

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.9
(351 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
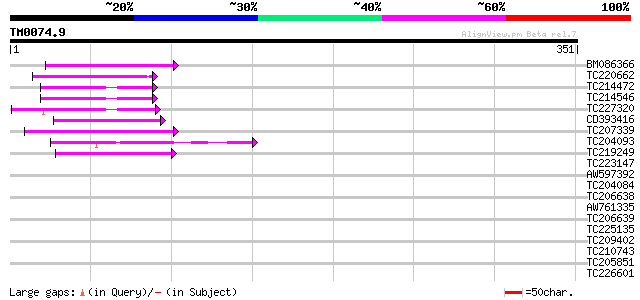
Score E
Sequences producing significant alignments: (bits) Value
BM086366 46 3e-05
TC220662 weakly similar to UP|RNP1_YEAST (P32385) Ribonucleoprot... 44 8e-05
TC214472 similar to PIR|T01932|T01932 RNA binding protein homolo... 44 1e-04
TC214546 similar to PIR|T01932|T01932 RNA binding protein homolo... 44 1e-04
TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding prot... 43 2e-04
CD393416 weakly similar to GP|15822703|gb| RNA-binding protein p... 43 2e-04
TC207339 similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif (RR... 42 3e-04
TC204093 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprot... 42 4e-04
TC219249 weakly similar to GB|AAC00006.1|2828149|AF042385 cyclop... 42 5e-04
TC223147 weakly similar to UP|Q9LZT1 (Q9LZT1) RNA binding protei... 41 0.001
AW597392 weakly similar to GP|7630030|emb| RNA binding protein-l... 41 0.001
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 40 0.001
TC206638 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor... 40 0.001
AW761335 similar to GP|5726567|gb|A glycine-rich RNA-binding pro... 40 0.002
TC206639 weakly similar to UP|SR35_HUMAN (Q8WXF0) 35 kDa SR repr... 39 0.003
TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, ... 39 0.004
TC209402 39 0.005
TC210743 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1,... 38 0.006
TC205851 similar to GB|AAG53636.1|12407751|AF291712 initiation f... 38 0.006
TC226601 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, parti... 38 0.006
>BM086366
Length = 427
Score = 45.8 bits (107), Expect = 3e-05
Identities = 23/82 (28%), Positives = 40/82 (48%)
Frame = +2
Query: 23 AEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGI 82
A+E R+FV GL + Q+ F R+G ++ + R FGF+ F +R
Sbjct: 77 AKEENRIFVGGLSWEVTERQLEHAFARYGKILECQIMMERDTGRPRGFGFITFADRRGME 256
Query: 83 AAMRSLHGTRLNSFFLSINPAR 104
A++ +HG + +S+N A+
Sbjct: 257 DAIKEMHGREIGDRIISVNKAQ 322
>TC220662 weakly similar to UP|RNP1_YEAST (P32385) Ribonucleoprotein 1,
partial (8%)
Length = 1128
Score = 44.3 bits (103), Expect = 8e-05
Identities = 29/77 (37%), Positives = 41/77 (52%)
Frame = +1
Query: 15 SCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVL 74
S +R E +L+V L + + Q+R LF RFG VVS V K+ + F+
Sbjct: 517 SSTKRISYYESPHKLYVGNLAKTVRPEQLRDLFSRFGNVVSARVLHDFKQGNSRVYAFLS 696
Query: 75 FQSKRDGIAAMRSLHGT 91
FQS+ + AAM SL+GT
Sbjct: 697 FQSEAERDAAM-SLNGT 744
>TC214472 similar to PIR|T01932|T01932 RNA binding protein homolog - common
tobacco (fragment) {Nicotiana tabacum;} , partial (29%)
Length = 767
Score = 43.5 bits (101), Expect = 1e-04
Identities = 27/72 (37%), Positives = 39/72 (53%)
Frame = +1
Query: 20 SEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKR 79
SE N +FV GLD DT +R+ F +FG VVSV +P + GFV F ++
Sbjct: 163 SEGDLNNTTIFVGGLDSDTSDEDLRQPFLQFGEVVSVKIPV------GKGCGFVQFADRK 324
Query: 80 DGIAAMRSLHGT 91
+ A+ +L+GT
Sbjct: 325 NAEEAIHALNGT 360
>TC214546 similar to PIR|T01932|T01932 RNA binding protein homolog - common
tobacco (fragment) {Nicotiana tabacum;} , partial (68%)
Length = 1756
Score = 43.5 bits (101), Expect = 1e-04
Identities = 27/72 (37%), Positives = 39/72 (53%)
Frame = +3
Query: 20 SEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKR 79
SE N +FV GLD DT +R+ F +FG VVSV +P + GFV F ++
Sbjct: 1107 SEGDINNTTIFVGGLDSDTSDEDLRQPFLQFGEVVSVKIPV------GKGCGFVQFADRK 1268
Query: 80 DGIAAMRSLHGT 91
+ A++ L+GT
Sbjct: 1269 NAEEAIQGLNGT 1304
Score = 28.1 bits (61), Expect = 6.2
Identities = 22/94 (23%), Positives = 40/94 (42%), Gaps = 10/94 (10%)
Frame = +3
Query: 10 PPPRWSCHERSEVA----------EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVP 59
PPP+ S H + A +E +++ L + + F G VVS V
Sbjct: 408 PPPQSSHHHHHKQAAAAAAAAASSDEIRTVWLGDLHHWMDENYLHNCFAHTGEVVSAKVI 587
Query: 60 KTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRL 93
+ + + E +GFV F S+ +++ +GT +
Sbjct: 588 RNKQTGQSEGYGFVEFYSRATAEKVLQNYNGTMM 689
>TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding protein, partial
(70%)
Length = 1724
Score = 42.7 bits (99), Expect = 2e-04
Identities = 29/102 (28%), Positives = 48/102 (46%), Gaps = 10/102 (9%)
Frame = +1
Query: 2 RPTGSARAPPPRWSCHER----------SEVAEENIRLFVDGLDRDTKHSQVRKLFERFG 51
RP A P + S H++ SE N +FV GLD + +R+ F ++G
Sbjct: 913 RPMRIGAATPRKSSGHQQGGLSNGTANQSEADSTNTTIFVGGLDPNVSDEDLRQPFSQYG 1092
Query: 52 TVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRL 93
+VSV +P + GFV F ++ + A++ L+GT +
Sbjct: 1093 EIVSVKIPV------GKGCGFVQFANRNNAEEALQKLNGTTI 1200
>CD393416 weakly similar to GP|15822703|gb| RNA-binding protein precursor
{Nicotiana tabacum}, partial (29%)
Length = 733
Score = 42.7 bits (99), Expect = 2e-04
Identities = 23/69 (33%), Positives = 36/69 (51%)
Frame = -2
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LFV G+ T +R+ F R+G V+ V V + R FGFV F + D +A++
Sbjct: 678 KLFVGGISYSTDDMSLRESFARYGEVIDVKVIMDRETGRSRGFGFVTFATSEDASSAIQG 499
Query: 88 LHGTRLNSF 96
+ G L+ F
Sbjct: 498 MDGQVLSVF 472
>TC207339 similar to UP|Q84ZB9 (Q84ZB9) RNA recognition motif
(RRM)-containing protein-like, partial (63%)
Length = 838
Score = 42.4 bits (98), Expect = 3e-04
Identities = 28/95 (29%), Positives = 42/95 (43%)
Frame = +3
Query: 10 PPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREK 69
PPP + AE N LFV GL + T ++R+ F +FG VV V +
Sbjct: 150 PPPGATPTPTRPTAEPNPNLFVSGLSKRTNTERLREEFAKFGEVVHARVVTDRVSGYSKG 329
Query: 70 FGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPAR 104
FGFV + + D + + G L+ + + AR
Sbjct: 330 FGFVQYATIEDAAKGIEGMDGKFLDGWVIFAEYAR 434
>TC204093 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A,
chloroplast precursor (CP29A), partial (63%)
Length = 1403
Score = 42.0 bits (97), Expect = 4e-04
Identities = 37/130 (28%), Positives = 55/130 (41%), Gaps = 2/130 (1%)
Frame = +2
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGT--VVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
+++LFV L + +Q+ +LFE G VV V KT R R FGFV S + A
Sbjct: 326 DLKLFVGNLPFNVDSAQLAELFESAGNVEVVEVIYDKTTGRSR--GFGFVTMSSVEEAEA 499
Query: 84 AMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAPFS 143
A + +G L+ L +N PP ++E P + R G+ F
Sbjct: 500 AAQQFNGYELDGRALRVNS----------GPPPARNESAP-----------RFRGGSSFG 616
Query: 144 GTGGFSQESQ 153
GG +S+
Sbjct: 617 SRGGGPSDSE 646
>TC219249 weakly similar to GB|AAC00006.1|2828149|AF042385 cyclophilin-33A
{Homo sapiens;} , partial (28%)
Length = 1185
Score = 41.6 bits (96), Expect = 5e-04
Identities = 24/75 (32%), Positives = 34/75 (45%)
Frame = +1
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V GL + S + F FG + V P + FGFV F + D AAM ++
Sbjct: 109 LYVGGLAEEVNESILHAAFIPFGDIKDVKTPLDQATQKHRSFGFVTFLEREDASAAMDNM 288
Query: 89 HGTRLNSFFLSINPA 103
G L L++N A
Sbjct: 289 DGAELYGRVLTVNYA 333
>TC223147 weakly similar to UP|Q9LZT1 (Q9LZT1) RNA binding protein-like
(At3g46020), partial (73%)
Length = 622
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/110 (25%), Positives = 47/110 (42%), Gaps = 19/110 (17%)
Frame = +3
Query: 13 RWSCHERS-EVAEENIRL------------------FVDGLDRDTKHSQVRKLFERFGTV 53
+W CH S + + +R+ F GL T Q++KLF FG V
Sbjct: 24 KWRCHRESGQSSSSTVRVILLNQIQAQPLPHHCFVSFFSGLSFYTTQEQLKKLFSPFGLV 203
Query: 54 VSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPA 103
+ R + FGFV F+S+ + A ++++G +N + + PA
Sbjct: 204 TQADLALDPITKRPKGFGFVSFKSEIEAEKACKAMNGRIVNGRLILVEPA 353
>AW597392 weakly similar to GP|7630030|emb| RNA binding protein-like
{Arabidopsis thaliana}, partial (78%)
Length = 392
Score = 40.8 bits (94), Expect = 0.001
Identities = 24/76 (31%), Positives = 39/76 (50%)
Frame = +2
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LFV L T Q++KLF FG V + R + FGFV F+S+ + A ++
Sbjct: 80 KLFVHRLSFYTTQEQLKKLFSPFGLVTQADLALDPITKRPKGFGFVSFKSEIEAEKACKA 259
Query: 88 LHGTRLNSFFLSINPA 103
++G +N + + PA
Sbjct: 260 MNGRIVNGRLILVEPA 307
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 40.4 bits (93), Expect = 0.001
Identities = 36/130 (27%), Positives = 54/130 (40%), Gaps = 2/130 (1%)
Frame = -1
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTV--VSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
+++LFV L +++ +LFE G V V V KT R R FGFV S + A
Sbjct: 884 DLKLFVGNLPFSVDSARLAELFESAGNVEVVEVIYDKTTGRSRG--FGFVTMSSVEEAEA 711
Query: 84 AMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAPFS 143
A + +G L+ L +N PP ++E P + R G+ F
Sbjct: 710 AAKQFNGYELDGRSLRVNS----------GPPPARNESAP-----------RFRGGSSFG 594
Query: 144 GTGGFSQESQ 153
GG +S+
Sbjct: 593 SRGGGPSDSE 564
>TC206638 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor SCL30a, 30a
kD, partial (56%)
Length = 1050
Score = 40.4 bits (93), Expect = 0.001
Identities = 41/175 (23%), Positives = 66/175 (37%)
Frame = +3
Query: 2 RPTGSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKT 61
R G R+P PR R + L V L D + +R+ F +FG + +++PK
Sbjct: 96 RRGGGGRSPSPRGRYPPRPRQQDLPTSLLVRNLRHDCRPEDLRRPFGQFGPLKDIYLPKD 275
Query: 62 IKRVRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEE 121
FGFV + D A + G L L++ F + K P + E
Sbjct: 276 YYTGEPRGFGFVQYVDPADAADAKYHMDGQVLLGRELTV---VFAEENRK-KPTEMRTRE 443
Query: 122 LPGRKKIRQEWRIKHRVGAPFSGTGGFSQESQKVWRVKKSTRKSPEKKVEEQERS 176
GR R+ ++ +S + +S+ R + +R K E RS
Sbjct: 444 RRGRFSDRRRSPPRYSRSPRYSRSPRYSRSPPPRHRSRSHSRDYYSPKRREYSRS 608
>AW761335 similar to GP|5726567|gb|A glycine-rich RNA-binding protein
{Glycine max}, partial (38%)
Length = 313
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/77 (28%), Positives = 41/77 (52%)
Frame = -1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
R VDGL R +H V++ F ++G + + + R + FGFV F S++ AA+
Sbjct: 289 RRMVDGLGRVIEHQAVQRAFSQYGDICESKIHQQS*DCRSKGFGFVTFASEQSMKAAIAV 110
Query: 88 LHGTRLNSFFLSINPAR 104
++G L+ +++ A+
Sbjct: 109 MNGQTLSRCNITVTTAQ 59
>TC206639 weakly similar to UP|SR35_HUMAN (Q8WXF0) 35 kDa SR repressor
protein (SRrp35), partial (23%)
Length = 438
Score = 39.3 bits (90), Expect = 0.003
Identities = 24/86 (27%), Positives = 35/86 (39%)
Frame = +1
Query: 5 GSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKR 64
G R+P PR R + L V L D + +R+ F +FG + +++PK
Sbjct: 169 GGGRSPSPRGRYPPRPRQQDLPTSLLVRNLRHDCRPEDLRRPFGQFGPLKDIYLPKDYYT 348
Query: 65 VRREKFGFVLFQSKRDGIAAMRSLHG 90
FGFV F D A + G
Sbjct: 349 GEPRGFGFVQFVDPADAADAKYHMDG 426
>TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(97%)
Length = 2392
Score = 38.9 bits (89), Expect = 0.004
Identities = 23/67 (34%), Positives = 37/67 (54%)
Frame = +3
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V LD + +Q+ LF + G VVSV V + + R +G+V F + +D A+ L
Sbjct: 186 LYVGDLDPNVTDAQLYDLFNQLGQVVSVRVCRDLTSRRSLGYGYVNFSNPQDAARALDVL 365
Query: 89 HGTRLNS 95
+ T LN+
Sbjct: 366 NFTPLNN 386
>TC209402
Length = 1163
Score = 38.5 bits (88), Expect = 0.005
Identities = 21/100 (21%), Positives = 46/100 (46%), Gaps = 2/100 (2%)
Frame = +1
Query: 7 ARAPPPRWSCHERSEVAEENI--RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKR 64
+ AP W+ SE + ++ ++V L + ++++LFE G + V +P
Sbjct: 151 SNAPTVSWADPRNSESSAISLVKSVYVKNLPENITQDRLKELFEHHGKITKVVLPSAKSG 330
Query: 65 VRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPAR 104
+ +FGFV F + + A+++ ++ L + A+
Sbjct: 331 QEKSRFGFVHFAERSSAMKALKNTEKYEIDGQLLECSLAK 450
>TC210743 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(53%)
Length = 452
Score = 38.1 bits (87), Expect = 0.006
Identities = 22/80 (27%), Positives = 40/80 (49%)
Frame = +3
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAA 84
E R F+ GL T +++ FE+FG ++ V R FGFV F K+ A
Sbjct: 75 EEYRCFIGGLAWSTSDRKLKDTFEKFGKLIEAKVVVDKFSGRSRGFGFVTFDDKKAMDEA 254
Query: 85 MRSLHGTRLNSFFLSINPAR 104
+ +++G L+ ++++ A+
Sbjct: 255 IDAMNGMDLDGRTITVDRAQ 314
>TC205851 similar to GB|AAG53636.1|12407751|AF291712 initiation factor 3g
{Arabidopsis thaliana;} , partial (45%)
Length = 641
Score = 38.1 bits (87), Expect = 0.006
Identities = 24/77 (31%), Positives = 37/77 (47%)
Frame = +1
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
E ++R V L DT+ + +LF FG V V+V K FGFV F ++ D
Sbjct: 205 ENSVR--VTNLSEDTREPDLLELFRPFGPVSRVYVAIDQKTGMSRGFGFVNFVNREDAQR 378
Query: 84 AMRSLHGTRLNSFFLSI 100
A+ L+G ++ L +
Sbjct: 379 AINKLNGYGYDNLILRV 429
>TC226601 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, partial (31%)
Length = 1584
Score = 38.1 bits (87), Expect = 0.006
Identities = 21/82 (25%), Positives = 40/82 (48%), Gaps = 4/82 (4%)
Frame = +1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
++FV GL +TK +++ F++FG ++ V R + +GFV+F+ I A +
Sbjct: 433 KIFVGGLAWETKRDTLKRYFDQFGEILEAVVITDRITGRSKGYGFVIFRDPNSAIRACHN 612
Query: 88 ----LHGTRLNSFFLSINPARF 105
+ G R N ++ +F
Sbjct: 613 PYPVIDGRRANCNLAALGAQKF 678
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,101,866
Number of Sequences: 63676
Number of extensions: 211481
Number of successful extensions: 1234
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 1228
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1233
length of query: 351
length of database: 12,639,632
effective HSP length: 98
effective length of query: 253
effective length of database: 6,399,384
effective search space: 1619044152
effective search space used: 1619044152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0074.9


