

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0065.6
(516 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
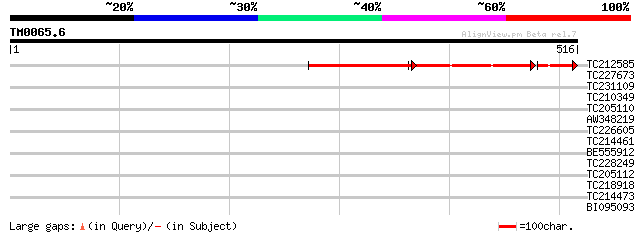
Score E
Sequences producing significant alignments: (bits) Value
TC212585 172 7e-88
TC227673 similar to UP|Q9LKG1 (Q9LKG1) Histone deacetylase, part... 33 0.24
TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, par... 31 1.5
TC210349 30 2.0
TC205110 30 2.0
AW348219 similar to PIR|B85431|B85 trichohyalin like protein [im... 30 2.0
TC226605 similar to UP|Q93YR3 (Q93YR3) HSP associated protein li... 30 2.6
TC214461 homologue to UP|Q39449 (Q39449) Specific tissue protein... 29 4.4
BE555912 29 4.4
TC228249 similar to UP|Q9M9V0 (Q9M9V0) F6A14.10 protein, partial... 29 4.4
TC205112 similar to UP|Q93ZP1 (Q93ZP1) AT5g61020/maf19_20, parti... 29 4.4
TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete 28 7.6
TC214473 similar to UP|Q39449 (Q39449) Specific tissue protein 1... 28 7.6
BI095093 similar to PIR|E85439|E854 thiol-disulfide interchange ... 28 9.9
>TC212585
Length = 1016
Score = 172 bits (437), Expect(3) = 7e-88
Identities = 84/98 (85%), Positives = 93/98 (94%)
Frame = +1
Query: 273 LGNLGPIVICTKVTNSIALLDPFTLRYCFLDADQYWRSSFKSLLTSRQLVEYIVLDVEVV 332
LGN+GP+VICTKVT+SIALLDPFTLR+CF+DADQYWR+SFKSLLTSRQLVEYIVLDVE V
Sbjct: 7 LGNIGPLVICTKVTSSIALLDPFTLRHCFIDADQYWRASFKSLLTSRQLVEYIVLDVETV 186
Query: 333 SSEVTVGGTKYVLADAQVARVSDFGKNDTIFSIKTHLG 370
SSEVT+GGTKY LADAQVARVSDFGKNDTIF+I+ G
Sbjct: 187 SSEVTIGGTKYRLADAQVARVSDFGKNDTIFNIQDSSG 300
Score = 147 bits (372), Expect(3) = 7e-88
Identities = 80/116 (68%), Positives = 90/116 (76%), Gaps = 1/116 (0%)
Frame = +2
Query: 364 SIKTHLGHLLNPGDYALGY-DLYGANNNDIELDKYKGLILPDAILVKKSYEEKRQKKRGK 422
+ KTHLGHLLNPGDYA ++G L KG I P+AIL+KKSYEEK QKKRGK
Sbjct: 281 TFKTHLGHLLNPGDYASWL*SVWG****TWNLTSNKGHI-PEAILIKKSYEEKHQKKRGK 457
Query: 423 ARSWKLKSLNMEVDDKVARADPDKMHSEYEQFLKDLEENPDLRFNISLYRNKEYQP 478
RSWKLKSL MEVDDK R D DKM SEYEQF++DLEENP++RFNISLYRNKEY+P
Sbjct: 458 LRSWKLKSLEMEVDDK-GRVDQDKMVSEYEQFIQDLEENPEMRFNISLYRNKEYKP 622
Score = 43.9 bits (102), Expect(3) = 7e-88
Identities = 22/36 (61%), Positives = 28/36 (77%)
Frame = +1
Query: 481 IASVTDGEELPSVPLEELLADLELGEDEDEEDDMME 516
+ + DG++ SVPLEELLADL+L E EDEED+M E
Sbjct: 613 VQAFLDGDDX-SVPLEELLADLDLSEHEDEEDNMTE 717
>TC227673 similar to UP|Q9LKG1 (Q9LKG1) Histone deacetylase, partial (88%)
Length = 1476
Score = 33.5 bits (75), Expect = 0.24
Identities = 34/107 (31%), Positives = 49/107 (45%), Gaps = 10/107 (9%)
Frame = +3
Query: 420 RGKARSWKLKS---LNMEVDDKVARADPDKMHSEYEQFLKD--LEENPDLRFNISLYRNK 474
R AR W ++ L +E+DDK+ + H YE F D L P N +N
Sbjct: 1032 RNVARCWCYETSVALGIELDDKMPQ------HEYYEYFGPDYTLHVAPSNMEN----KNS 1181
Query: 475 EYQPSEI-ASVTDG----EELPSVPLEELLADLELGEDEDEEDDMME 516
+ EI A + D + PSVP +E D EL E ++++DD E
Sbjct: 1182 RHLLDEIRAKLLDNLSRLQHAPSVPFQERPPDAELLERDEDQDDRDE 1322
>TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, partial (8%)
Length = 1061
Score = 30.8 bits (68), Expect = 1.5
Identities = 28/91 (30%), Positives = 43/91 (46%)
Frame = +3
Query: 229 RSDKQLVSHDTKSNEYNYKHTFSVEISPICREDLICLPPKVALSLGNLGPIVICTKVTNS 288
R L S D KS E N VE+S L+C + + LG+ ++ + V +
Sbjct: 246 RHRSHLYSLDLKSPEQN-----PVELS----HPLMCYSNSIKV-LGSSNGLLCISNVADD 395
Query: 289 IALLDPFTLRYCFLDADQYWRSSFKSLLTSR 319
IAL +PF ++ L AD++ R SL +R
Sbjct: 396 IALWNPFLRKHRILPADRFHRPQ-SSLFAAR 485
>TC210349
Length = 605
Score = 30.4 bits (67), Expect = 2.0
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 3/111 (2%)
Frame = +3
Query: 409 KKSYEEKRQKKRGKARSWKLKSLNMEVDDKVARADPDKMHSEYEQF-LKDLEENPDLRFN 467
+K E +R +R R+ L + N+ +A A S QF L ENP
Sbjct: 21 EKERERERAMRRTLVRNGSLYTRNLLHHSALALAA-----SSRPQFRLFSSNENPPPVPQ 185
Query: 468 ISLYRNKEYQPSEIASVTDGEE--LPSVPLEELLADLELGEDEDEEDDMME 516
I NKE + ++I + G+E LPS+ +E +L G ED +D++ME
Sbjct: 186 IDDVNNKELK-AQIETYFKGDEKVLPSI-MEAILKRKLSGNHEDTDDELME 332
>TC205110
Length = 313
Score = 30.4 bits (67), Expect = 2.0
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Frame = -3
Query: 22 CCKCGIPM-QPNAANMCVKCLRSEVDITEGLLKRLVLVHCPECE 64
CC IP QP + + C+ S I +G +KRL + P+CE
Sbjct: 218 CCSRSIPF*QPFCIHSSICCICSSRFIDQGCIKRLRKCNRPKCE 87
>AW348219 similar to PIR|B85431|B85 trichohyalin like protein [imported] -
Arabidopsis thaliana, partial (1%)
Length = 432
Score = 30.4 bits (67), Expect = 2.0
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +2
Query: 31 PNAANMCVKCLRSEVDITEGLLKRLVLVHCPECE 64
P A N + C+ S I +G +KRL +CP+CE
Sbjct: 236 PXAFNSSLCCICSXSFIHQGCIKRLWKCNCPKCE 337
>TC226605 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like, partial
(37%)
Length = 670
Score = 30.0 bits (66), Expect = 2.6
Identities = 18/74 (24%), Positives = 39/74 (52%), Gaps = 6/74 (8%)
Frame = +2
Query: 449 SEYEQFLKDLEENPDLRFNISLYRNKEYQPSEIASVTDGEELPSVPLE------ELLADL 502
++ + F++ + NP L + SL ++Y S A + + S +E ++ +
Sbjct: 122 NQLKHFIEQCKSNPSLLSDPSLSFFRDYPESLGAKLPESAYSESTGVESDEDIKDVTEEQ 301
Query: 503 ELGEDEDEEDDMME 516
E GE+E+E+D+++E
Sbjct: 302 EKGEEEEEDDEIIE 343
>TC214461 homologue to UP|Q39449 (Q39449) Specific tissue protein 1, partial
(11%)
Length = 849
Score = 29.3 bits (64), Expect = 4.4
Identities = 36/133 (27%), Positives = 58/133 (43%), Gaps = 8/133 (6%)
Frame = +3
Query: 389 NNDIE----LDKYKGLILPDAILVKKSYEEKRQKKRGKARSWKLKSLNMEVDDKVARADP 444
N+D E + KY L +I+ +E + K ++LKS + +D +P
Sbjct: 291 NSDFEPIPSVTKYDDLEFKSSIIKNDDFESRPSVT--KYDDFELKSSVTKNND----FEP 452
Query: 445 DKMHSEYEQF--LKDLEENPDL--RFNISLYRNKEYQPSEIASVTDGEELPSVPLEELLA 500
++Y+ F + EN D R NIS Y + E + +SVT+ ++ S P
Sbjct: 453 RPSATKYDDFELRSSVTENDDFEPRPNISKYDDFELR----SSVTENDDFESRPSVTKYD 620
Query: 501 DLELGEDEDEEDD 513
DLEL + DD
Sbjct: 621 DLELKSSITKNDD 659
>BE555912
Length = 411
Score = 29.3 bits (64), Expect = 4.4
Identities = 22/93 (23%), Positives = 37/93 (39%)
Frame = +1
Query: 369 LGHLLNPGDYALGYDLYGANNNDIELDKYKGLILPDAILVKKSYEEKRQKKRGKARSWKL 428
L +L G A YG + I+LD + L + K Y K ++++
Sbjct: 91 LDQVLELGPIAKSNKKYGIEKSQIDLDTEQYLEQSKTAVRDKPYISKAERRK-------- 246
Query: 429 KSLNMEVDDKVARADPDKMHSEYEQFLKDLEEN 461
++ + K D + H +YE LKD+ N
Sbjct: 247 ----LKKEQKHGEEDLNVEHGKYESKLKDISAN 333
>TC228249 similar to UP|Q9M9V0 (Q9M9V0) F6A14.10 protein, partial (82%)
Length = 1066
Score = 29.3 bits (64), Expect = 4.4
Identities = 22/74 (29%), Positives = 30/74 (39%)
Frame = +3
Query: 443 DPDKMHSEYEQFLKDLEENPDLRFNISLYRNKEYQPSEIASVTDGEELPSVPLEELLADL 502
D D++H E + +KD DL N Y N E ++ S D EE
Sbjct: 657 DVDEIHDEVAELIKD-----DLWPNPLTYFNNELDEEDVDSEADDEEND----------- 788
Query: 503 ELGEDEDEEDDMME 516
ED+ E+DD E
Sbjct: 789 ---EDDSEDDDQEE 821
>TC205112 similar to UP|Q93ZP1 (Q93ZP1) AT5g61020/maf19_20, partial (38%)
Length = 1757
Score = 29.3 bits (64), Expect = 4.4
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Frame = -3
Query: 22 CCKCGIPM-QPNAANMCVKCLRSEVDITEGLLKRLVLVHCPECE 64
C IP QP + + C+ S I +G +KRL +CP+CE
Sbjct: 1536 CWGRSIPFYQPFCIHSSLCCICSSSFIHQGCIKRLWKCNCPKCE 1405
>TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete
Length = 1507
Score = 28.5 bits (62), Expect = 7.6
Identities = 34/122 (27%), Positives = 52/122 (41%), Gaps = 1/122 (0%)
Frame = +1
Query: 393 ELDKYKGLILPDAILVKKSYE-EKRQKKRGKARSWKLKSLNMEVDDKVARADPDKMHSEY 451
E++ Y G L +L KK + K K K S + N +V D D
Sbjct: 700 EIEWYPGKCLTQKVLKKKPKKGSKNAKPITKTESCE-SFFNFFKPPEVPEDDADIDEDLA 876
Query: 452 EQFLKDLEENPDLRFNISLYRNKEYQPSEIASVTDGEELPSVPLEELLADLELGEDEDEE 511
E+ +E++ D+ S R+K P ++ T GE E+L D + EDEDE+
Sbjct: 877 EELQNQMEQDYDIG---STLRDKII-PHAVSWFT-GEAAQGDEFEDLEDDEDEEEDEDED 1041
Query: 512 DD 513
+D
Sbjct: 1042ED 1047
>TC214473 similar to UP|Q39449 (Q39449) Specific tissue protein 1, partial
(19%)
Length = 946
Score = 28.5 bits (62), Expect = 7.6
Identities = 36/139 (25%), Positives = 59/139 (41%), Gaps = 14/139 (10%)
Frame = +3
Query: 389 NNDIE----LDKYKGLILPDAILVKKSYEEKRQKKRG---KARSWKLKSLNMEVDDKVAR 441
N+D E + KY L +I+ +E + + + +S K+ + E V +
Sbjct: 273 NSDFEPIPSVTKYDDLEFKSSIIKNDDFESRPSVTKHDDFELKSSVTKNNDFEPRPSVTK 452
Query: 442 AD---PDKMHSEYEQF--LKDLEENPDL--RFNISLYRNKEYQPSEIASVTDGEELPSVP 494
D P ++Y+ F + EN D R NIS Y + E + +SVT+ ++ S P
Sbjct: 453 YDNFEPRPSVTKYDDFELRSSVTENDDFEPRPNISKYDDFELR----SSVTENDDFESRP 620
Query: 495 LEELLADLELGEDEDEEDD 513
DLEL + DD
Sbjct: 621 SVTKYDDLELKSSITKNDD 677
>BI095093 similar to PIR|E85439|E854 thiol-disulfide interchange like protein
[imported] - Arabidopsis thaliana, partial (34%)
Length = 457
Score = 28.1 bits (61), Expect = 9.9
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +1
Query: 24 KCGIPMQPNAANMCVKCLRSEVDITE 49
KCGIP PN+++ C + S +IT+
Sbjct: 376 KCGIPSMPNSSSSCSHFVLSTFNITK 453
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,380,875
Number of Sequences: 63676
Number of extensions: 338285
Number of successful extensions: 1825
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 1796
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1811
length of query: 516
length of database: 12,639,632
effective HSP length: 101
effective length of query: 415
effective length of database: 6,208,356
effective search space: 2576467740
effective search space used: 2576467740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0065.6


