

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0076c.7
(656 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
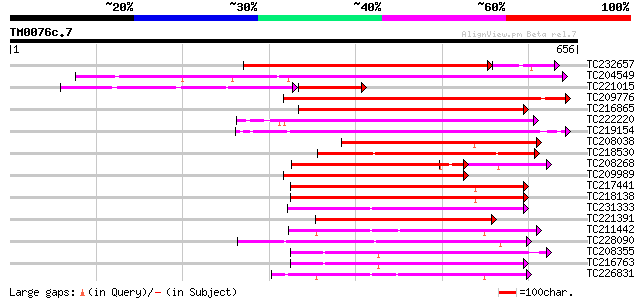
Score E
Sequences producing significant alignments: (bits) Value
TC232657 weakly similar to UP|O22580 (O22580) Receptor-like seri... 454 e-138
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 323 1e-88
TC221015 similar to UP|Q9FNE1 (Q9FNE1) Receptor-like serine/thre... 172 6e-73
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 260 1e-69
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 257 1e-68
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 246 2e-65
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 245 5e-65
TC208038 weakly similar to UP|Q9FNE1 (Q9FNE1) Receptor-like seri... 224 7e-59
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 224 7e-59
TC208268 weakly similar to UP|Q9LMB9 (Q9LMB9) F14D16.24, partial... 221 8e-58
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 214 1e-55
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 213 2e-55
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 212 4e-55
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 208 7e-54
TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-l... 204 8e-53
TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicite... 204 1e-52
TC228090 similar to UP|Q6QLL5 (Q6QLL5) WAK-like kinase, partial ... 201 1e-51
TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete 200 2e-51
TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete 200 2e-51
TC226831 similar to PIR|T52285|T52285 serine/threonine-specific ... 199 3e-51
>TC232657 weakly similar to UP|O22580 (O22580) Receptor-like serine/threonine
kinase, partial (35%)
Length = 1208
Score = 454 bits (1169), Expect(2) = e-138
Identities = 221/288 (76%), Positives = 254/288 (87%)
Frame = +3
Query: 271 IVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATN 330
I+A S AA+A LL+V T++FF RKNV+ +RRERRQFGA L + N SKLNMPYEVLEKATN
Sbjct: 27 ILAASSAALALLLVVVTVVFFTRKNVVTRRRERRQFGALLATVNKSKLNMPYEVLEKATN 206
Query: 331 YFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVK 390
YF+++NKLG+GGSGSVYKG +PDG TVAIKRLS+NTTQWA+HFF EVNLI + HKNLVK
Sbjct: 207 YFNEANKLGQGGSGSVYKGVMPDGNTVAIKRLSYNTTQWAEHFFTEVNLISGIHHKNLVK 386
Query: 391 LLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQ 450
LLGCSITGPESLLVYEYVPN SLHDH SVRR Q LTWE+R KIILG AEG+AYLHEES
Sbjct: 387 LLGCSITGPESLLVYEYVPNQSLHDHFSVRRTSQPLTWEMRQKIILGIAEGMAYLHEESH 566
Query: 451 LKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLGKL 510
++IIHRDIKLSNILL+++FTPKIADFGLARLFPED+S ISTAI GTLGYMAPEYIV GKL
Sbjct: 567 VRIIHRDIKLSNILLEEDFTPKIADFGLARLFPEDKSHISTAIAGTLGYMAPEYIVRGKL 746
Query: 511 TEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDI 558
TEKAD+YSFGVLVIE++SGK +S++ NS S+L VWSLYGSNRL ++
Sbjct: 747 TEKADVYSFGVLVIEIVSGKKISSYIMNSSSLLQTVWSLYGSNRLYEV 890
Score = 58.2 bits (139), Expect(2) = e-138
Identities = 37/84 (44%), Positives = 46/84 (54%), Gaps = 6/84 (7%)
Frame = +1
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINN----IHKIADPTQPPF 614
VDP LEG +PAEEAC+LL+IGLLC + + ++ V N K P QPPF
Sbjct: 892 VDPTLEGAFPAEEACQLLQIGLLCCTSICRVE---TINVSSS*NG**QSTKFLSPAQPPF 1062
Query: 615 LNFGSGEFSRSSLP--EESLQPGS 636
+ S EF +S LP SL P S
Sbjct: 1063IXSSSSEFXQSGLPGYNFSLDPHS 1134
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 323 bits (828), Expect = 1e-88
Identities = 213/599 (35%), Positives = 317/599 (52%), Gaps = 30/599 (5%)
Frame = +3
Query: 77 GSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTRVL-RCSPIQRGVEGGRFFFDGCYLR 135
G T+ + VY C D+ + C C R+ +CS ++ V ++D C +R
Sbjct: 6 GLGTSPSDRVYGLFMCRGDVPSALCQQCVVNATGRLRSQCSLAKQAV----IWYDECTVR 173
Query: 136 YDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVGVLSKR 195
Y +FF+ + + L + S S + + R A + G +++
Sbjct: 174 YSNRSFFSTVDTRPRVGLLNTANISNQDSFMRLLFQTINRTADEAANFSVGLKKYAVNQA 353
Query: 196 NVS----VYGLAQCWNFVNESVCQNCLVEAVTRIDSCAL-KEEGRVLNAGCYLRFSTDKF 250
N+S +Y LAQC +++ C++CL + + C K+ GRVL C +R+ F
Sbjct: 354 NISGFQSLYCLAQCTPDLSQENCRSCLSGVIGDLPWCCQGKQGGRVLYPSCNVRYELYPF 533
Query: 251 YDNSSN-----------IAPPGTKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQ 299
Y +++ + PP + S I A + A+ + VA +IF V L +
Sbjct: 534 YRVTASPPSSSPSPPTLLPPPTSPISPGSSGISAGTIVAIVVPITVAVLIFIVGICFLSR 713
Query: 300 RRERRQFGAFLFSENNSKLNMP--------YEVLEKATNYFHDSNKLGEGGSGSVYKGAL 351
R ++Q G+ E + ++P + +E ATN F NKLGEGG G VYKG L
Sbjct: 714 RARKKQQGSV--KEGKTAYDIPTVDSLQFDFSTIEAATNKFSADNKLGEGGFGEVYKGTL 887
Query: 352 PDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNL 411
G VA+KRLS ++ Q + F NEV ++ LQH+NLV+LLG + G E +LVYEYVPN
Sbjct: 888 SSGQVVAVKRLSKSSGQGGEEFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNK 1067
Query: 412 SLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTP 471
SL L ++L W R+KII G A G+ YLHE+S+L+IIHRD+K SNILLD + P
Sbjct: 1068SLDYILFDPEKQRELDWGRRYKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMNP 1247
Query: 472 KIADFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK 530
KI+DFG+AR+F DQ+Q +T+ I GT GYMAPEY + G+ + K+D+YSFGVL++E+LSGK
Sbjct: 1248KISDFGMARIFGVDQTQGNTSRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGK 1427
Query: 531 SRTSFVQNSCS--ILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAE 588
+SF Q + +L W L+ +++DPIL +Y E + + IGLLC Q
Sbjct: 1428KNSSFYQTDGAEDLLSYAWQLWKDGTPLELMDPILRESYNQNEVIRSIHIGLLCVQEDPA 1607
Query: 589 LRPPMSVVVKMI-NNIHKIADPTQPP-FLNFGSGEFSRSSLPEESLQPGSNTQSSGDSM 645
RP M+ +V M+ +N + PTQP F++ G+ LP + P S S M
Sbjct: 1608DRPTMATIVLMLDSNTVTLPTPTQPAFFVHSGTDPNMPKELPFDQSIPMSVNDMSISEM 1784
Score = 39.7 bits (91), Expect = 0.004
Identities = 21/64 (32%), Positives = 35/64 (53%), Gaps = 1/64 (1%)
Frame = +3
Query: 189 VGVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRIDS-CALKEEGRVLNAGCYLRFST 247
VG+ + + VYGL C V ++CQ C+V A R+ S C+L ++ + C +R+S
Sbjct: 3 VGLGTSPSDRVYGLFMCRGDVPSALCQQCVVNATGRLRSQCSLAKQAVIWYDECTVRYSN 182
Query: 248 DKFY 251
F+
Sbjct: 183 RSFF 194
>TC221015 similar to UP|Q9FNE1 (Q9FNE1) Receptor-like serine/threonine
kinase, partial (33%)
Length = 1110
Score = 172 bits (435), Expect(2) = 6e-73
Identities = 103/280 (36%), Positives = 150/280 (52%), Gaps = 5/280 (1%)
Frame = +1
Query: 59 LESLTSLVTTQRHGT-VVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTRVLRCSP 117
+ESL+ LVT+ GT VK S + + +Y F +C +DLS +DC +C+A +TR+ RC P
Sbjct: 34 MESLSQLVTSNNWGTHSVKISGSGSSIPIYGFAQCFRDLSHTDCLLCYAASRTRLPRCLP 213
Query: 118 IQRGVEGGRFFFDGCYLRYDAYNFFNQSLS-SQDLTLCGNIDFS--GNRSVHKANVVELV 174
R + DGC+LRYD Y+F+++ S+D C + R + V +V
Sbjct: 214 SV----SARIYLDGCFLRYDNYSFYSEGTDPSRDAVNCTGVAAGDEAERVELQERVGRVV 381
Query: 175 RNL-SVEAPKNDGFFVGVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRIDSCALKEE 233
N+ ++ +GF VG + VY LAQCWN + C+ CL +A + C K+E
Sbjct: 382 DNVVNIAERDGNGFGVGEVE----GVYALAQCWNTLGSGGCRECLRKAGREVKGCLPKKE 549
Query: 234 GRVLNAGCYLRFSTDKFYDNSSNIAPPGTKGSTKVGIIVAESFAAVATLLIVATIIFFVR 293
GR LNAGCYLR+ST KFY N A G + G+IVAE AA A +++ + +
Sbjct: 550 GRALNAGCYLRYSTQKFY-NEDGDAGGGNGFLRRRGVIVAEVLAAAAVIMLALSASYAAF 726
Query: 294 KNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFH 333
+ ++E G S + S LN YE LEKAT+YF+
Sbjct: 727 TKFSKIKKENNNLGQISSSISKSSLNYKYETLEKATDYFN 846
Score = 121 bits (304), Expect(2) = 6e-73
Identities = 56/79 (70%), Positives = 65/79 (81%)
Frame = +2
Query: 335 SNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGC 394
S K+G+GG+GSV+KG LP+G VA+KRL FN QW D FFNEVNLI ++HKNLVKLLGC
Sbjct: 851 SRKVGQGGAGSVFKGILPNGKVVAVKRLIFNNRQWVDEFFNEVNLISGIEHKNLVKLLGC 1030
Query: 395 SITGPESLLVYEYVPNLSL 413
SI GPESLLVYEY+P SL
Sbjct: 1031SIEGPESLLVYEYLPKKSL 1087
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 260 bits (665), Expect = 1e-69
Identities = 143/336 (42%), Positives = 204/336 (60%), Gaps = 4/336 (1%)
Frame = +2
Query: 318 LNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEV 377
L + +E AT F ++NKLGEGG G VYKG LP G VA+KRLS + Q + F NEV
Sbjct: 38 LRFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGGEEFKNEV 217
Query: 378 NLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILG 437
++ LQH+NLV+LLG + G E +LVYE+V N SL L + L W R+KI+ G
Sbjct: 218 EIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKIVEG 397
Query: 438 TAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGT 496
A G+ YLHE+S+LKIIHRD+K SN+LLD + PKI+DFG+AR+F DQ+Q +T I GT
Sbjct: 398 IARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVGT 577
Query: 497 LGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCS--ILHMVWSLYGSNR 554
GYM+PEY + G+ + K+D+YSFGVL++E++SGK +SF + + +L W L+
Sbjct: 578 YGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLWKDEA 757
Query: 555 LCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIH-KIADPTQPP 613
+++D L +Y E + + IGLLC Q RP M+ VV M+++ + P QP
Sbjct: 758 PLELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDSYSVTLQVPNQPA 937
Query: 614 FLNFGSGEFSRSSLPEESLQPGSNTQSSGDSMTESS 649
F + ++P+ S T S+ S+ + S
Sbjct: 938 FY---INSRTEPNMPKGLKIDQSTTNSTSKSVNDMS 1036
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 257 bits (656), Expect = 1e-68
Identities = 136/269 (50%), Positives = 178/269 (65%), Gaps = 3/269 (1%)
Frame = +1
Query: 335 SNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGC 394
+NK+GEGG G VYKG DGT +A+K+LS + Q F NE+ +I LQH +LVKL GC
Sbjct: 4 ANKIGEGGFGPVYKGCFSDGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYGC 183
Query: 395 SITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEVRHKIILGTAEGLAYLHEESQLKI 453
+ G + LLVYEY+ N SL L Q +L W R+KI +G A GLAYLHEES+LKI
Sbjct: 184 CVEGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLKI 363
Query: 454 IHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLGKLTEK 513
+HRDIK +N+LLD + PKI+DFGLA+L ED + IST I GT GYMAPEY + G LT+K
Sbjct: 364 VHRDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHISTRIAGTFGYMAPEYAMHGYLTDK 543
Query: 514 ADIYSFGVLVIELLSGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEE 571
AD+YSFG++ +E+++G+S T Q S S+L L + D+VD L + EE
Sbjct: 544 ADVYSFGIVALEIINGRSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNKEE 723
Query: 572 ACKLLKIGLLCTQASAELRPPMSVVVKMI 600
A ++K+ LLCT +A LRP MS VV M+
Sbjct: 724 ALVMIKVALLCTNVTAALRPTMSSVVSML 810
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 246 bits (628), Expect = 2e-65
Identities = 143/360 (39%), Positives = 214/360 (58%), Gaps = 10/360 (2%)
Frame = +3
Query: 263 KGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFL---FSENNS--- 316
K ++V +IV V +++++ + +F+ K R R+++ L F E ++
Sbjct: 27 KSVSRVVLIVVP---VVVSIILLLCVCYFILK------RSRKKYNTLLRENFGEESATLE 179
Query: 317 KLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNE 376
L +E ATN F ++GEGG G VYKG LPDG +A+K+LS ++ Q A+ F NE
Sbjct: 180 SLQFGLVTIEAATNKFSYEKRIGEGGFGVVYKGVLPDGREIAVKKLSKSSGQGANEFKNE 359
Query: 377 VNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIIL 436
+ LI LQH+NLV LLG + E +L+YE+V N SL L +QL W R+KII
Sbjct: 360 ILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSNKSLDYFLFDSHRSKQLNWSERYKIIE 539
Query: 437 GTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICG 495
G A+G++YLHE S+LK+IHRD+K SN+LLD N PKI+DFG+AR+ DQ Q T I G
Sbjct: 540 GIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMNPKISDFGMARIVAIDQLQGKTNRIVG 719
Query: 496 TLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRT-SFVQNSCSILHMVWSLYGSNR 554
T GYM+PEY + G+ +EK+D++SFGV+V+E++S K T S + +L W +
Sbjct: 720 TYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIISAKRNTRSVFSDHDDLLSYTWEQWMDEA 899
Query: 555 LCDIVDPILEGNY-PAEEACKLLKIGLLCTQASAELRPPMSVVVKMIN-NIHKIADPTQP 612
+I D +E + E K ++IGLLC Q + RP ++ V+ +N +I ++ P +P
Sbjct: 900 PLNIFDQSIEAEFCDHSEVVKCIQIGLLCVQEKPDDRPKITQVISYLNSSITELPLPKKP 1079
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 245 bits (625), Expect = 5e-65
Identities = 150/390 (38%), Positives = 222/390 (56%), Gaps = 3/390 (0%)
Frame = +1
Query: 262 TKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMP 321
+K S+K II+ S AA L +V I F R+N+ + + ++ L + +M
Sbjct: 157 SKKSSK--IIIGTSVAA--PLGVVLAICFIYRRNIADKSKTKKSIDRQLQDVDVPLFDML 324
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIK 381
+ AT+ F +NK+GEGG G VYKG L G +A+KRLS + Q F EV LI
Sbjct: 325 --TITAATDNFLLNNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLSGQGITEFITEVKLIA 498
Query: 382 DLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEG 441
LQH+NLVKLLGC I G E LLVYEYV N SL+ + + + L W R IILG A G
Sbjct: 499 KLQHRNLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIKSKLLDWPRRFNIILGIARG 678
Query: 442 LAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYM 500
L YLH++S+L+IIHRD+K SN+LLD+ PKI+DFG+AR F DQ++ +T + GT GYM
Sbjct: 679 LLYLHQDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTYGYM 858
Query: 501 APEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFV--QNSCSILHMVWSLYGSNRLCDI 558
APEY G + K+D++SFG+L++E++ G S + +++ W+L+ +
Sbjct: 859 APEYAFDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLNLVGYAWALWKEQNALQL 1038
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPFLNFG 618
+D ++ + E + + + LLC Q E RP M+ V++M+ + + +P +P F
Sbjct: 1039IDSGIKDSCVIPEVLRCIHVSLLCVQQYPEDRPTMTSVIQMLGSEMDMVEPKEPGF---- 1206
Query: 619 SGEFSRSSLPEESLQPGSNTQSSGDSMTES 648
F R L E +L+ +S D +T S
Sbjct: 1207---FPRRILKEGNLK----EMTSNDELTIS 1275
>TC208038 weakly similar to UP|Q9FNE1 (Q9FNE1) Receptor-like serine/threonine
kinase, partial (25%)
Length = 797
Score = 224 bits (572), Expect = 7e-59
Identities = 120/237 (50%), Positives = 155/237 (64%), Gaps = 6/237 (2%)
Frame = +1
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAY 444
H+NLV+LLGC GPE +LVYEY+ N SL L L W+ R+ IILGTA GLAY
Sbjct: 4 HRNLVRLLGCCSRGPERILVYEYMANSSLDKFLFAGSKKGSLNWKQRYDIILGTARGLAY 183
Query: 445 LHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEY 504
LHEE + IIHRDIK NILLDD PKIADFGLARL P D+S +ST GTLGY APEY
Sbjct: 184 LHEEFHVSIIHRDIKTGNILLDDYLQPKIADFGLARLLPRDRSHLSTKFAGTLGYTAPEY 363
Query: 505 IVLGKLTEKADIYSFGVLVIELLSGKSRTSFV---QNSCSILHMVWSLYGSNRLCDIVDP 561
+ G+L+EKAD YS+G++V+E+LSG+ T+ + +L W LY ++VD
Sbjct: 364 AMQGQLSEKADTYSYGIVVLEILSGQKSTNVKVDDEGREYLLQRAWKLYERGMQLELVDK 543
Query: 562 ILEGN-YPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIAD--PTQPPFL 615
++ N Y AEEA K+++I LLCTQASA RP MS ++ ++ + + PT P F+
Sbjct: 544 DIDPNEYDAEEAKKIIEIALLCTQASAAARPTMSELIVLLKSKSLVEHLRPTMPVFV 714
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 224 bits (572), Expect = 7e-59
Identities = 120/259 (46%), Positives = 165/259 (63%), Gaps = 2/259 (0%)
Frame = +3
Query: 357 VAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDH 416
+A+K+LS + Q F E+ I +QH+NLVKL GC I G + LLVYEY+ N SL
Sbjct: 6 IAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQA 185
Query: 417 LSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADF 476
L + L W R+ I LG A GL YLHEES+L+I+HRD+K SNILLD PKI+DF
Sbjct: 186 LFGK--CLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISDF 359
Query: 477 GLARLFPEDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK--SRTS 534
GLA+L+ + ++ IST + GT+GY+APEY + G LTEKAD++SFGV+ +EL+SG+ S +S
Sbjct: 360 GLAKLYDDKKTHISTGVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSDSS 539
Query: 535 FVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMS 594
+L W L+ N + D+VD L + EE +++ I LLCTQ S LRP MS
Sbjct: 540 LEGEKVYLLEWAWQLHEKNCIIDLVDDRL-SEFNEEEVKRVVGIALLCTQTSPTLRPSMS 716
Query: 595 VVVKMINNIHKIADPTQPP 613
VV M++ +++ T P
Sbjct: 717 RVVAMLSGDIEVSTVTSKP 773
>TC208268 weakly similar to UP|Q9LMB9 (Q9LMB9) F14D16.24, partial (32%)
Length = 1457
Score = 221 bits (563), Expect = 8e-58
Identities = 115/207 (55%), Positives = 147/207 (70%), Gaps = 3/207 (1%)
Frame = +2
Query: 327 KATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHK 386
KAT FH++NKLG+GG G+VYKG L DG +A+KRL FN A F+NEVN+I ++HK
Sbjct: 2 KATESFHENNKLGQGGFGTVYKGVLADGREIAVKRLFFNNRHRAADFYNEVNIISSVEHK 181
Query: 387 NLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLH 446
NLV+LLGCS +GPESLLVYE++PN SL ++ + ++L WE R++II+GTAEGL YLH
Sbjct: 182 NLVRLLGCSCSGPESLLVYEFLPNRSLDRYIFDKNKGKELNWEKRYEIIIGTAEGLVYLH 361
Query: 447 EESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTL---GYMAPE 503
E S+ +IIHRDIK SNILLD KIADFGLAR F ED+S ISTAI GTL GY
Sbjct: 362 ENSKTRIIHRDIKASNILLDAKLRAKIADFGLARSFQEDKSHISTAIAGTLEVYGY*IHN 541
Query: 504 YIVLGKLTEKADIYSFGVLVIELLSGK 530
+ K K D GVL++E+++ +
Sbjct: 542 SYSVNK--SKEDAKRHGVLLLEIVTAR 616
Score = 80.9 bits (198), Expect = 2e-15
Identities = 47/139 (33%), Positives = 81/139 (57%), Gaps = 10/139 (7%)
Frame = +1
Query: 498 GYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK--SRTSFVQNSCSILHMVWSLYGSNRL 555
G +APEY+ G+LTEKAD+YSFGVL++E+++ + +R+ + S S++ + W + +
Sbjct: 661 GTVAPEYLAHGQLTEKADVYSFGVLLLEIVTARQNNRSKASEYSDSLVTVAWKHFQAGTA 840
Query: 556 CDIVDPILE-------GNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHK-IA 607
+ DP L+ +E +++ IGLLCTQ + LRP MS ++M+ + +
Sbjct: 841 EQLFDPNLDLQEDHNSNVNVKDEILRVVHIGLLCTQEVSSLRPSMSKALQMLTKKEEDLV 1020
Query: 608 DPTQPPFLNFGSGEFSRSS 626
P+ PPFL+ + E +S
Sbjct: 1021APSNPPFLDESTMELHDTS 1077
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 214 bits (545), Expect = 1e-55
Identities = 110/214 (51%), Positives = 146/214 (67%), Gaps = 1/214 (0%)
Frame = +3
Query: 318 LNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEV 377
L + ++ ATN F D+NKLG+GG G VYKG L DG +AIKRLS N+ Q F NE+
Sbjct: 93 LQFEFATIKFATNNFSDANKLGQGGFGIVYKGTLSDGQEIAIKRLSINSNQGETEFKNEI 272
Query: 378 NLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILG 437
L LQH+NLV+LLG E LL+YE+VPN SL + L E+R+KII G
Sbjct: 273 LLTGRLQHRNLVRLLGFCFARRERLLIYEFVPNKSLDYFIFDPNKRVNLN*EIRYKIIRG 452
Query: 438 TAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIS-TAICGT 496
A GL YLHE+S+L ++HRD+K SNILLD PKI+DFG+ARLF +Q++ S T I GT
Sbjct: 453 IARGLLYLHEDSRLNVVHRDLKTSNILLDGELNPKISDFGMARLFEINQTEASTTTIVGT 632
Query: 497 LGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK 530
GYMAPEYI G+ + K+D++SFGV+++E++ G+
Sbjct: 633 FGYMAPEYIKHGQFSIKSDVFSFGVMILEIVCGQ 734
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 213 bits (542), Expect = 2e-55
Identities = 120/282 (42%), Positives = 173/282 (60%), Gaps = 6/282 (2%)
Frame = +2
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWAD-HFFNEVNLIKDL 383
L+ AT+ F + + LG GG G VYKG L DG+ VA+KRL TQ F EV +I
Sbjct: 134 LQVATDNFSNKHILGRGGFGKVYKGRLADGSLVAVKRLKEERTQGG*LQFQTEVEMISMA 313
Query: 384 QHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ-LTWEVRHKIILGTAEGL 442
H+NL++L G +T E LLVY Y+ N S+ L R+ Q L W R +I LG+A GL
Sbjct: 314 VHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPERKRIALGSARGL 493
Query: 443 AYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAP 502
AYLH+ KIIHRD+K +NILLD+ F + DFGLA+L + ++TA+ GT+G++AP
Sbjct: 494 AYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRGTIGHIAP 673
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ----NSCSILHMVWSLYGSNRLCDI 558
EY+ GK +EK D++ +GV+++EL++G+ + + +L V L +L +
Sbjct: 674 EYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKDRKLETL 853
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
VD L+G+Y EE +L+++ LLCTQ S RP MS VV+M+
Sbjct: 854 VDADLQGSYNDEEVEQLIQVALLCTQGSPMERPKMSEVVRML 979
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 212 bits (540), Expect = 4e-55
Identities = 117/282 (41%), Positives = 172/282 (60%), Gaps = 6/282 (2%)
Frame = +1
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWAD-HFFNEVNLIKDL 383
L+ AT+ F + N LG GG G VYKG L DG+ VA+KRL T + F EV +I
Sbjct: 136 LQVATDSFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQFQTEVEMISMA 315
Query: 384 QHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ-LTWEVRHKIILGTAEGL 442
H+NL++L G +T E LLVY Y+ N S+ L R Q+ L W R ++ LG+A GL
Sbjct: 316 VHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTRKRVALGSARGL 495
Query: 443 AYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAP 502
+YLH+ KIIHRD+K +NILLD+ F + DFGLA+L + ++TA+ GT+G++AP
Sbjct: 496 SYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRGTIGHIAP 675
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ----NSCSILHMVWSLYGSNRLCDI 558
EY+ GK +EK D++ +G++++EL++G+ + + +L V L +L +
Sbjct: 676 EYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKEKKLEML 855
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
VDP L+ NY E +L+++ LLCTQ S RP MS VV+M+
Sbjct: 856 VDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRML 981
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 208 bits (529), Expect = 7e-54
Identities = 112/282 (39%), Positives = 163/282 (57%), Gaps = 3/282 (1%)
Frame = +3
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIK 381
+ L+ AT F +N L EGG GSV++G LPDG +A+K+ +TQ F +EV ++
Sbjct: 12 FSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQGDKEFCSEVEVLS 191
Query: 382 DLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEG 441
QH+N+V L+G + LLVYEY+ N SL H+ RR L W R KI +G A G
Sbjct: 192 CAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHI-YRRKQNVLEWSARQKIAVGAARG 368
Query: 442 LAYLHEESQLK-IIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYM 500
L YLHEE ++ I+HRD++ +NILL +F + DFGLAR P+ + T + GT GY+
Sbjct: 369 LRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARWQPDGDMGVETRVIGTFGYL 548
Query: 501 APEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWS--LYGSNRLCDI 558
APEY G++TEKAD+YSFG++++EL++G+ + W+ L +
Sbjct: 549 APEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEWARPLLEKQATYKL 728
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
+DP L Y +E ++LK LC LRP MS V++M+
Sbjct: 729 IDPSLRNCYVDQEVYRMLKCSSLCIGRDPHLRPRMSQVLRML 854
>TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-like protein
, partial (25%)
Length = 642
Score = 204 bits (520), Expect = 8e-53
Identities = 106/212 (50%), Positives = 146/212 (68%), Gaps = 2/212 (0%)
Frame = +2
Query: 354 GTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSL 413
G VA+KRLS ++Q + F NE+ LI LQH+NLV+LLGC I G E +LVYEY+PN SL
Sbjct: 2 GEEVAVKRLSRKSSQGLEEFKNEMVLIAKLQHRNLVRLLGCCIQGEEKILVYEYLPNKSL 181
Query: 414 HDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKI 473
L QL W R +II G A GL YLH +S+L+IIHRD+K SNILLD++ PKI
Sbjct: 182 DCFLFDPVKQTQLDWAKRFEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDESMNPKI 361
Query: 474 ADFGLARLFPEDQSQIST-AICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSR 532
+DFGLAR+F +Q++ +T + GT GYM+PEY + G + K+D+YSFGVL++E++SG+
Sbjct: 362 SDFGLARIFGGNQNEANTNRVVGTYGYMSPEYAMEGLFSIKSDVYSFGVLLLEIMSGRKN 541
Query: 533 TSFVQ-NSCSILHMVWSLYGSNRLCDIVDPIL 563
TSF + S++ W L+ R+ ++VDP L
Sbjct: 542 TSFRDTDDSSLIGYAWHLWSEQRVMELVDPSL 637
>TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicited protein
264, partial (73%)
Length = 1047
Score = 204 bits (518), Expect = 1e-52
Identities = 122/304 (40%), Positives = 176/304 (57%), Gaps = 11/304 (3%)
Frame = +2
Query: 323 EVLEKATNYFHDSNKLGEGGSGSVYKGALPD-------GTTVAIKRLSFNTTQWADHFFN 375
E L +ATN F SN LGEGG G VYKG + D TVA+KRL + Q +
Sbjct: 44 EELREATNSFSWSNMLGEGGFGPVYKGFVDDKLRSGLKAQTVAVKRLDLDGLQGHREWLA 223
Query: 376 EVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKII 435
E+ + L+H +LVKL+G LL+YEY+P SL + L RR + W R KI
Sbjct: 224 EIIFLGQLRHPHLVKLIGYCYEDEHRLLMYEYMPRGSLENQL-FRRYSAAMPWSTRMKIA 400
Query: 436 LGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPE-DQSQISTAIC 494
LG A+GLA+LHE + +I+RD K SNILLD +FT K++DFGLA+ PE + + ++T I
Sbjct: 401 LGAAKGLAFLHEADK-PVIYRDFKASNILLDSDFTAKLSDFGLAKDGPEGEDTHVTTRIM 577
Query: 495 GTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWS---LYG 551
GT GY APEYI+ G LT K+D+YS+GV+++ELL+G+ +++ + W+ L
Sbjct: 578 GTQGYAAPEYIMTGHLTTKSDVYSYGVVLLELLTGRRVVDKSRSNEGKSLVEWARPLLRD 757
Query: 552 SNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQ 611
++ +I+D LEG +P + A K+ + C RP MS V+K++ + D
Sbjct: 758 QKKVYNIIDRRLEGQFPMKGAMKVAMLAFKCLSHHPNARPTMSDVIKVLEPLQDYDDVFI 937
Query: 612 PPFL 615
PF+
Sbjct: 938 GPFV 949
>TC228090 similar to UP|Q6QLL5 (Q6QLL5) WAK-like kinase, partial (51%)
Length = 1533
Score = 201 bits (510), Expect = 1e-51
Identities = 120/348 (34%), Positives = 189/348 (53%), Gaps = 8/348 (2%)
Frame = +1
Query: 264 GSTKVGIIVAESFAAVATLLIVATII-FFVRKNVLRQRRERRQFGAFLFSENNSKLNMPY 322
G+T+ +++ V+ ++ + ++ F+ R++ LR ++ +N+ + PY
Sbjct: 13 GTTRFIVLIGGFVVGVSLMVTLGSLCCFYRRRSKLRVTNSTKRRLTEATGKNSVPI-YPY 189
Query: 323 EVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKD 382
+ +EKATN F + +LG G G+VY G L + VAIKR+ T + NE+ L+
Sbjct: 190 KDIEKATNSFSEKQRLGTGAYGTVYAGKLYNNEWVAIKRIKHRDTDSIEQVMNEIKLLSS 369
Query: 383 LQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGL 442
+ H NLV+LLGCSI E +LVYE++PN +L HL R L W +R I TA+ +
Sbjct: 370 VSHTNLVRLLGCSIEYGEQILVYEFMPNGTLSQHLQKERG-SGLPWPIRLTIATETAQAI 546
Query: 443 AYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAP 502
AYLH I HRDIK SNILLD NF K+ADFGL+RL + S IST GT GY+ P
Sbjct: 547 AYLHSAICPPIYHRDIKSSNILLDYNFRSKVADFGLSRLGMTEISHISTTPQGTPGYVDP 726
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVD 560
+Y L++K+D+YS GV+++E+++G+ F + N ++ + G L +I+D
Sbjct: 727 QYHQDFHLSDKSDVYSLGVVLVEIITGQKVVDFSRPHNEVNLASLAADKIGKGLLNEIID 906
Query: 561 PILEGN-----YPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
P LE + K+ ++ C ++RP M+ V + +
Sbjct: 907 PFLEPEVRSDAWTLSSIHKVAELAFRCIAFHRDMRPSMTEVASELEQL 1050
>TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete
Length = 1536
Score = 200 bits (508), Expect = 2e-51
Identities = 117/311 (37%), Positives = 176/311 (55%), Gaps = 9/311 (2%)
Frame = +2
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L+ T+ F +GEG G VY+ L +G V IK+L ++ Q F ++V+++ L+
Sbjct: 383 LKPLTDNFGSKCFIGEGAYGKVYQATLKNGRAVVIKKLD-SSNQPEHEFLSQVSIVSRLK 559
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ------LTWEVRHKIILGT 438
H+N+V+L+ + GP L YEY P SLHD L R+ ++ L+W R KI +G
Sbjct: 560 HENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVGA 739
Query: 439 AEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI-STAICGTL 497
A GL YLHE++++ IIHR IK SNILL D+ KIADF L+ P+ +++ ST + GT
Sbjct: 740 ARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKIADFDLSNQAPDAAARLHSTRVLGTF 919
Query: 498 GYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRT--SFVQNSCSILHMVWSLYGSNRL 555
GY APEY + G+LT K+D+YSFGV+++ELL+G+ + + S++ +++
Sbjct: 920 GYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 1099
Query: 556 CDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPFL 615
VD L+G YP++ K+ + LC Q AE RP MS++VK + P L
Sbjct: 1100KQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKALQ-----------PLL 1246
Query: 616 NFGSGEFSRSS 626
N S SS
Sbjct: 1247NTRSSHSKESS 1279
>TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete
Length = 1634
Score = 200 bits (508), Expect = 2e-51
Identities = 110/285 (38%), Positives = 170/285 (59%), Gaps = 9/285 (3%)
Frame = +1
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L+ T+ F +GEG G VY+ L +G V IK+L ++ Q F ++V+++ L+
Sbjct: 373 LKSLTDNFGSKYFIGEGAYGKVYQATLKNGHAVVIKKLD-SSNQPEQEFLSQVSIVSRLK 549
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ------LTWEVRHKIILGT 438
H+N+V+L+ + GP L YEY P SLHD L R+ ++ L+W R KI +G
Sbjct: 550 HENVVELVNYCVDGPFRALAYEYAPKGSLHDILHGRKGVKGAQPGPVLSWAQRVKIAVGA 729
Query: 439 AEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI-STAICGTL 497
A GL YLHE++++ IIHR IK SNILL D+ K+ADF L+ P+ +++ ST + GT
Sbjct: 730 ARGLEYLHEKAEIHIIHRYIKSSNILLFDDDVAKVADFDLSNQAPDAAARLHSTRVLGTF 909
Query: 498 GYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRT--SFVQNSCSILHMVWSLYGSNRL 555
GY APEY + G+LT K+D+YSFGV+++ELL+G+ + + S++ +++
Sbjct: 910 GYHAPEYAMTGQLTSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKV 1089
Query: 556 CDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
VD L+G YP++ K+ + LC Q AE RP MS++VK +
Sbjct: 1090KQCVDVRLKGEYPSKSVAKMAAVAALCVQYEAEFRPNMSIIVKAL 1224
>TC226831 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (82%)
Length = 1693
Score = 199 bits (506), Expect = 3e-51
Identities = 122/314 (38%), Positives = 174/314 (54%), Gaps = 14/314 (4%)
Frame = +1
Query: 304 RQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPD---------- 353
R G L S N + L+ AT F + LGEGG G VYKG + +
Sbjct: 397 RSEGEILSSPNLKAFT--FNELKNATRNFRPDSLLGEGGFGYVYKGWIDEHTFTASKPGS 570
Query: 354 GTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSL 413
G VA+K+L Q + EV+ + L H+NLVKL+G G LLVYE++ SL
Sbjct: 571 GMVVAVKKLKPEGLQGHKEWLTEVDYLGQLHHQNLVKLIGYCADGENRLLVYEFMSKGSL 750
Query: 414 HDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKI 473
+HL RR Q L+W VR K+ +G A GL++LH ++ ++I+RD K SNILLD F K+
Sbjct: 751 ENHL-FRRGPQPLSWSVRMKVAIGAARGLSFLHN-AKSQVIYRDFKASNILLDAEFNAKL 924
Query: 474 ADFGLARLFPE-DQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSR 532
+DFGLA+ P D++ +ST + GT GY APEY+ G+LT K+D+YSFGV+++ELLSG+
Sbjct: 925 SDFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLSGRRA 1104
Query: 533 TSFVQNSCSILHMVWS---LYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAEL 589
+ + W+ L RL I+D L G YP + A + L C A+
Sbjct: 1105VDRSKAGVEQNLVEWAKPYLGDKRRLFRIMDTKLGGQYPQKGAYMAATLALKCLNREAKG 1284
Query: 590 RPPMSVVVKMINNI 603
RPP++ V++ + I
Sbjct: 1285RPPITEVLQTLEQI 1326
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,472,828
Number of Sequences: 63676
Number of extensions: 483587
Number of successful extensions: 5101
Number of sequences better than 10.0: 1022
Number of HSP's better than 10.0 without gapping: 4205
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4296
length of query: 656
length of database: 12,639,632
effective HSP length: 103
effective length of query: 553
effective length of database: 6,081,004
effective search space: 3362795212
effective search space used: 3362795212
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0076c.7


