1e-32
Identities = 68/87 (78%), Positives = 76/87 (87%)
Frame = +3
Query: 53 ASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEK 112
+ S S NKGNK +NG PG VVTTSLNP RNVKAAPSSL++GKRKRPGEK KV+S+EEK
Sbjct: 42 SQSKSDENKGNKFNNGSSPGPVVTTSLNPKRNVKAAPSSLAVGKRKRPGEKSKVISDEEK 221
Query: 113 SALKAREAARKRVQEREKSLLGLYSKP 139
+ALKAREAARKRVQEREK LLGLY+KP
Sbjct: 222 AALKAREAARKRVQEREKPLLGLYNKP 302
>TC228562 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (18%)
Length = 696
Score = 134 bits (337), Expect = 1e-32
Identities = 68/85 (80%), Positives = 75/85 (88%)
Frame = +2
Query: 55 STSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSA 114
S S NKGNK +NG PG VVTTSLNP RNVKAAPSSL++GKRKRPGEK KV+S+EEK+A
Sbjct: 14 SKSDENKGNKFNNGSSPGPVVTTSLNPKRNVKAAPSSLAVGKRKRPGEKSKVISDEEKAA 193
Query: 115 LKAREAARKRVQEREKSLLGLYSKP 139
LKAREAARKRVQEREK LLGLY+KP
Sbjct: 194 LKAREAARKRVQEREKPLLGLYNKP 268
>TC217431 similar to GB|AAN18131.1|23308323|BT000562 At5g09390/T5E8_190
{Arabidopsis thaliana;} , partial (64%)
Length = 1524
Score = 46.6 bits (109), Expect = 4e-06
Identities = 18/37 (48%), Positives = 24/37 (64%)
Frame = +2
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEE 39
++ SSGYYY G+YYDP + YYS A G+W + E
Sbjct: 1127 YDESSGYYYSSNLGYYYDPNTRLYYSAASGQWYSFNE 1237
>BE331652 similar to GP|15528704|dbj P0583G08.4 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 428
Score = 38.1 bits (87), Expect = 0.001
Identities = 14/34 (41%), Positives = 24/34 (70%)
Frame = +2
Query: 6 SSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEE 39
+S Y+H ++GFY+DP +G+YYS+ G + E+
Sbjct: 17 NSQLYFHASSGFYHDPNAGWYYSNKDGAYYKFED 118
Score = 33.9 bits (76), Expect = 0.026
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = +2
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWV 35
F +SSG+Y+ G+YY K G YY G +V
Sbjct: 32 FHASSGFYHDPNAGWYYSNKDGAYYKFEDGNYV 130
>TC210374
Length = 859
Score = 30.8 bits (68), Expect = 0.22
Identities = 18/66 (27%), Positives = 34/66 (51%), Gaps = 1/66 (1%)
Frame = +3
Query: 44 PHFASTTKKASSTSQSNKGNKPHNGPLPGRV-VTTSLNPTRNVKAAPSSLSLGKRKRPGE 102
P +S TK + ST+ + H PL +V + +NPT+ ++ + S + ++ + E
Sbjct: 204 PAVSSLTKPSISTNLPSDPKNKHATPLADKVPLKQDVNPTKELQVSNPSTTPKEKLQENE 383
Query: 103 KPKVVS 108
PKV +
Sbjct: 384 PPKVTT 401
>TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, partial (4%)
Length = 582
Score = 29.3 bits (64), Expect = 0.64
Identities = 18/64 (28%), Positives = 28/64 (43%)
Frame = +1
Query: 36 TQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLG 95
TQ S H T + +ST+ + N+P P P ++N +N K ++
Sbjct: 127 TQNPPNPSSHSNRTMRLLNSTNSNPSPNQP*KTPTPWPSTNQNMNKKKNRKTPKANKR*R 306
Query: 96 KRKR 99
KRKR
Sbjct: 307 KRKR 318
>TC232525 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type, partial
(73%)
Length = 1242
Score = 29.3 bits (64), Expect = 0.64
Identities = 28/102 (27%), Positives = 42/102 (40%), Gaps = 6/102 (5%)
Frame = +1
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG 101
A+ A+ K SST +N +G G V P R V AA K+P
Sbjct: 460 ANAQAATEKNKKSSTEVNNGAVVATDGVGVGNFV-----PARKVGAA---------KKPK 597
Query: 102 EKPKVV------SEEEKSALKAREAARKRVQEREKSLLGLYS 137
E+P+V+ EEK A K ++ K ++ K+ + S
Sbjct: 598 EEPEVIVISSDDESEEKQAAKGKKEREKSARKNAKAFSSVLS 723
>AW350457
Length = 458
Score = 28.9 bits (63), Expect = 0.83
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = -3
Query: 40 AYASPHFASTTKKASSTSQSNKGNKPHN 67
A+ASP A TTK +S S+K N HN
Sbjct: 282 AFASPMKAFTTKGSSKRDASDKNNDTHN 199
>TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (25%)
Length = 2651
Score = 28.9 bits (63), Expect = 0.83
Identities = 31/122 (25%), Positives = 46/122 (37%), Gaps = 10/122 (8%)
Frame = +2
Query: 13 KTNGFYYDPKSG----------FYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
KT G P+SG FYY + + +WV +E A A+ +T+ G
Sbjct: 1127 KTVGLVLKPRSGRQAKLGDKNKFYYDEKLKRWV-EEGAEVPAEEAAALTPPPTTAAFQNG 1303
Query: 63 NKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAAR 122
+ +N L + T S P SSL L PG P + + + + R R
Sbjct: 1304 STEYN--LRSALKTESSPPIEGSSIRTSSLELS----PG-MPLIPPSANQFSARGRLGVR 1462
Query: 123 KR 124
R
Sbjct: 1463 SR 1468
>TC226049 similar to UP|O24078 (O24078) Protein phosphatase 2C, partial (34%)
Length = 604
Score = 28.5 bits (62), Expect = 1.1
Identities = 24/101 (23%), Positives = 43/101 (41%), Gaps = 2/101 (1%)
Frame = +2
Query: 40 AYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVT--TSLNPTRNVKAAPSSLSLGKR 97
++ P +STT SS S + + + P R+ T + + + + + ++ KR
Sbjct: 233 SHLKPSNSSTTSSCSSPSAAASSSSSPSSPFRLRLPKPPTVFSASSSASSTSPNGTVLKR 412
Query: 98 KRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSK 138
KRP VS + AA + + E E+ G+Y K
Sbjct: 413 KRPARLDIPVSSLTFAVPPTPSAAARDLVEAEEDGFGVYCK 535
>TC218521 similar to UP|Q8W596 (Q8W596) Fe-superoxide dismutase , partial
(83%)
Length = 1248
Score = 28.5 bits (62), Expect = 1.1
Identities = 19/52 (36%), Positives = 22/52 (41%)
Frame = -1
Query: 53 ASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKP 104
A STS H P P V LN +RN P SLS+ R+ P P
Sbjct: 657 AFSTSSLLSLYASHAHPEPNCVEAAVLNSSRNFSNEPKSLSIRFRRSPDGLP 502
>TC230281 similar to UP|Q9FNY4 (Q9FNY4) DNA polymerase lambda, partial (21%)
Length = 791
Score = 28.5 bits (62), Expect = 1.1
Identities = 17/45 (37%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Frame = -1
Query: 96 KRKRPG--EKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSK 138
+ RPG +KPK VS E A RE K+++ +K L Y+K
Sbjct: 308 RNSRPG*VQKPK*VSTPESVACLDREKLPKKIEILQKHLADRYTK 174
>TC222405 similar to UP|Q6NQ64 (Q6NQ64) At4g27340, partial (8%)
Length = 505
Score = 28.1 bits (61), Expect = 1.4
Identities = 29/90 (32%), Positives = 39/90 (43%), Gaps = 6/90 (6%)
Frame = -1
Query: 48 STTKKASSTSQSNKGNKPHNGPLPG------RVVTTSLNPTRNVKAAPSSLSLGKRKRPG 101
S T++ SSTS S+K N PLPG RV + S TR + L R+R
Sbjct: 445 SETRRLSSTSFSSKS----NSPLPG*GKRGARVGSMSSPGTRAILRRRGQLRRWPRRR-- 284
Query: 102 EKPKVVSEEEKSALKAREAARKRVQEREKS 131
+S E S R AAR + + + S
Sbjct: 283 -----LSRAEHSEEGTRSAARWKTRVKASS 209
>BE023814
Length = 401
Score = 28.1 bits (61), Expect = 1.4
Identities = 18/75 (24%), Positives = 32/75 (42%)
Frame = +1
Query: 56 TSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSAL 115
T + +KP G P SL +K + S +RKR E+ K E++
Sbjct: 1 TREEPGSSKPSLGDTPAEEENVSLPSQPAMKTSDDSEDESRRKRRREERKEEKREKREKH 180
Query: 116 KAREAARKRVQEREK 130
+RE ++ +E+ +
Sbjct: 181 HSREDRKQEKREKHE 225
>TC225198 similar to UP|Q6Z6A4 (Q6Z6A4) Oxidation protection protein-like,
partial (5%)
Length = 630
Score = 28.1 bits (61), Expect = 1.4
Identities = 14/31 (45%), Positives = 17/31 (54%)
Frame = +1
Query: 34 WVTQEEAYASPHFASTTKKASSTSQSNKGNK 64
WVTQE +P +TT K S SQ + NK
Sbjct: 28 WVTQESESCNPPTKNTTTKLLSCSQMCRRNK 120
>TC205134 similar to UP|C862_ARATH (O23066) Cytochrome P450 86A2 , partial
(87%)
Length = 2056
Score = 27.7 bits (60), Expect = 1.9
Identities = 17/75 (22%), Positives = 34/75 (44%)
Frame = +3
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKV 106
A TT ++++++ KP + P P V ++ +P+R A P + S PK
Sbjct: 351 ACTTGSPTTSARAAARTKPASAPFPSSPVNSASSPSR---ATPKTSSTSSSSASTTTPKA 521
Query: 107 VSEEEKSALKAREAA 121
+S + + +A+
Sbjct: 522 PRGSPRSTICSAKAS 566
>TC226532 similar to GB|AAM51367.1|21436395|AY117292 IAA-amino acid hydrolase
{Arabidopsis thaliana;} , partial (15%)
Length = 486
Score = 27.7 bits (60), Expect = 1.9
Identities = 16/64 (25%), Positives = 29/64 (45%)
Frame = +1
Query: 43 SPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGE 102
SP + +T++S P PLP R++ S +PT + + G+R+R
Sbjct: 229 SPRAGPRSAPPPNTTRSTPHTPP---PLPQRILVASCSPTPSSGIGGPGKARGRRRRKES 399
Query: 103 KPKV 106
P++
Sbjct: 400 APRM 411
>TC234598 similar to UP|Q9LMI5 (Q9LMI5) T2D23.4 protein, partial (17%)
Length = 640
Score = 27.7 bits (60), Expect = 1.9
Identities = 16/47 (34%), Positives = 21/47 (44%)
Frame = +2
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLS 93
ASTT S + + HN PLP + P+R+V P S S
Sbjct: 191 ASTTASTSYATTRSSKTTHHNNPLPKPFSPKNSVPSRHVSPKPISRS 331
>AI939150
Length = 223
Score = 27.3 bits (59), Expect = 2.4
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = +3
Query: 96 KRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGL 135
+R+R E+ + E E+ + RE R+R +ERE+ LGL
Sbjct: 24 ERERERERER---ERERERERERERERERERERERESLGL 134
>TC227961 similar to UP|Q9SAF0 (Q9SAF0) F3F19.19 protein
(At1g13170/F3F19_19), partial (36%)
Length = 1122
Score = 26.9 bits (58), Expect = 3.2
Identities = 27/124 (21%), Positives = 49/124 (38%), Gaps = 7/124 (5%)
Frame = +1
Query: 12 HKTNGFYYDPKSGFYYSDAIGKW------VTQEEAYASPHFASTTKKASSTSQSNKGNKP 65
H+ +GF D ++G + IGKW V + + T+ A + N K
Sbjct: 406 HQVHGFVQDNRTGEKVAMLIGKWDEAMYYVLGDPTTKPKGYDPMTEAALLWERDNHLPKT 585
Query: 66 HNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKV-VSEEEKSALKAREAARKR 124
P + + P + K P+ L +R E + ++ EK L+ + ++
Sbjct: 586 RYNLSPFAISLNEILPGLSEKLPPTDSRLRPDQRHLENGEYELANAEKLRLEQLQRQARK 765
Query: 125 VQER 128
+QER
Sbjct: 766 MQER 777
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,012,511
Number of Sequences: 63676
Number of extensions: 76809
Number of successful extensions: 463
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 455
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 460
length of query: 139
length of database: 12,639,632
effective HSP length: 87
effective length of query: 52
effective length of database: 7,099,820
effective search space: 369190640
effective search space used: 369190640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0079a.1


