

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.3
(267 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
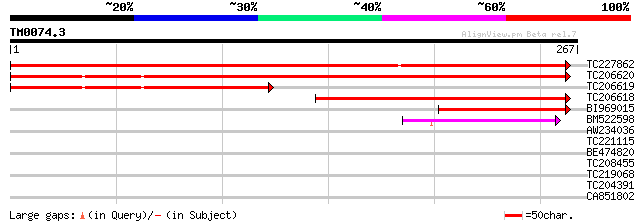
TC227862 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extr... 491 e-139
TC206620 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extr... 414 e-116
TC206619 weakly similar to UP|Q70FG8 (Q70FG8) 4,5 dioxygenase ex... 186 7e-48
TC206618 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extr... 184 3e-47
BI969015 105 3e-23
BM522598 44 6e-05
AW234036 similar to GP|21952793|dbj P0018C10.10 {Oryza sativa (j... 39 0.002
TC221115 similar to UP|Q9MSB5 (Q9MSB5) Phosphoenolpyruvate/phosp... 28 4.3
BE474820 similar to GP|6635838|gb|A amino acid/peptide transport... 28 5.6
TC208455 similar to UP|Q9LHJ0 (Q9LHJ0) Arabidopsis thaliana geno... 27 7.3
TC219068 similar to UP|O81138 (O81138) AP2 domain family transcr... 27 9.6
TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elo... 27 9.6
CA851802 27 9.6
>TC227862 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extradiol,
partial (78%)
Length = 1061
Score = 491 bits (1265), Expect = e-139
Identities = 234/264 (88%), Positives = 251/264 (94%)
Frame = +2
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
MALKDTFYISHGSPTLSIDES+ ARKFLQSWKK+VFP RP+SILVISGHW+TAVPTVNVV
Sbjct: 86 MALKDTFYISHGSPTLSIDESIQARKFLQSWKKDVFPQRPSSILVISGHWETAVPTVNVV 265
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
DS NDTIYDFYGFPK MYQLKYPAPGAP LA+RVKELLK+ GFS VDED KRGLDHGAWV
Sbjct: 266 DSINDTIYDFYGFPKQMYQLKYPAPGAPQLARRVKELLKKSGFSHVDEDTKRGLDHGAWV 445
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERH 180
PL LMYPEADIPVCQ+S+QS DGT+HYN+GKALAPLKDEGVLI+GSGSAVHNLRALE H
Sbjct: 446 PLFLMYPEADIPVCQISIQSQQDGTYHYNLGKALAPLKDEGVLIMGSGSAVHNLRALEPH 625
Query: 181 ATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGEN 240
+TV APWA+EFDNWLK+ALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGE+
Sbjct: 626 STV-APWALEFDNWLKDALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGED 802
Query: 241 SKAKLIHSSIDLGSLSYASYQFTS 264
+KAKLIHSSI+LGSLSYASYQFTS
Sbjct: 803 AKAKLIHSSIELGSLSYASYQFTS 874
>TC206620 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extradiol,
partial (78%)
Length = 1189
Score = 414 bits (1064), Expect = e-116
Identities = 196/264 (74%), Positives = 226/264 (85%)
Frame = +1
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
+ LK+TFY+SHG+PTL++D+S+ A F SWK++ FP RP+SIL+IS HWDT VPTVNVV
Sbjct: 130 LRLKETFYLSHGAPTLAVDDSIPAWNFFNSWKEQ-FPTRPSSILIISAHWDTHVPTVNVV 306
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
NDTIYDFYGFPK MY+LKYPAPGAPHLAKRVKELL GFS VDEDKKRGLDHGAWV
Sbjct: 307 HQ-NDTIYDFYGFPKSMYKLKYPAPGAPHLAKRVKELLLGSGFSHVDEDKKRGLDHGAWV 483
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERH 180
PLLLMYPEADIPVCQLS+ SN GT+HYN+GKALAPLKDEGVLIVGSGSA HNLRA+
Sbjct: 484 PLLLMYPEADIPVCQLSISSNKGGTYHYNMGKALAPLKDEGVLIVGSGSATHNLRAIAPR 663
Query: 181 ATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGEN 240
T APWA F +WLK +LL+GRYE+VN YE+KAP+AK AHPWPDHF+PLHVA+GAAGEN
Sbjct: 664 GTPPAPWASAFMSWLKTSLLDGRYEEVNEYEEKAPYAKMAHPWPDHFFPLHVAMGAAGEN 843
Query: 241 SKAKLIHSSIDLGSLSYASYQFTS 264
SKAK++H S D GS+SYAS+ FT+
Sbjct: 844 SKAKVVHDSWDGGSMSYASFGFTT 915
>TC206619 weakly similar to UP|Q70FG8 (Q70FG8) 4,5 dioxygenase extradiol,
partial (36%)
Length = 455
Score = 186 bits (473), Expect = 7e-48
Identities = 91/124 (73%), Positives = 103/124 (82%)
Frame = +1
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
+ LK+TFY+SHG+P+L ID+S+ A F SWK++ FP +P+SILVIS HWDT VPTVNVV
Sbjct: 88 LKLKETFYLSHGAPSLVIDDSIPAWHFFNSWKEQ-FPTKPSSILVISAHWDTHVPTVNVV 264
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
D NDTIYDF GFPK MY+LKYPAPGAP LAKRVKELL GFS VDEDKKRGLDHGAWV
Sbjct: 265 DQ-NDTIYDFSGFPKSMYKLKYPAPGAPQLAKRVKELLLGSGFSHVDEDKKRGLDHGAWV 441
Query: 121 PLLL 124
PL L
Sbjct: 442 PLFL 453
>TC206618 similar to UP|Q70FG7 (Q70FG7) 4,5-DOPA dioxygenase extradiol,
partial (40%)
Length = 456
Score = 184 bits (468), Expect = 3e-47
Identities = 82/120 (68%), Positives = 103/120 (85%)
Frame = +3
Query: 145 THHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAAPWAVEFDNWLKEALLEGRY 204
T+HYN+GKALAPLKDEGVLI+GSGSA HNLRA+ + APWA F +WL+ +LL+GRY
Sbjct: 3 TYHYNMGKALAPLKDEGVLIIGSGSATHNLRAMAPRGSPPAPWASAFMSWLETSLLDGRY 182
Query: 205 EDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKLIHSSIDLGSLSYASYQFTS 264
E+VN +E+KAP+AK AHPWPDHF+PLHVA+GAAGEN+KAK++H S D GS+SYAS+ FT+
Sbjct: 183 EEVNEFEEKAPYAKMAHPWPDHFFPLHVAMGAAGENAKAKVVHDSWDAGSISYASFGFTT 362
>BI969015
Length = 733
Score = 105 bits (261), Expect = 3e-23
Identities = 44/62 (70%), Positives = 56/62 (89%)
Frame = -3
Query: 203 RYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKLIHSSIDLGSLSYASYQF 262
RYE+VN +E+KAP+AK AHPWPDHF+PLHVA+GAAGEN+KAK++H S D GS+SYAS+ F
Sbjct: 299 RYEEVNEFEEKAPYAKMAHPWPDHFFPLHVAMGAAGENAKAKVVHDSWDAGSISYASFGF 120
Query: 263 TS 264
T+
Sbjct: 119 TT 114
>BM522598
Length = 373
Score = 44.3 bits (103), Expect = 6e-05
Identities = 25/76 (32%), Positives = 39/76 (50%), Gaps = 2/76 (2%)
Frame = +1
Query: 186 PWAVEFDNWLKE--ALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
PWA EF W+ + AL E K P ++AHP +H+ P+ A+GAA K
Sbjct: 85 PWAKEFAAWVHKSAALPAPEREAALLQYAKQPTFRRAHPREEHWMPIFFALGAADSQEKV 264
Query: 244 KLIHSSIDLGSLSYAS 259
+++ I GSL+ ++
Sbjct: 265 QILADRILCGSLAVST 312
>AW234036 similar to GP|21952793|dbj P0018C10.10 {Oryza sativa (japonica
cultivar-group)}, partial (6%)
Length = 407
Score = 39.3 bits (90), Expect = 0.002
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = +2
Query: 78 YQLKYPAPGAPHLAKRVKE 96
+QLKYPAPGAP LAKRVKE
Sbjct: 350 WQLKYPAPGAPQLAKRVKE 406
>TC221115 similar to UP|Q9MSB5 (Q9MSB5) Phosphoenolpyruvate/phosphate
translocator, partial (18%)
Length = 515
Score = 28.1 bits (61), Expect = 4.3
Identities = 10/32 (31%), Positives = 19/32 (59%)
Frame = +2
Query: 224 PDHFYPLHVAIGAAGENSKAKLIHSSIDLGSL 255
P HF+P H + E+S + ++++LG+L
Sbjct: 218 PSHFHPFHARATSVPESSAGNTLLNTLELGAL 313
>BE474820 similar to GP|6635838|gb|A amino acid/peptide transporter {Prunus
dulcis}, partial (17%)
Length = 319
Score = 27.7 bits (60), Expect = 5.6
Identities = 12/26 (46%), Positives = 17/26 (65%), Gaps = 1/26 (3%)
Frame = +2
Query: 209 HYEQKAP-HAKKAHPWPDHFYPLHVA 233
H ++K P +A K WP HF P+HV+
Sbjct: 164 HRQRKGPFNAAKNGNWPFHFSPVHVS 241
>TC208455 similar to UP|Q9LHJ0 (Q9LHJ0) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MMP21, partial (18%)
Length = 1350
Score = 27.3 bits (59), Expect = 7.3
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +1
Query: 116 HGAWVPLLLMYPEADIPVCQLS 137
H W+PL+ M PE + CQLS
Sbjct: 355 HQIWLPLVQMLPELLVRKCQLS 420
>TC219068 similar to UP|O81138 (O81138) AP2 domain family transcription
factor homolog (AP2 domain transcription factor)
(ABI4:abscisic acid-insensitive 4 ) (ABI4), partial
(25%)
Length = 1064
Score = 26.9 bits (58), Expect = 9.6
Identities = 16/53 (30%), Positives = 23/53 (43%)
Frame = -2
Query: 186 PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAG 238
PW + + + +L RY DV + K PH AH +Y + AAG
Sbjct: 679 PWKTKPNPRRRNSLPPPRYSDVAAVDSKKPHVALAHAHGYCYY*YYPENHAAG 521
>TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation
factor EF-2), complete
Length = 2941
Score = 26.9 bits (58), Expect = 9.6
Identities = 16/39 (41%), Positives = 19/39 (48%), Gaps = 1/39 (2%)
Frame = -2
Query: 3 LKDTFYISHGSPTLSIDE-SLVARKFLQSWKKEVFPPRP 40
L D PTL+ S++A FL W KEVF P P
Sbjct: 963 LLDGLVTELNKPTLAGSRTSILAGPFLCGWIKEVFTPEP 847
>CA851802
Length = 612
Score = 26.9 bits (58), Expect = 9.6
Identities = 12/27 (44%), Positives = 17/27 (62%), Gaps = 1/27 (3%)
Frame = +1
Query: 211 EQKAPH-AKKAHPWPDHFYPLHVAIGA 236
E++APH A +PDHF+ LH I +
Sbjct: 64 EKRAPHPAATGTSFPDHFFVLHCRIAS 144
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,031,523
Number of Sequences: 63676
Number of extensions: 194057
Number of successful extensions: 925
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 901
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 920
length of query: 267
length of database: 12,639,632
effective HSP length: 96
effective length of query: 171
effective length of database: 6,526,736
effective search space: 1116071856
effective search space used: 1116071856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0074.3


