

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0066.12
(163 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
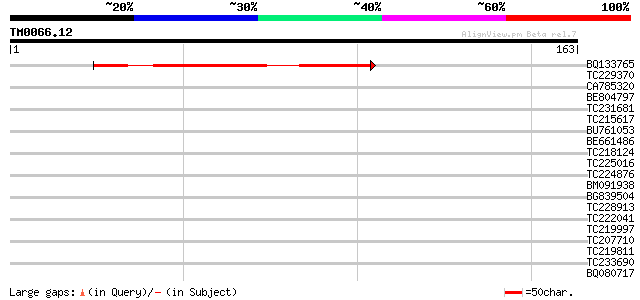
Score E
Sequences producing significant alignments: (bits) Value
BQ133765 90 4e-19
TC229370 similar to UP|Q9LSA0 (Q9LSA0) Emb|CAB62463.1, partial (7%) 35 0.012
CA785320 35 0.021
BE804797 33 0.046
TC231681 32 0.13
TC215617 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase ... 32 0.13
BU761053 similar to GP|22530930|gb| vacuolar sorting receptor-li... 32 0.13
BE661486 similar to PIR|T51625|T516 MAP3K alpha protein kinase (... 32 0.13
TC218124 32 0.18
TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%) 31 0.23
TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial ... 31 0.23
BM091938 31 0.23
BG839504 similar to GP|20466754|gb Isp4-like protein {Arabidopsi... 31 0.30
TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partia... 31 0.30
TC222041 similar to UP|Q8LBQ6 (Q8LBQ6) Isp4-like protein, partia... 30 0.39
TC219997 similar to PIR|T12954|T12954 dUTP pyrophosphatase homol... 30 0.51
TC207710 30 0.51
TC219811 similar to UP|Q6NQC3 (Q6NQC3) At3g48120, partial (13%) 30 0.67
TC233690 weakly similar to UP|Q6K393 (Q6K393) Ankyrin 3, epithel... 30 0.67
BQ080717 29 0.87
>BQ133765
Length = 426
Score = 90.1 bits (222), Expect = 4e-19
Identities = 46/81 (56%), Positives = 56/81 (68%)
Frame = +3
Query: 25 WFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNTLTRPLRLHDQDTDPSS 84
WFD ++ LP+ PF+L+QAVCSHGLFMM PN WDPLS TL RPLR S
Sbjct: 225 WFDFDMELPS-------PFQLEQAVCSHGLFMMPPNHWDPLSKTLIRPLR---------S 356
Query: 85 SSPSFIVTVSQRSESIAVRVH 105
S SF+V++SQ S+S+AVRVH
Sbjct: 357 SPSSFLVSLSQHSQSLAVRVH 419
>TC229370 similar to UP|Q9LSA0 (Q9LSA0) Emb|CAB62463.1, partial (7%)
Length = 1134
Score = 35.4 bits (80), Expect = 0.012
Identities = 31/90 (34%), Positives = 43/90 (47%), Gaps = 1/90 (1%)
Frame = +2
Query: 2 EELEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNS 61
+E + PP +P+ P SP T+F P+ T SET P + +V ++PN
Sbjct: 107 DEYQLSPPFSPS---PSPSPP-TYFCGGNPMKTRMCSETAPVTIPHSVMGKSSRNLSPNF 274
Query: 62 WDPLSNTLTRPLRLHDQDTDPSS-SSPSFI 90
DP N+L L + TD SS SPS I
Sbjct: 275 SDPSRNSLP---PLSPRRTDGSSQESPSGI 355
>CA785320
Length = 409
Score = 34.7 bits (78), Expect = 0.021
Identities = 41/140 (29%), Positives = 60/140 (42%), Gaps = 2/140 (1%)
Frame = +3
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLS 66
PPP NP CSK S S PTT +S T R + AP+S PL+
Sbjct: 24 PPPANP-CSKTSSPSSPVTASALSAWPTTATSST--IRPSEPTSK----TTAPSSTPPLT 182
Query: 67 NTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLL--SPHEVRALMVPTS 124
RL + P ++ PSF +++ V+ G LL SP R ++P
Sbjct: 183 P------RLSSTSSPPPNTDPSFFESLT------PVKSSKGLILLFSSP---RTSLLPFE 317
Query: 125 LTASLNSWVFIGKLISVVLF 144
+ L W++ G+ +V+LF
Sbjct: 318 TLSGLGLWLW-GEGATVLLF 374
>BE804797
Length = 370
Score = 33.5 bits (75), Expect = 0.046
Identities = 19/53 (35%), Positives = 28/53 (51%)
Frame = +1
Query: 49 VCSHGLFMMAPNSWDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIA 101
+C+H ++ P S D + LTRP + DTDP SS +TV+ R + A
Sbjct: 85 ICTH*SCILKPESTDTKLHKLTRP---NAVDTDPHSSHSLHHITVANRKQPFA 234
>TC231681
Length = 784
Score = 32.0 bits (71), Expect = 0.13
Identities = 17/37 (45%), Positives = 21/37 (55%)
Frame = +1
Query: 8 PPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFR 44
PP +P+ SSP S W D+ LP PT T+S T R
Sbjct: 100 PPLSPSLP*SASSPPSIW-DL*LPRPTLTASTTTQIR 207
>TC215617 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase kinase
kinase alpha, partial (51%)
Length = 1451
Score = 32.0 bits (71), Expect = 0.13
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +3
Query: 11 NPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAV 49
+P CS+ SS ++ D LPLP + S T+ F ++Q +
Sbjct: 192 SPRCSRDFSSSAAAAVDQGLPLPRPSVSSTQSFGIEQGL 308
>BU761053 similar to GP|22530930|gb| vacuolar sorting receptor-like protein
{Arabidopsis thaliana}, partial (12%)
Length = 446
Score = 32.0 bits (71), Expect = 0.13
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = -2
Query: 19 SSPSSTWFDMELPLPTTTSSETEPFRLDQAVC 50
SSP TW P PTTT+ + +P ++ VC
Sbjct: 193 SSPPQTWHQQPPPPPTTTTKQPQPRKVSWFVC 98
>BE661486 similar to PIR|T51625|T516 MAP3K alpha protein kinase (EC 2.7.1.-)
[imported] - Arabidopsis thaliana, partial (6%)
Length = 805
Score = 32.0 bits (71), Expect = 0.13
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +3
Query: 11 NPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAV 49
+P CS+ SS ++ D LPLP + S T+ F ++Q +
Sbjct: 279 SPRCSRDFSSSAAAAVDQGLPLPRPSVSSTQSFGIEQGL 395
>TC218124
Length = 658
Score = 31.6 bits (70), Expect = 0.18
Identities = 23/71 (32%), Positives = 30/71 (41%), Gaps = 9/71 (12%)
Frame = +2
Query: 62 WDPLSNTLTRPLRLHDQDTDPSS---------SSPSFIVTVSQRSESIAVRVHHGTHLLS 112
W PLS + + P RL T P S SSP+ T S RSE + + T
Sbjct: 203 WVPLSPSTSPPPRLRTTPTSPVSPPPPTDRTDSSPATPPTASPRSEPLTTAMTTKTMSFR 382
Query: 113 PHEVRALMVPT 123
P E +A + T
Sbjct: 383 PKETKATKMKT 415
>TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%)
Length = 973
Score = 31.2 bits (69), Expect = 0.23
Identities = 22/67 (32%), Positives = 33/67 (48%)
Frame = +1
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L P P P+ P S+PS + + LPLPTT+S E + C L +P+
Sbjct: 280 LPSPSPSPPSSPSPTSTPSPS---LRLPLPTTSS*EAIQVPISATSCP-SLPSPSPSLRP 447
Query: 64 PLSNTLT 70
LS+T++
Sbjct: 448 LLSSTIS 468
>TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial (66%)
Length = 1340
Score = 31.2 bits (69), Expect = 0.23
Identities = 22/67 (32%), Positives = 33/67 (48%)
Frame = +2
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L P P P+ P S+PS + + LPLPTT+S E + C L +P+
Sbjct: 872 LPSPSPSPPSSPSPTSTPSPS---LRLPLPTTSS*EAIQVPISATSCP-SLPSPSPSLRP 1039
Query: 64 PLSNTLT 70
LS+T++
Sbjct: 1040LLSSTIS 1060
>BM091938
Length = 421
Score = 31.2 bits (69), Expect = 0.23
Identities = 22/67 (32%), Positives = 33/67 (48%)
Frame = +1
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L P P P+ P S+PS + + LPLPTT+S E + C L +P+
Sbjct: 154 LPSPSPSPPSSPSPTSTPSPS---LRLPLPTTSS*EAIQVPISATSCP-SLPSPSPSLRP 321
Query: 64 PLSNTLT 70
LS+T++
Sbjct: 322 LLSSTIS 342
>BG839504 similar to GP|20466754|gb Isp4-like protein {Arabidopsis thaliana},
partial (15%)
Length = 499
Score = 30.8 bits (68), Expect = 0.30
Identities = 9/30 (30%), Positives = 20/30 (66%)
Frame = +3
Query: 120 MVPTSLTASLNSWVFIGKLISVVLFRFRLR 149
M+P + + N+W+F+G + + +FR+R +
Sbjct: 72 MMPPATPLNYNAWIFVGTIFNFFIFRYRXK 161
>TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partial (19%)
Length = 692
Score = 30.8 bits (68), Expect = 0.30
Identities = 23/81 (28%), Positives = 31/81 (37%)
Frame = +2
Query: 8 PPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSN 67
PP +P S P SSP P P+TT + + P + S + P+ P S+
Sbjct: 68 PPASPRSSSPPSSP---------PSPSTTPTSSAPANAPPSSPSASTRLQTPSG--PSSD 214
Query: 68 TLTRPLRLHDQDTDPSSSSPS 88
T P S SPS
Sbjct: 215 ASTSPKPTSTSSRAAPSKSPS 277
Score = 30.4 bits (67), Expect = 0.39
Identities = 28/107 (26%), Positives = 39/107 (36%), Gaps = 1/107 (0%)
Frame = +2
Query: 8 PPCNPNCSKPESSPSSTWFDMEL-PLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLS 66
PP +P+ S +PS D P PT+TSS P + +P++W S
Sbjct: 155 PPSSPSASTRLQTPSGPSSDASTSPKPTSTSSRAAPSK-------------SPSTWPSAS 295
Query: 67 NTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSP 113
+ Q PS+S+ S S S S T SP
Sbjct: 296 LATSMSSPASPQPRVPSASTSSTTSAASPASASSVANTASVTTAPSP 436
>TC222041 similar to UP|Q8LBQ6 (Q8LBQ6) Isp4-like protein, partial (38%)
Length = 1012
Score = 30.4 bits (67), Expect = 0.39
Identities = 9/28 (32%), Positives = 19/28 (67%)
Frame = +2
Query: 120 MVPTSLTASLNSWVFIGKLISVVLFRFR 147
M+P + + N+W+F+G + + +FR+R
Sbjct: 569 MMPPATPLNYNAWIFVGTIFNFFIFRYR 652
>TC219997 similar to PIR|T12954|T12954 dUTP pyrophosphatase homolog T6H20.30
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (84%)
Length = 772
Score = 30.0 bits (66), Expect = 0.51
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = +3
Query: 13 NCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAP 59
+CSK +SS + F LP + SS T PF C GL + P
Sbjct: 81 HCSKSQSSTKTATFPQPLPFSASRSSPTRPF------CPLGLPLSPP 203
>TC207710
Length = 1546
Score = 30.0 bits (66), Expect = 0.51
Identities = 16/46 (34%), Positives = 24/46 (51%), Gaps = 4/46 (8%)
Frame = +2
Query: 75 LHDQDTDPSSSSPSFIVTVSQRSESIAV----RVHHGTHLLSPHEV 116
LHD+DT P++ SP + S +E AV HH T+ H++
Sbjct: 767 LHDRDTAPTTCSPKAALAGSCTAEKSAVVAVGSTHHSTNTQVHHQI 904
>TC219811 similar to UP|Q6NQC3 (Q6NQC3) At3g48120, partial (13%)
Length = 567
Score = 29.6 bits (65), Expect = 0.67
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = +1
Query: 5 EKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEP 42
+ PPP +PN SKP T P+TTSS P
Sbjct: 196 QNPPPSSPNSSKPPPLRLQTLRSKRSFNPSTTSSRKSP 309
>TC233690 weakly similar to UP|Q6K393 (Q6K393) Ankyrin 3, epithelial isoform
a-like, partial (13%)
Length = 1024
Score = 29.6 bits (65), Expect = 0.67
Identities = 13/37 (35%), Positives = 18/37 (48%)
Frame = +1
Query: 8 PPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFR 44
P C+ SKP + S W L +P +TS+ P R
Sbjct: 268 PKCHSASSKPPTPAISPWHSAALSIPPSTSTSPAPSR 378
>BQ080717
Length = 426
Score = 29.3 bits (64), Expect = 0.87
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 2/46 (4%)
Frame = +2
Query: 7 PPPCNPNCSKPESSPS--STWFDMELPLPTTTSSETEPFRLDQAVC 50
P PC + P SSPS ST P P T++S + FR + C
Sbjct: 188 PSPCPAMSTSPPSSPSAPSTPSPSGSPTPPTSTSPSPSFRCSKPSC 325
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.132 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,054,878
Number of Sequences: 63676
Number of extensions: 157678
Number of successful extensions: 1527
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 1491
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1524
length of query: 163
length of database: 12,639,632
effective HSP length: 90
effective length of query: 73
effective length of database: 6,908,792
effective search space: 504341816
effective search space used: 504341816
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0066.12


