

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0067a.13
(212 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
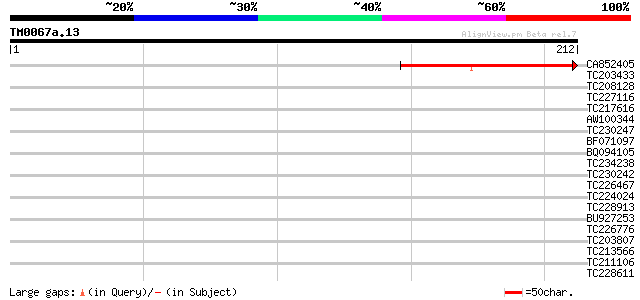
Score E
Sequences producing significant alignments: (bits) Value
CA852405 weakly similar to GP|24461860|gb CTV.20 {Poncirus trifo... 55 2e-08
TC203433 similar to PIR|T09390|T09390 21K protein precursor - al... 37 0.007
TC208128 homologue to UP|Q8V7F3 (Q8V7F3) ORF1 (Fragment), partia... 36 0.011
TC227116 similar to UP|WAS2_HUMAN (Q9Y6W5) Wiskott-Aldrich syndr... 36 0.011
TC217616 similar to GB|AAM63479.1|21554372|AY086477 phospholipas... 35 0.025
AW100344 similar to PIR|T09390|T093 21K protein precursor - alfa... 35 0.033
TC230247 weakly similar to GB|AAP88326.1|32815835|BT009692 At1g6... 35 0.033
BF071097 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 35 0.033
BQ094105 homologue to GP|13591696|gb| histone acetyltransferase ... 33 0.073
TC234238 weakly similar to UP|Q9ZUS6 (Q9ZUS6) Chloroplast lumen ... 33 0.073
TC230242 weakly similar to UP|Q9FQZ6 (Q9FQZ6) Avr9/Cf-9 rapidly ... 33 0.096
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 33 0.096
TC224024 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, part... 33 0.096
TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partia... 33 0.096
BU927253 similar to GP|14194119|gb| At1g01540/F22L4_6 {Arabidops... 33 0.12
TC226776 similar to GB|AAP68287.1|31711862|BT008848 At1g02720 {A... 33 0.12
TC203807 similar to UP|Q8L9H4 (Q8L9H4) CONSTANS-like B-box zinc ... 33 0.12
TC213566 weakly similar to UP|Q79FY9 (Q79FY9) PE-PGRS FAMILY PRO... 32 0.16
TC211106 similar to UP|RS2_HUMAN (P15880) 40S ribosomal protein ... 32 0.16
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 32 0.16
>CA852405 weakly similar to GP|24461860|gb CTV.20 {Poncirus trifoliata},
partial (0%)
Length = 674
Score = 55.5 bits (132), Expect = 2e-08
Identities = 29/68 (42%), Positives = 43/68 (62%), Gaps = 2/68 (2%)
Frame = +3
Query: 147 YEFILVDTDSIEVTHNKDKNGEIIC--SKVKILKVLNMQDWKHPLFQSKQFSREFTPTGH 204
YEFIL+DT+S+ + H KD + S ++ILKV+ + + L Q K+FS F G+
Sbjct: 30 YEFILIDTNSVSMKHFKDPKDPNLNTHSTIQILKVMQPRHYGSNLNQPKKFSAPFDLAGY 209
Query: 205 NYFDYMDA 212
NY+DY+DA
Sbjct: 210 NYWDYIDA 233
>TC203433 similar to PIR|T09390|T09390 21K protein precursor - alfalfa
{Medicago sativa;} , partial (88%)
Length = 825
Score = 37.0 bits (84), Expect = 0.007
Identities = 25/71 (35%), Positives = 35/71 (49%), Gaps = 3/71 (4%)
Frame = +2
Query: 6 TSRDQYGPP---LGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQ 62
+S PP L S+ P+S + +AP + P + P PSTP SS+ PT S
Sbjct: 65 SSSSSSSPPST*LPPSQPPPTSSSPPAAPPSTQP--SASSPSPSTPPPSSKTPTSSSRQP 238
Query: 63 SPAKSKSPLVP 73
SP+ S +P P
Sbjct: 239 SPSPSTTPRPP 271
>TC208128 homologue to UP|Q8V7F3 (Q8V7F3) ORF1 (Fragment), partial (19%)
Length = 711
Score = 36.2 bits (82), Expect = 0.011
Identities = 24/63 (38%), Positives = 31/63 (49%)
Frame = -1
Query: 1 MSEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQI 60
MS K S PP S + P ++T S PVA L++I P PS PT S + P V
Sbjct: 543 MSAKSCSLSWLKPPPIS--RPPPTLTHKSPPVATQILQSIPSPSPSPPTTSFRSPQVPSF 370
Query: 61 VQS 63
Q+
Sbjct: 369 SQT 361
>TC227116 similar to UP|WAS2_HUMAN (Q9Y6W5) Wiskott-Aldrich syndrome protein
family member 2 (WASP-family protein member 2)
(Verprolin homology domain-containing protein 2),
partial (5%)
Length = 1154
Score = 36.2 bits (82), Expect = 0.011
Identities = 24/87 (27%), Positives = 36/87 (40%)
Frame = +3
Query: 4 KPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQS 63
+P+ + PP SS +++ SS P+ P + PP S P S +
Sbjct: 183 QPSPKTTPNPP--SSTSFSKTLSYSSTPLPQNPAPS--PPSTSPPPSSPPSTPTASFAAP 350
Query: 64 PAKSKSPLVPLSNKYSILDYQNTISSK 90
AK+K P P N Y Y I+S+
Sbjct: 351 SAKTKLPPTPKPNNYPANTYTTPIASR 431
>TC217616 similar to GB|AAM63479.1|21554372|AY086477 phospholipase-like
protein {Arabidopsis thaliana;} , partial (95%)
Length = 1400
Score = 35.0 bits (79), Expect = 0.025
Identities = 21/68 (30%), Positives = 32/68 (46%)
Frame = +1
Query: 14 PLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSPLVP 73
P S P+S GSS+ T + + PP PST T++ PT S + + S
Sbjct: 358 PSPSCTDTPASPAGSSSSPPSTSPRPVSPPAPST-TRAMASPTASSLTSPTSTPSSTTAS 534
Query: 74 LSNKYSIL 81
S++ S+L
Sbjct: 535 PSSRISVL 558
>AW100344 similar to PIR|T09390|T093 21K protein precursor - alfalfa,
partial (26%)
Length = 364
Score = 34.7 bits (78), Expect = 0.033
Identities = 21/56 (37%), Positives = 30/56 (53%)
Frame = +3
Query: 15 LGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSP 70
L S+ P+S + +AP + P + P PSTP SS+ PT S SP+ S +P
Sbjct: 33 LPPSQPPPTSSSPPAAPPSTQP--SAYSPSPSTPPPSSKTPTSSSRQSSPSPSTTP 194
>TC230247 weakly similar to GB|AAP88326.1|32815835|BT009692 At1g61870
{Arabidopsis thaliana;} , partial (56%)
Length = 1291
Score = 34.7 bits (78), Expect = 0.033
Identities = 27/98 (27%), Positives = 37/98 (37%), Gaps = 18/98 (18%)
Frame = +1
Query: 13 PPLGSSRKLPSSITGSSAPVAVT-------------PLKTIKPPGPSTPTKSSQRPT--- 56
PP S PSS AP + T P+ +I P PS T P+
Sbjct: 250 PPTTSPASAPSSTISKPAPTSATKNSSPTPSSSTARPICSITPSAPSPKTSPPLAPSKP* 429
Query: 57 --VSQIVQSPAKSKSPLVPLSNKYSILDYQNTISSKTP 92
S + SP +KS L SN ++ + T + TP
Sbjct: 430 IPSSLLPSSPRTTKSSLESTSNSPKLIQFNQT*THTTP 543
>BF071097 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (18%)
Length = 366
Score = 34.7 bits (78), Expect = 0.033
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Frame = +2
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTV-SQIVQS 63
P+S PP SS LP S S+ +P +++ PP TP ++P V S+ V +
Sbjct: 158 PSSPPSSKPPASSSPPLPYSSCASTTHPPPSPPRSLPPPPSPTPPSPRRKPRVFSKSVSA 337
Query: 64 PAKS 67
P ++
Sbjct: 338 PTRA 349
>BQ094105 homologue to GP|13591696|gb| histone acetyltransferase GCN5
{Arabidopsis thaliana}, partial (12%)
Length = 341
Score = 33.5 bits (75), Expect = 0.073
Identities = 16/53 (30%), Positives = 24/53 (45%)
Frame = +1
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPA 65
PP P T +S+ + T L +++PP P TP + PT + PA
Sbjct: 145 PPTPPPPPPPPPSTSASSRLRTTRLPSLRPPSPPTPATAPSPPTTTSRASPPA 303
>TC234238 weakly similar to UP|Q9ZUS6 (Q9ZUS6) Chloroplast lumen common
protein family, partial (21%)
Length = 461
Score = 33.5 bits (75), Expect = 0.073
Identities = 17/51 (33%), Positives = 25/51 (48%)
Frame = +3
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRP 55
P+S PP SS PSS S+ +PL+++ PP TP ++P
Sbjct: 105 PSSPPSSKPPASSSPPPPSSSCASTTRPPSSPLRSLPPPRKPTPPSPRRKP 257
>TC230242 weakly similar to UP|Q9FQZ6 (Q9FQZ6) Avr9/Cf-9 rapidly elicited
protein 146, partial (30%)
Length = 713
Score = 33.1 bits (74), Expect = 0.096
Identities = 14/33 (42%), Positives = 21/33 (63%)
Frame = +3
Query: 23 SSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRP 55
SS + S+AP++ T PP PS+PT ++ RP
Sbjct: 318 SSTSSSAAPISTVTTTTAAPPPPSSPTTTTFRP 416
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 33.1 bits (74), Expect = 0.096
Identities = 20/71 (28%), Positives = 30/71 (42%)
Frame = +3
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
++ P + PP + PSS G+ V + P PP STP +SQ P +
Sbjct: 45 AQAPATAPANSPPTTPAATSPSS--GTPLAVTLPPTVVASPPTTSTPPTTSQPPAIVAPK 218
Query: 62 QSPAKSKSPLV 72
+P +P V
Sbjct: 219 PAPVTPPAPKV 251
>TC224024 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, partial (3%)
Length = 425
Score = 33.1 bits (74), Expect = 0.096
Identities = 25/86 (29%), Positives = 38/86 (44%), Gaps = 4/86 (4%)
Frame = +3
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQI- 60
+ P+S PPL S PSS T S + +P PP PS+ + S+ PT +
Sbjct: 153 TRSPSSSTSPPPPLSS----PSSPTPRSPSPSPSPTPPPSPPPPSSASSSANAPTYPALS 320
Query: 61 ---VQSPAKSKSPLVPLSNKYSILDY 83
+ S + S S +P + SI +
Sbjct: 321 SVQLSSSSVSASVFIPSFSTISIKSF 398
>TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partial (19%)
Length = 692
Score = 33.1 bits (74), Expect = 0.096
Identities = 24/71 (33%), Positives = 32/71 (44%), Gaps = 6/71 (8%)
Frame = +2
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTP------LKTIKPPGPSTPTKSSQRPTVS 58
P S PP SS PS+ SSAP P + P GPS+ +S +PT +
Sbjct: 71 PASPRSSSPP--SSPPSPSTTPTSSAPANAPPSSPSASTRLQTPSGPSSDASTSPKPTST 244
Query: 59 QIVQSPAKSKS 69
+P+KS S
Sbjct: 245 SSRAAPSKSPS 277
Score = 28.1 bits (61), Expect = 3.1
Identities = 17/46 (36%), Positives = 24/46 (51%)
Frame = +2
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVS 58
P SS +S+T + +P + TP T PP + T SS PT+S
Sbjct: 380 PASASSVANTASVTTAPSPPS-TPSTTTTPPPTARSTPSSSNPTLS 514
>BU927253 similar to GP|14194119|gb| At1g01540/F22L4_6 {Arabidopsis
thaliana}, partial (13%)
Length = 309
Score = 32.7 bits (73), Expect = 0.12
Identities = 25/69 (36%), Positives = 30/69 (43%), Gaps = 6/69 (8%)
Frame = +3
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAP------VAVTPLKTIKPPGPSTPTKSSQRPTVS 58
P+S G SS SS SS+P A T TI+ P P+TP P S
Sbjct: 105 PSSASASGSSSASSSDPSSSSPSSSSPSASPPAAATTTTTTIRTPAPATP------PPPS 266
Query: 59 QIVQSPAKS 67
+ QSP KS
Sbjct: 267 PLPQSPKKS 293
>TC226776 similar to GB|AAP68287.1|31711862|BT008848 At1g02720 {Arabidopsis
thaliana;} , partial (68%)
Length = 1307
Score = 32.7 bits (73), Expect = 0.12
Identities = 29/70 (41%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Frame = +2
Query: 2 SEKPTSRDQYG--PPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVS- 58
S P SR Y P SS LPSS APV V L+ + PP +TPT S P S
Sbjct: 962 SSNPASRGLYTSTPIWFSSTTLPSS----GAPV*V--LEPLAPPSTATPTSPSTSPQGSG 1123
Query: 59 QIVQSPAKSK 68
+ SPA+S+
Sbjct: 1124 RTCASPARSR 1153
>TC203807 similar to UP|Q8L9H4 (Q8L9H4) CONSTANS-like B-box zinc finger
protein-like, partial (49%)
Length = 1780
Score = 32.7 bits (73), Expect = 0.12
Identities = 24/81 (29%), Positives = 37/81 (45%), Gaps = 2/81 (2%)
Frame = +2
Query: 17 SSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSPLVPLSN 76
SS+ P ++GSS+P + T PP TP S R + S S +P
Sbjct: 470 SSKPPPPPLSGSSSPPTMALPPTSSPPTTPTPPPGSSRTPI-----SGPNSWTPPKSSPR 634
Query: 77 KYSILDY--QNTISSKTPGKT 95
+YS L + + + ++TP KT
Sbjct: 635 RYSSLKWTPSSILITQTPSKT 697
>TC213566 weakly similar to UP|Q79FY9 (Q79FY9) PE-PGRS FAMILY PROTEIN,
partial (4%)
Length = 676
Score = 32.3 bits (72), Expect = 0.16
Identities = 31/129 (24%), Positives = 48/129 (37%), Gaps = 29/129 (22%)
Frame = -3
Query: 13 PPLGSSRKLPSSITGSSAPVA-VTPLKTIKPPGPSTPTKS-------------------- 51
PP+ + LP I PV VTP + KPP P P K
Sbjct: 614 PPV--PKTLPPKIAPVKPPVPKVTPALSPKPPSPKIPPKKAPISPPPLPLPPADAPPLPT 441
Query: 52 -------SQRPTVSQIVQSPAKSKSPLVPLSNKYSILDYQNTIS-SKTPGKTEQYEYQEK 103
S+ PT ++ +PA + PL++ + D + T S + +P + Y +
Sbjct: 440 PKVSPTPSKAPTPAKETPAPAPAHKKKAPLADDIATTDSEETPSPAPSPNANGAHSYYRQ 261
Query: 104 PECLPSVMI 112
PS+ I
Sbjct: 260 GNMWPSINI 234
>TC211106 similar to UP|RS2_HUMAN (P15880) 40S ribosomal protein S2 (S4)
(LLREP3 protein), partial (6%)
Length = 978
Score = 32.3 bits (72), Expect = 0.16
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 13/76 (17%)
Frame = +2
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKT-----IKPPGPS---TPTKSSQRPTVSQIVQSP 64
PPL ++ PSS GSS P P K+ + PP S PT T+S+ +
Sbjct: 113 PPLTAAAAAPSSPPGSSPPTPAPPSKSPLHRRLPPPATSP*CPPTTPKNASTLSKTRSAT 292
Query: 65 AKSKS-----PLVPLS 75
A S+S P+ P S
Sbjct: 293 AASESHSPTPPVTPFS 340
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 32.3 bits (72), Expect = 0.16
Identities = 26/72 (36%), Positives = 31/72 (42%), Gaps = 14/72 (19%)
Frame = +3
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKTIKP--------------PGPSTPTKSSQRPTVS 58
PP R SS T SS P AVTP + P P P+TP+ SS P
Sbjct: 174 PPPPPPRSASSSTTASS-PTAVTPPSSSPPRPSPGPSATPTSTSPSPATPSTSSSAPRPP 350
Query: 59 QIVQSPAKSKSP 70
+ SPA + SP
Sbjct: 351 --LSSPAPTPSP 380
Score = 27.7 bits (60), Expect = 4.0
Identities = 18/56 (32%), Positives = 26/56 (46%)
Frame = +3
Query: 4 KPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQ 59
+P S Y PP PS + +S+P A +P K P TP+ S P+ S+
Sbjct: 471 EPPSPPPY-PPTSPPPPCPSLKSPNSSPNAASPSKNSSPSVGPTPSDSLTAPSSSR 635
Score = 26.6 bits (57), Expect = 8.9
Identities = 21/70 (30%), Positives = 28/70 (40%), Gaps = 1/70 (1%)
Frame = +3
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRP-TVSQI 60
S PTS +S P S AP +P T PP P+T + S P + S
Sbjct: 279 SATPTSTSPSPATPSTSSSAPRPPLSSPAPTP-SPAPTFSPPPPATSSPCSAAPSSPSSS 455
Query: 61 VQSPAKSKSP 70
+ A+ SP
Sbjct: 456 AAATAEPPSP 485
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.130 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,979,111
Number of Sequences: 63676
Number of extensions: 152068
Number of successful extensions: 1349
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 1229
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1328
length of query: 212
length of database: 12,639,632
effective HSP length: 93
effective length of query: 119
effective length of database: 6,717,764
effective search space: 799413916
effective search space used: 799413916
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0067a.13


