

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0062.9
(271 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
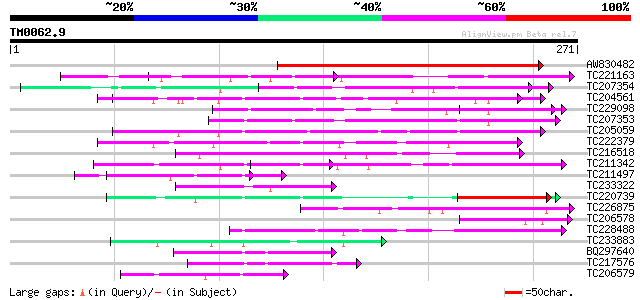
Score E
Sequences producing significant alignments: (bits) Value
AW830482 similar to GP|22022579|gb AT3g05090/T12H1_5 {Arabidopsi... 207 5e-54
TC221163 65 4e-11
TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 62 2e-10
TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 60 8e-10
TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory pr... 58 4e-09
TC207353 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 57 7e-09
TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 55 3e-08
TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, par... 51 6e-07
TC216518 weakly similar to UP|Q9LW87 (Q9LW87) Coatomer protein c... 50 8e-07
TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana geno... 49 2e-06
TC211497 weakly similar to UP|Q9FNZ2 (Q9FNZ2) Zfwd1 protein (Fra... 49 2e-06
TC233322 47 7e-06
TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nu... 47 9e-06
TC226875 similar to UP|Q94JT6 (Q94JT6) At1g78070/F28K19_28, part... 45 3e-05
TC206578 weakly similar to UP|Q8MSF3 (Q8MSF3) GM21396p, partial ... 45 5e-05
TC228488 similar to UP|Q8LF96 (Q8LF96) PRL1 protein, partial (70%) 44 6e-05
TC233883 similar to UP|Q9FG99 (Q9FG99) Similarity to GTP-binding... 44 1e-04
BQ297640 similar to GP|10178169|db contains similarity to WD-con... 44 1e-04
TC217576 44 1e-04
TC206579 weakly similar to UP|TAF5_YEAST (P38129) Transcription ... 43 1e-04
>AW830482 similar to GP|22022579|gb AT3g05090/T12H1_5 {Arabidopsis thaliana},
partial (14%)
Length = 388
Score = 207 bits (526), Expect = 5e-54
Identities = 100/127 (78%), Positives = 116/127 (90%)
Frame = +3
Query: 129 VTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLT 188
VTCL A+EKN+NIVASGGLGGEVF+WD+EAA A +KC D T D++SNGINGSGN LPLT
Sbjct: 6 VTCLTAAEKNNNIVASGGLGGEVFIWDIEAALAPVSKCNDDTVDESSNGINGSGNLLPLT 185
Query: 189 SLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPR 248
SLR+I+SS+++S+HTTQ QGYIPIAAKGHK+SVYALAMNE GT+LVSGGTEKVVRVWD R
Sbjct: 186 SLRTINSSDNMSMHTTQTQGYIPIAAKGHKDSVYALAMNESGTILVSGGTEKVVRVWDTR 365
Query: 249 SGSKTMK 255
SGSKT+K
Sbjct: 366 SGSKTLK 386
>TC221163
Length = 832
Score = 64.7 bits (156), Expect = 4e-11
Identities = 55/218 (25%), Positives = 89/218 (40%), Gaps = 14/218 (6%)
Frame = +3
Query: 67 WALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSD-TTLKTWDAFSTGTCTRTLR-- 123
W D A TF H D V D ++ SD TL+ W+ TG T +R
Sbjct: 18 WMWNTDNAALLNTFIGHGDSVTCGDFTPDGKIICTGSDDATLRIWNP-KTGESTHVVRGH 194
Query: 124 -QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE----------AAHASAAKCIDATDD 172
H++ +TCL + S + SG G V + ++ A+H+ + +C+
Sbjct: 195 PYHTEGLTCLTINS-TSTLALSGSKDGSVHIVNITTGRVVDNNALASHSDSIECVGFAPS 371
Query: 173 DTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTL 232
+ + G L + + + +P H++ V LA G +
Sbjct: 372 GSWAAVGGMDKKLIIWDIEHL----------------LPRGTCEHEDGVTCLAWL-GASY 500
Query: 233 LVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ SG + VR+WD RSG LKGH+D I++L + S
Sbjct: 501 VASGCVDGKVRLWDSRSGECVKTLKGHSDAIQSLSVSS 614
Score = 42.4 bits (98), Expect = 2e-04
Identities = 35/135 (25%), Positives = 57/135 (41%), Gaps = 2/135 (1%)
Frame = +3
Query: 25 VLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGE--DAATCSATFES 82
V++++ H I C+ S + G D +L W + TC
Sbjct: 306 VVDNNALASHSDSIECVGFAPSG-----SWAAVGGMDKKLIIWDIEHLLPRGTCE----- 455
Query: 83 HVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIV 142
H D V +G S + S D ++ WD+ S G C +TL+ HSD + L+ S N N +
Sbjct: 456 HEDGVTCLAWLGASYVASGCVDGKVRLWDSRS-GECVKTLKGHSDAIQSLSVS-SNRNYL 629
Query: 143 ASGGLGGEVFVWDLE 157
S + G +++E
Sbjct: 630 VSASVDGTACAFEVE 674
>TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (57%)
Length = 1124
Score = 62.4 bits (150), Expect = 2e-10
Identities = 60/255 (23%), Positives = 102/255 (39%), Gaps = 10/255 (3%)
Frame = +1
Query: 6 SAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLK 65
S+G + T+ K R L D ++ C + +DG+ L + S D L
Sbjct: 64 SSGGGGTGTQTFKPYRHLKTLTDHENAVSCVKFS---------NDGT-LLASASLDKTLI 213
Query: 66 RWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAFSTGTCTRTLRQ 124
W+ T H + ++D DS + S S D TL+ WDA G C + LR
Sbjct: 214 IWSSA--TLTLCHRLVGHSEGISDLAWSSDSHYICSASDDRTLRIWDATVGGGCIKILRG 387
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H D V C+ + ++S IV SG + VWD++ KC+ G++
Sbjct: 388 HDDAVFCVNFNPQSSYIV-SGSFDETIKVWDVK-----TGKCVHTI----------KGHT 519
Query: 185 LPLTSLR---------SISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVS 235
+P+TS+ S S S + T+ + + +V + G L+++
Sbjct: 520 MPVTSVHYNRDGNLIISASHDGSCKIWDTETGNLLKTLIEDKAPAVSFAKFSPNGKLILA 699
Query: 236 GGTEKVVRVWDPRSG 250
+++W+ SG
Sbjct: 700 ATLNDTLKLWNYGSG 744
Score = 50.8 bits (120), Expect = 6e-07
Identities = 37/142 (26%), Positives = 67/142 (47%), Gaps = 1/142 (0%)
Frame = +1
Query: 120 RTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGIN 179
+TL H + V+C+ S + ++AS L + +W +SA + S GI+
Sbjct: 118 KTLTDHENAVSCVKFSN-DGTLLASASLDKTLIIW------SSATLTLCHRLVGHSEGIS 276
Query: 180 GSGNSLPLTSLRSISSSNSISV-HTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGT 238
S + S S ++ + T G I I +GH ++V+ + N + +VSG
Sbjct: 277 DLAWSSDSHYICSASDDRTLRIWDATVGGGCIKIL-RGHDDAVFCVNFNPQSSYIVSGSF 453
Query: 239 EKVVRVWDPRSGSKTMKLKGHT 260
++ ++VWD ++G +KGHT
Sbjct: 454 DETIKVWDVKTGKCVHTIKGHT 519
Score = 32.0 bits (71), Expect = 0.30
Identities = 15/54 (27%), Positives = 27/54 (49%)
Frame = +1
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
H+ +V + + GTLL S +K + +W + + +L GH++ I L S
Sbjct: 133 HENAVSCVKFSNDGTLLASASLDKTLIIWSSATLTLCHRLVGHSEGISDLAWSS 294
Score = 30.8 bits (68), Expect = 0.68
Identities = 15/55 (27%), Positives = 25/55 (45%), Gaps = 2/55 (3%)
Frame = +2
Query: 105 TTLKTWDAFSTGTCTRTLRQHSDYVTCLAA--SEKNSNIVASGGLGGEVFVWDLE 157
TTL ++ G C + H + V C+ + S N + G V++WDL+
Sbjct: 710 TTLLSYGTMGVGKCLKIYSGHVNRVYCITSTFSVTNGKYIVGGSEDHCVYIWDLQ 874
>TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1419
Score = 60.5 bits (145), Expect = 8e-10
Identities = 69/224 (30%), Positives = 96/224 (42%), Gaps = 21/224 (9%)
Frame = +1
Query: 43 VLKSAVSDGSDYLFTGSRDGRLKRW-ALGEDAATCSATFE---SHVDWVNDAVLVGDSTL 98
VL A S + + + SRD +K W LGE C T + +H DWV+ V STL
Sbjct: 466 VLSVAFSIDNRQIVSASRDRTIKLWNTLGE----CKYTIQDGDAHSDWVS-CVRFSPSTL 630
Query: 99 ----VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVW 154
VS S D T+K W+ + TL H+ YV +A S S + ASGG G + +W
Sbjct: 631 QPTIVSASWDRTVKVWN-LTNCKLRNTLAGHNGYVNTVAVSPDGS-LCASGGKDGVILLW 804
Query: 155 DLEAAHASAAKCIDATDDDTSNGINGSGN----------SLPLTSLRSISSSNSISVHTT 204
DL A +DA + + S N S+ + L S S + V
Sbjct: 805 DL--AEGKRLYSLDA--GSIIHALCFSPNRYWLCAATEQSIKIWDLESKSIVEDLKVDLK 972
Query: 205 QNQGYIPIAAKGHKESV-YALAMN--EGGTLLVSGGTEKVVRVW 245
+K+ V Y ++N G+ L SG T+ VVRVW
Sbjct: 973 TEADATSGGGNANKKKVIYCTSLNWSADGSTLFSGYTDGVVRVW 1104
Score = 55.5 bits (132), Expect = 3e-08
Identities = 60/211 (28%), Positives = 88/211 (41%), Gaps = 4/211 (1%)
Frame = +1
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSAT---FESHVDWVNDAVLVGDSTL-VSCSSDT 105
D SD + T SRD + W L ++ T H +V D VL D +S S D
Sbjct: 220 DNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDG 399
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
L+ WD + GT R H+ V +A S N IV S + +W+ +
Sbjct: 400 ELRLWD-LAAGTSARRFVGHTKDVLSVAFSIDNRQIV-SASRDRTIKLWNTLGECKYTIQ 573
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
DA D S + S ++L T + S S ++ V N A GH V +A
Sbjct: 574 DGDAHSDWVSC-VRFSPSTLQPTIV-SASWDRTVKVWNLTNCKLRNTLA-GHNGYVNTVA 744
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKL 256
++ G+L SGG + V+ +WD G + L
Sbjct: 745 VSPDGSLCASGGKDGVILLWDLAEGKRLYSL 837
Score = 38.5 bits (88), Expect = 0.003
Identities = 29/143 (20%), Positives = 53/143 (36%)
Frame = +1
Query: 121 TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGING 180
T+R H+D VT +A NS+++ + + +W L T +D + G+
Sbjct: 172 TMRAHTDVVTAIATPIDNSDMIVTASRDKSIILWHL-------------TKEDKTYGV-- 306
Query: 181 SGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEK 240
P L GH V + ++ G +SG +
Sbjct: 307 -----PRRRLT------------------------GHSHFVQDVVLSSDGQFALSGSWDG 399
Query: 241 VVRVWDPRSGSKTMKLKGHTDNI 263
+R+WD +G+ + GHT ++
Sbjct: 400 ELRLWDLAAGTSARRFVGHTKDV 468
>TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (59%)
Length = 580
Score = 58.2 bits (139), Expect = 4e-09
Identities = 49/175 (28%), Positives = 74/175 (42%), Gaps = 11/175 (6%)
Frame = +3
Query: 98 LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
+VS S D T++ WD S G C + L HSD VT + + S IV+S G +WD
Sbjct: 99 IVSGSFDETVRVWDVKS-GKCLKVLPAHSDPVTAVDFNRDGSLIVSSS-YDGLCRIWDAS 272
Query: 158 AAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQ--------GY 209
H C+ DD ++ P++ ++ ++ I V T N G
Sbjct: 273 TGH-----CMKTLIDD---------DNPPVSFVKFSPNAKFILVGTLDNTLRLWNYSTGK 410
Query: 210 IPIAAKGHKESVYALAMN---EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTD 261
GH S Y ++ G +V G E + +WD +S KL+GH+D
Sbjct: 411 FLKTYTGHVNSKYCISSTFSTTNGKYIVGGSEENYIYLWDLQSRKIVQKLEGHSD 575
Score = 41.2 bits (95), Expect = 5e-04
Identities = 18/51 (35%), Positives = 28/51 (54%)
Frame = +3
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
GH V+ + N ++VSG ++ VRVWD +SG L H+D + A+
Sbjct: 48 GHTNYVFCVNFNPQSNIIVSGSFDETVRVWDVKSGKCLKVLPAHSDPVTAV 200
>TC207353 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (58%)
Length = 898
Score = 57.4 bits (137), Expect = 7e-09
Identities = 40/171 (23%), Positives = 80/171 (46%), Gaps = 3/171 (1%)
Frame = +3
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
S +VS S D T+K WD TG C T++ H+ VT + + ++ ++ S G +WD
Sbjct: 33 SYIVSGSFDETIKVWDV-KTGKCVHTIKGHTMPVTSVHYN-RDGTLIISASHDGSCKIWD 206
Query: 156 LEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAK 215
+ +D + ++ + S + + + ++++ + + ++ I +
Sbjct: 207 -----TGTGNLLKTLIEDKAPAVSFAKFSPNGKFILAATLNDTLKLWNYGSGKFLKIYS- 368
Query: 216 GHKESVYALAMN---EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
GH VY + G +VSG ++ V +WD ++ + KL+GHTD +
Sbjct: 369 GHVNRVYCITSTFSVTNGRYIVSGSEDRCVYIWDLQAKNMIQKLEGHTDTV 521
Score = 35.0 bits (79), Expect = 0.036
Identities = 13/40 (32%), Positives = 24/40 (59%)
Frame = +3
Query: 221 VYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHT 260
V+ + N + +VSG ++ ++VWD ++G +KGHT
Sbjct: 3 VFCVNFNPQSSYIVSGSFDETIKVWDVKTGKCVHTIKGHT 122
>TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1492
Score = 55.5 bits (132), Expect = 3e-08
Identities = 60/211 (28%), Positives = 88/211 (41%), Gaps = 4/211 (1%)
Frame = +2
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSAT---FESHVDWVNDAVLVGDSTL-VSCSSDT 105
D SD + T SRD + W L ++ T H +V D VL D +S S D
Sbjct: 296 DNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDG 475
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
L+ WD + GT R H+ V +A S N IV S + +W+ +
Sbjct: 476 ELRLWD-LAAGTSARRFVGHTKDVLSVAFSIDNRQIV-SASRDRTIKLWNTLGECKYTIQ 649
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
DA D S + S ++L T + S S ++ V N A GH V +A
Sbjct: 650 DGDAHSDWVSC-VRFSPSTLQPTIV-SASWDRTVKVWNLTNCKLRNTLA-GHNGYVNTVA 820
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKL 256
++ G+L SGG + V+ +WD G + L
Sbjct: 821 VSPDGSLCASGGKDGVILLWDLAEGKRLYSL 913
Score = 38.5 bits (88), Expect = 0.003
Identities = 29/143 (20%), Positives = 53/143 (36%)
Frame = +2
Query: 121 TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGING 180
T+R H+D VT +A NS+++ + + +W L T +D + G+
Sbjct: 248 TMRAHTDVVTAIATPIDNSDMIVTASRDKSIILWHL-------------TKEDKTYGV-- 382
Query: 181 SGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEK 240
P L GH V + ++ G +SG +
Sbjct: 383 -----PRRRLT------------------------GHSHFVQDVVLSSDGQFALSGSWDG 475
Query: 241 VVRVWDPRSGSKTMKLKGHTDNI 263
+R+WD +G+ + GHT ++
Sbjct: 476 ELRLWDLAAGTSARRFVGHTKDV 544
Score = 26.9 bits (58), Expect = 9.8
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 3/38 (7%)
Frame = +3
Query: 211 PIAAKGHKESV-YALAMN--EGGTLLVSGGTEKVVRVW 245
P+ + +K+ V Y ++N G+ L SG T+ VVRVW
Sbjct: 1068 PVVSNPNKKKVIYCTSLNWSSDGSTLFSGYTDGVVRVW 1181
>TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, partial (84%)
Length = 950
Score = 50.8 bits (120), Expect = 6e-07
Identities = 54/216 (25%), Positives = 90/216 (41%), Gaps = 13/216 (6%)
Frame = +1
Query: 43 VLKSAVSDGSDYLFTGSRDGRLKRW-ALGEDAATCSATFESHVDWVNDAVLVGDS---TL 98
VL + S + + +GSRD +K W LG+ T T + H +WV+ + +
Sbjct: 169 VLSVSFSADNRQIVSGSRDRTIKLWNTLGDCKYTI--TDKGHTEWVSCVRFSPNPQNPVI 342
Query: 99 VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDL-E 157
VS D +K W+ + T + H+ Y+ + S S + ASGG G +WDL E
Sbjct: 343 VSAGWDKFVKVWELATCRIQTDHIG-HTGYINTVTISPDGS-LCASGGKDGTTMLWDLNE 516
Query: 158 AAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQG--------Y 209
+ H T D + + S N ++S+SI V + + Y
Sbjct: 517 SKH-----LYSLTAGDEIHALVFSPNRY----WXCAATSSSIIVFDLEKKSKVDELKPEY 669
Query: 210 IPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVW 245
+ + K + +LA + G L +G T+ +R W
Sbjct: 670 VEVGKKAREPECISLAWSADGQTLFAGYTDNKIRAW 777
Score = 33.9 bits (76), Expect = 0.080
Identities = 14/56 (25%), Positives = 27/56 (48%)
Frame = +1
Query: 208 GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
GY + GH V ++ G +S +K +R+W+ +G T + GH +++
Sbjct: 4 GYPKRSLHGHSHIVSDCVISSDGAYALSSSWDKTLRLWELSTGQTTRRFVGHNNDV 171
Score = 30.8 bits (68), Expect = 0.68
Identities = 13/47 (27%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Frame = +1
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSK-TMKLKGHTD 261
GH V +++ + +VSG ++ +++W+ K T+ KGHT+
Sbjct: 154 GHNNDVLSVSFSADNRQIVSGSRDRTIKLWNTLGDCKYTITDKGHTE 294
>TC216518 weakly similar to UP|Q9LW87 (Q9LW87) Coatomer protein complex, beta
prime; beta'-COP protein, partial (29%)
Length = 1189
Score = 50.4 bits (119), Expect = 8e-07
Identities = 39/177 (22%), Positives = 73/177 (41%), Gaps = 10/177 (5%)
Frame = +1
Query: 80 FESHVDWVND-AVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKN 138
F H D++ AV ++S S D LK W+ +C HS YV +A + K+
Sbjct: 334 FAEHKDYIRSLAVHPVLPYVISASDDQVLKLWNWRKGWSCYENFEGHSHYVMQVAFNPKD 513
Query: 139 SNIVASGGLGGEVFVWDLEAA--------HASAAKCIDA-TDDDTSNGINGSGNSLPLTS 189
+ AS L G + +W L+++ H C+D +D ++GS +
Sbjct: 514 PSTFASASLDGTLKIWSLDSSAPNFTLEGHQKGVNCVDYFITNDKQYLLSGSDDYT--AK 687
Query: 190 LRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWD 246
+ S N + +GH+ +V A+ + ++++ + V++WD
Sbjct: 688 VWDYHSRNCVQT------------LEGHENNVTAICAHPELPIIITASEDSTVKIWD 822
>TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MIL23, partial (29%)
Length = 862
Score = 48.9 bits (115), Expect = 2e-06
Identities = 41/168 (24%), Positives = 70/168 (41%), Gaps = 17/168 (10%)
Frame = +1
Query: 116 GTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD---------LEAAHASAAKC 166
GTC TL H VT L ++ S ++ASG +V +WD L AAK
Sbjct: 10 GTCETTLNGHKGAVTTLRYNKAGS-LLASGSRDNDVILWDVVGETGLFRLRGHRDQAAKQ 186
Query: 167 IDAT--------DDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHK 218
+ + DD I+ + + L S ++ VH + ++ GHK
Sbjct: 187 LTVSNVSTMKMNDDALVVAISPDAKYIAVALLDS-----TVKVHFADTFKFF-LSLYGHK 348
Query: 219 ESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
V + ++ G L+V+G +K +++W G + H D++ A+
Sbjct: 349 LPVLCMDISSDGDLIVTGSADKNIKIWGLDFGDCHKSIFAHADSVMAV 492
Score = 43.9 bits (102), Expect = 8e-05
Identities = 37/116 (31%), Positives = 48/116 (40%), Gaps = 1/116 (0%)
Frame = +1
Query: 41 LAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV- 99
L VL +S D + TGS D +K W L D C + +H D V V + V
Sbjct: 349 LPVLCMDISSDGDLIVTGSADKNIKIWGL--DFGDCHKSIFAHADSVMAVQFVPKTHYVF 522
Query: 100 SCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
S D +K WDA TL H + CLA S + IV +G + WD
Sbjct: 523 SVGKDRLVKYWDA-DKFELLLTLEGHHADIWCLAVSNRGDFIV-TGSHDRSIRRWD 684
Score = 39.7 bits (91), Expect = 0.001
Identities = 19/56 (33%), Positives = 29/56 (50%)
Frame = +1
Query: 206 NQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTD 261
++G GHK +V L N+ G+LL SG + V +WD + +L+GH D
Sbjct: 4 DKGTCETTLNGHKGAVTTLRYNKAGSLLASGSRDNDVILWDVVGETGLFRLRGHRD 171
Score = 30.8 bits (68), Expect = 0.68
Identities = 47/222 (21%), Positives = 87/222 (39%), Gaps = 17/222 (7%)
Frame = +1
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRW-ALGE----------DAATCSATFE--SHVDWVN 88
AV + L +GSRD + W +GE D A T S + +
Sbjct: 46 AVTTLRYNKAGSLLASGSRDNDVILWDVVGETGLFRLRGHRDQAAKQLTVSNVSTMKMND 225
Query: 89 DAVLVGDST----LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVAS 144
DA++V S + D+T+K A T +L H V C+ S + +++ +
Sbjct: 226 DALVVAISPDAKYIAVALLDSTVKVHFA-DTFKFFLSLYGHKLPVLCMDISS-DGDLIVT 399
Query: 145 GGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTT 204
G + +W L+ K I A D + +P T + + +
Sbjct: 400 GSADKNIKIWGLD--FGDCHKSIFAHADSVM-----AVQFVPKTHYVFSVGKDRLVKYWD 558
Query: 205 QNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWD 246
++ + + +GH ++ LA++ G +V+G ++ +R WD
Sbjct: 559 ADKFELLLTLEGHHADIWCLAVSNRGDFIVTGSHDRSIRRWD 684
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +1
Query: 34 HCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGED 72
H A I CLAV S+ D++ TGS D ++RW E+
Sbjct: 595 HHADIWCLAV-----SNRGDFIVTGSHDRSIRRWDRTEE 696
>TC211497 weakly similar to UP|Q9FNZ2 (Q9FNZ2) Zfwd1 protein (Fragment),
partial (31%)
Length = 840
Score = 48.9 bits (115), Expect = 2e-06
Identities = 28/86 (32%), Positives = 41/86 (47%)
Frame = +3
Query: 32 TKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAV 91
T H + CL + G L++GS D +K W + D C+ T H D V ++
Sbjct: 165 TGHTKAVVCLTI-------GCKMLYSGSMDQSIKVWDM--DTLQCTMTLNEHTDIVT-SL 314
Query: 92 LVGDSTLVSCSSDTTLKTWDAFSTGT 117
+ D L+SCSSD T+K W G+
Sbjct: 315 ICWDQYLLSCSSDCTIKVWACTEVGS 392
Score = 45.8 bits (107), Expect = 2e-05
Identities = 31/89 (34%), Positives = 44/89 (48%), Gaps = 3/89 (3%)
Frame = +3
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAAT---CSATFESHVDWVNDAVLVGDSTLVSCSS 103
A++ G+D LF G+ DG + W A + A+ H V + +G L S S
Sbjct: 54 AMTVGNDTLFAGAEDGVIFAWRGSSGAKSPFELVASLTGHTKAVV-CLTIGCKMLYSGSM 230
Query: 104 DTTLKTWDAFSTGTCTRTLRQHSDYVTCL 132
D ++K WD T CT TL +H+D VT L
Sbjct: 231 DQSIKVWD-MDTLQCTMTLNEHTDIVTSL 314
Score = 35.0 bits (79), Expect = 0.036
Identities = 18/52 (34%), Positives = 30/52 (57%)
Frame = +3
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALL 267
GH ++V L + G +L SG ++ ++VWD + TM L HTD + +L+
Sbjct: 168 GHTKAVVCLTI--GCKMLYSGSMDQSIKVWDMDTLQCTMTLNEHTDIVTSLI 317
>TC233322
Length = 725
Score = 47.4 bits (111), Expect = 7e-06
Identities = 26/79 (32%), Positives = 38/79 (47%), Gaps = 2/79 (2%)
Frame = +1
Query: 80 FESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLR--QHSDYVTCLAASEK 137
F H V D TL+S SSD T++ W C++ L H DYVTC+ +
Sbjct: 442 FSGHASDVLDLAWSNSDTLLSSSSDKTVRLWKI----GCSQCLSVFHHKDYVTCIQFNPV 609
Query: 138 NSNIVASGGLGGEVFVWDL 156
+ N SG + G+V +W +
Sbjct: 610 DENYFISGSIDGKVRIWGI 666
Score = 27.3 bits (59), Expect = 7.5
Identities = 11/36 (30%), Positives = 19/36 (52%)
Frame = +2
Query: 209 YIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
YI + HK ++ + + G L SGG + V+R+
Sbjct: 167 YIGQEIRAHKGLIWTMKFSPNGQYLASGGEDGVIRI 274
>TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nuclear
ribonucleoprotein Prp4 (U4/U6 snRNP 60 kDa protein) (WD
splicing factor Prp4) (hPrp4), partial (34%)
Length = 1206
Score = 47.0 bits (110), Expect = 9e-06
Identities = 20/45 (44%), Positives = 28/45 (61%)
Frame = +1
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
+GH SVY LA + G+L S G + + RVWD R+G + L+GH
Sbjct: 436 EGHSRSVYGLAFHNDGSLAASCGLDSLARVWDLRTGRSILALEGH 570
Score = 43.1 bits (100), Expect = 1e-04
Identities = 55/218 (25%), Positives = 74/218 (33%), Gaps = 1/218 (0%)
Frame = +1
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWV-NDAVLVGDSTLVSCSSDT 105
A S D+L T S D K W G + TFE H+D + A L + S D
Sbjct: 220 AYSPVHDHLATASADRTAKYWNQG----SLLKTFEGHLDRLARIAFHPSGKYLGTASFDK 387
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
T + WD TG HS V LA S + AS GL VWDL
Sbjct: 388 TWRLWD-IETGDELLLQEGHSRSVYGLAFHNDGS-LAASCGLDSLARVWDLRTG------ 543
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
RSI +A +GH + V ++
Sbjct: 544 -------------------------RSI------------------LALEGHVKPVLGIS 594
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
+ G L +GG + R+WD R + H++ I
Sbjct: 595 FSPNGYHLATGGEDNTCRIWDLRKKKSFYTIPAHSNLI 708
Score = 35.4 bits (80), Expect = 0.028
Identities = 16/61 (26%), Positives = 32/61 (52%)
Frame = +1
Query: 206 NQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRA 265
NQG + +GH + + +A + G L + +K R+WD +G + + +GH+ ++
Sbjct: 283 NQGSLLKTFEGHLDRLARIAFHPSGKYLGTASFDKTWRLWDIETGDELLLQEGHSRSVYG 462
Query: 266 L 266
L
Sbjct: 463 L 465
>TC226875 similar to UP|Q94JT6 (Q94JT6) At1g78070/F28K19_28, partial (84%)
Length = 1542
Score = 45.4 bits (106), Expect = 3e-05
Identities = 47/176 (26%), Positives = 72/176 (40%), Gaps = 45/176 (25%)
Frame = +1
Query: 140 NIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTS--NGINGSGNSLPLTSLRSISSSN 197
N++ +GG GE+ +L+ H C T DD + N ++ N P SLR I+++N
Sbjct: 520 NLMVAGGFQGELICKNLK--HPGVLFCGKITTDDNAITNAVDVYSN--PAGSLRVITANN 687
Query: 198 SI--------------------SVHTTQ----------------------NQGYIPIAAK 215
SV+ T N G I + K
Sbjct: 688 DFQVRVFDAENFASLGCFKYDWSVNNTSVSPDGKLLAVLGDSTECLIADANTGKITGSLK 867
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMK-LKGHTDNIRALLLDS 270
GH + ++ A + G +L +G +K R+WD R+ S++M LKG IRAL S
Sbjct: 868 GHLDYSFSSAWHPDGQILATGNQDKTCRLWDIRNLSQSMAVLKGRMGAIRALRFTS 1035
>TC206578 weakly similar to UP|Q8MSF3 (Q8MSF3) GM21396p, partial (67%)
Length = 1363
Score = 44.7 bits (104), Expect = 5e-05
Identities = 23/58 (39%), Positives = 35/58 (59%), Gaps = 4/58 (6%)
Frame = +1
Query: 216 GHKESVYALAMNEGGTLLVSGGTEKVVRVW--DPRSGSKT--MKLKGHTDNIRALLLD 269
GHK+ V+++A N GT L SG ++ R+W +P K ++LKGHTD++ L D
Sbjct: 199 GHKKKVHSVAWNCIGTKLASGSVDQTARIWHIEPHGHGKVKDIELKGHTDSVDQLCWD 372
Score = 34.3 bits (77), Expect = 0.061
Identities = 21/55 (38%), Positives = 30/55 (54%), Gaps = 3/55 (5%)
Frame = +1
Query: 212 IAAKGHKESVYALAMN-EGGTLLVSGGTEKVVRVWDPRSG--SKTMKLKGHTDNI 263
I KGH +SV L + + L+ + +K VR+WD RSG S+ +L G NI
Sbjct: 325 IELKGHTDSVDQLCWDPKHADLIATASGDKTVRLWDARSGKCSQQAELSGENINI 489
>TC228488 similar to UP|Q8LF96 (Q8LF96) PRL1 protein, partial (70%)
Length = 1303
Score = 44.3 bits (103), Expect = 6e-05
Identities = 40/169 (23%), Positives = 70/169 (40%), Gaps = 8/169 (4%)
Frame = +3
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA------- 158
T+K WD ++G TL H + V LA S +++ + ++G +V WDLE
Sbjct: 579 TIKIWD-LASGVLKLTLTGHIEQVRGLAVSNRHTYMFSAGD-DKQVKCWDLEQNKVIRSY 752
Query: 159 -AHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH 217
H S C+ A + G +S+ R + + +H A GH
Sbjct: 753 HGHLSGVYCL-ALHPTIDVLLTGGRDSV----CRVWDIRSKMQIH----------ALSGH 887
Query: 218 KESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+V ++ +V+G + +++WD R G L H ++RA+
Sbjct: 888 DNTVCSVFTRPTDPQVVTGSHDTTIKMWDLRYGKTMSTLTNHKKSVRAM 1034
Score = 37.7 bits (86), Expect = 0.006
Identities = 19/64 (29%), Positives = 27/64 (41%)
Frame = +3
Query: 208 GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALL 267
G + + GH E V LA++ T + S G +K V+ WD GH + L
Sbjct: 606 GVLKLTLTGHIEQVRGLAVSNRHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLA 785
Query: 268 LDST 271
L T
Sbjct: 786 LHPT 797
Score = 36.6 bits (83), Expect = 0.012
Identities = 32/114 (28%), Positives = 42/114 (36%), Gaps = 10/114 (8%)
Frame = +3
Query: 34 HCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWAL----------GEDAATCSATFESH 83
H +G+ CLA+ + D L TG RD + W + G D CS
Sbjct: 759 HLSGVYCLALHPTI-----DVLLTGGRDSVCRVWDIRSKMQIHALSGHDNTVCSVFTRPT 923
Query: 84 VDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEK 137
D +V+ S DTT+K WD G TL H V +A K
Sbjct: 924 -----------DPQVVTGSHDTTIKMWD-LRYGKTMSTLTNHKKSVRAMAQHPK 1049
Score = 35.0 bits (79), Expect = 0.036
Identities = 48/222 (21%), Positives = 85/222 (37%), Gaps = 6/222 (2%)
Frame = +3
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGD-STLVSCSSDT 105
AVS+ Y+F+ D ++K W L ++ S + H+ V L L++ D+
Sbjct: 657 AVSNRHTYMFSAGDDKQVKCWDLEQNKVIRS--YHGHLSGVYCLALHPTIDVLLTGGRDS 830
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
+ WD S L H D C + V +G + +WDL
Sbjct: 831 VCRVWDIRSK-MQIHALSGH-DNTVCSVFTRPTDPQVVTGSHDTTIKMWDLRYGKTM--- 995
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
+T + + P + +S+++I + K + A+A
Sbjct: 996 ---STLTNHKKSVRAMAQH-PKEQAFASASADNIKKFNLPKGEFCHNMLSQQKTIINAMA 1163
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSG-----SKTMKLKGHTDN 262
+NE G ++V+GG + WD +SG S+T+ G D+
Sbjct: 1164VNEEG-VMVTGGDNGSMWFWDWKSGHNFQQSQTIVQPGSLDS 1286
Score = 27.7 bits (60), Expect = 5.7
Identities = 9/31 (29%), Positives = 18/31 (58%)
Frame = +3
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ +++WD SG + L GH + +R L + +
Sbjct: 576 RTIKIWDLASGVLKLTLTGHIEQVRGLAVSN 668
>TC233883 similar to UP|Q9FG99 (Q9FG99) Similarity to GTP-binding regulatory
protein and WD-repeat protein, partial (21%)
Length = 887
Score = 43.5 bits (101), Expect = 1e-04
Identities = 38/154 (24%), Positives = 58/154 (36%), Gaps = 22/154 (14%)
Frame = +3
Query: 49 SDGSDYLFTGSRDGRLKRWAL--GEDAATCSATFESHVDWVNDAVLVGD-STLVSCSSDT 105
++ + ++TGS D ++K W G+ + T E H VN L D S L S + D
Sbjct: 258 NNNNGIVYTGSADTKIKMWKKLEGDKKHSLIGTLEKHKSAVNALALNSDGSVLYSGACDR 437
Query: 106 TLKTW----DAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA--- 158
++ W D + LR H+ + CL S++V SG V +W
Sbjct: 438 SILVWEGDEDNNNNMVVVGALRGHTKAILCLVV---ESDLVCSGSADNSVRIWRRSVENE 608
Query: 159 ------------AHASAAKCIDATDDDTSNGING 180
+H KC+ D S G G
Sbjct: 609 KKSYYSCLAVLESHRRPVKCLAMAVDSNSGGGGG 710
Score = 38.5 bits (88), Expect = 0.003
Identities = 20/59 (33%), Positives = 33/59 (55%), Gaps = 5/59 (8%)
Frame = +3
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTM-----KLKGHTDNIRALLLDS 270
HK +V ALA+N G++L SG ++ + VW+ + L+GHT I L+++S
Sbjct: 366 HKSAVNALALNSDGSVLYSGACDRSILVWEGDEDNNNNMVVVGALRGHTKAILCLVVES 542
>BQ297640 similar to GP|10178169|db contains similarity to WD-containing
protein~gene_id:K18G13.8 {Arabidopsis thaliana}, partial
(16%)
Length = 416
Score = 43.5 bits (101), Expect = 1e-04
Identities = 24/78 (30%), Positives = 39/78 (49%)
Frame = +3
Query: 79 TFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKN 138
+F+ H+ V D L+S S D T++ W S+ +C + HSDYVTC+ + +
Sbjct: 108 SFKGHLHDVLDLSWSKSQRLLSSSMDKTVRLWH-LSSKSCLKVF-SHSDYVTCIQFNPVD 281
Query: 139 SNIVASGGLGGEVFVWDL 156
SG L +V +W +
Sbjct: 282 DRYFISGSLDAKVRIWSI 335
>TC217576
Length = 1432
Score = 43.5 bits (101), Expect = 1e-04
Identities = 25/83 (30%), Positives = 41/83 (49%)
Frame = +1
Query: 86 WVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASG 145
+++ A ++ L+S S D T++ W C R H++YVTC+ + N N SG
Sbjct: 85 FLHQAPILKPQFLLSSSVDKTVRLWHV-GIDRCLRVF-SHNNYVTCVNFNPVNDNFFISG 258
Query: 146 GLGGEVFVWDLEAAHASAAKCID 168
+ G+V +W E H + ID
Sbjct: 259 SIDGKVRIW--EVVHCRVSDYID 321
>TC206579 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 835
Score = 43.1 bits (100), Expect = 1e-04
Identities = 29/81 (35%), Positives = 38/81 (46%), Gaps = 1/81 (1%)
Frame = +1
Query: 54 YLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVL-VGDSTLVSCSSDTTLKTWDA 112
YL T S D +K W + D T T H WV D V V + L++ SSDTT + W +
Sbjct: 340 YLATASSDHTVKIWNV--DGFTLEKTLIGHQRWVWDCVFSVDGAYLITASSDTTARLW-S 510
Query: 113 FSTGTCTRTLRQHSDYVTCLA 133
STG + + H C A
Sbjct: 511 MSTGEDIKVYQGHHKATICCA 573
Score = 27.7 bits (60), Expect = 5.7
Identities = 41/228 (17%), Positives = 77/228 (32%), Gaps = 10/228 (4%)
Frame = +1
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSC 101
AV V L +G ++G ++ W L ++ +C E + V+ S +V+
Sbjct: 16 AVXXVVVHPDXXELISGGQNGNIRVWDLTANSCSCELVPEVDTAVRSLTVMWDGSLVVAA 195
Query: 102 SSDTTLKTWDAFSTGTCTRT-------LRQHSDYVT-CLAASE--KNSNIVASGGLGGEV 151
++ T W GT T T L+ H Y+ CL + E + +A+ V
Sbjct: 196 NNHGTCYVWRLLR-GTQTMTNFEPLHKLQAHKGYILKCLLSPEFCEPHRYLATASSDHTV 372
Query: 152 FVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIP 211
+W+++ G +L T +
Sbjct: 373 KIWNVD------------------------GFTLEKTLI--------------------- 417
Query: 212 IAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
GH+ V+ + G L++ ++ R+W +G +GH
Sbjct: 418 ----GHQRWVWDCVFSVDGAYLITASSDTTARLWSMSTGEDIKVYQGH 549
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.127 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,875,722
Number of Sequences: 63676
Number of extensions: 162126
Number of successful extensions: 922
Number of sequences better than 10.0: 139
Number of HSP's better than 10.0 without gapping: 749
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 902
length of query: 271
length of database: 12,639,632
effective HSP length: 96
effective length of query: 175
effective length of database: 6,526,736
effective search space: 1142178800
effective search space used: 1142178800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0062.9


