

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
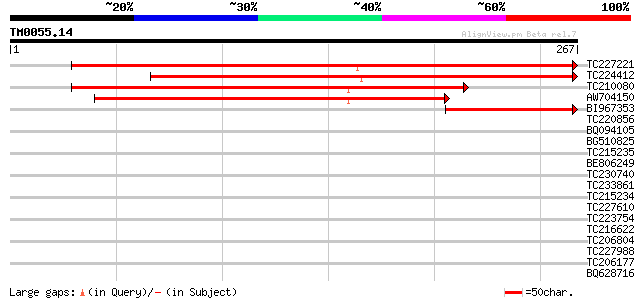
Query= TM0055.14
(267 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC227221 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_1... 357 4e-99
TC224412 282 1e-76
TC210080 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_1... 277 3e-75
AW704150 177 3e-45
BI967353 80 1e-15
TC220856 similar to GB|AAS20952.1|42565013|AY531115 calmodulin b... 39 0.002
BQ094105 homologue to GP|13591696|gb| histone acetyltransferase ... 37 0.009
BG510825 35 0.035
TC215235 similar to UP|O24447 (O24447) Carbamoyl phosphate synth... 35 0.046
BE806249 similar to GP|2462781|gb| carbamoyl phosphate synthetas... 34 0.060
TC230740 similar to UP|Q9FZA1 (Q9FZA1) F3H9.7 protein, partial (... 34 0.060
TC233861 weakly similar to UP|Q41192 (Q41192) NaPRP3, partial (21%) 34 0.060
TC215234 similar to UP|O24447 (O24447) Carbamoyl phosphate synth... 34 0.060
TC227610 similar to UP|Q8L9I8 (Q8L9I8) Anthranilate synthase bet... 34 0.060
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 34 0.078
TC216622 33 0.17
TC206804 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gat... 33 0.17
TC227988 similar to UP|Q9SWE2 (Q9SWE2) Alanine:glyoxylate aminot... 33 0.17
TC206177 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 32 0.23
BQ628716 GP|23093168|g CG4059-PB {Drosophila melanogaster}, part... 32 0.23
>TC227221 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_110, partial
(47%)
Length = 1042
Score = 357 bits (915), Expect = 4e-99
Identities = 168/241 (69%), Positives = 197/241 (81%), Gaps = 3/241 (1%)
Frame = +1
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KRYALLMCGEDSEYL+K HGG +G F ++L E+ E+WDLYK+V GEFP +LPLYDGFV
Sbjct: 61 KRYALLMCGEDSEYLVKMHGGTYGVFMKLLQEQEEKWDLYKLVQGEFPEQHDLPLYDGFV 240
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GSC+DAHAN PWI DL++LV LDS++KKILGICFGHQI+GR LGGKVGRS GWDIG
Sbjct: 241 ISGSCYDAHANDPWILDLIALVIKLDSMHKKILGICFGHQIIGRALGGKVGRSPNGWDIG 420
Query: 150 VTTMKLASSSSIA---LNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHM 206
V + ++SS +A L LPS+LSI+KCHRDE+ LPPKAE+I WSE TG+EMF YGDHM
Sbjct: 421 VKAINVSSSLPLAFSSLKLPSKLSIYKCHRDEILELPPKAEVIAWSEMTGVEMFSYGDHM 600
Query: 207 LGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLKDR 266
GIQGHPEFT D+L FIDR+ RNL+QE FAVD KVKAA QEP+ E KRLCV+FLK R
Sbjct: 601 FGIQGHPEFTYDLLLFFIDRIIQRNLVQEAFAVDAKVKAASQEPDKEILKRLCVDFLKGR 780
Query: 267 L 267
L
Sbjct: 781 L 783
>TC224412
Length = 828
Score = 282 bits (721), Expect = 1e-76
Identities = 134/204 (65%), Positives = 158/204 (76%), Gaps = 3/204 (1%)
Frame = +3
Query: 67 DLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICF 126
D+YKV GEFP D+L LYDGFVITGSC DAH N W+ DLL+L+ LDSIN KILGICF
Sbjct: 6 DVYKVARGEFPDEDDLGLYDGFVITGSCSDAHGNDTWVSDLLNLLRKLDSINTKILGICF 185
Query: 127 GHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALN---LPSELSIFKCHRDEVRNLP 183
GHQILGR LGGKV RS+TGWD+GV T+ L+ S AL+ LPS LSI +CHRDE+R LP
Sbjct: 186 GHQILGRALGGKVTRSTTGWDLGVRTITLSPSLRFALSSLDLPSRLSIIECHRDEIRELP 365
Query: 184 PKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKV 243
KAE+I WS+KTGIEMFRYGDH++GIQGHPE++ DIL H IDRL RN + + F +
Sbjct: 366 AKAEVIAWSDKTGIEMFRYGDHIMGIQGHPEYSKDILLHLIDRLIQRNYIIDAFGKQARE 545
Query: 244 KAALQEPNTEAWKRLCVNFLKDRL 267
+AAL EP+ EAWK+LCV FLK RL
Sbjct: 546 RAALWEPDREAWKKLCVTFLKGRL 617
>TC210080 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_110, partial
(45%)
Length = 693
Score = 277 bits (709), Expect = 3e-75
Identities = 129/190 (67%), Positives = 155/190 (80%), Gaps = 3/190 (1%)
Frame = +2
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+ +LMC EDSEY+ K +GG G F RMLAEEGE WD+YKV G+FP D+L LYDGFV
Sbjct: 119 KRFGVLMCAEDSEYVKKVYGGYSGVFVRMLAEEGETWDVYKVACGDFPDEDDLGLYDGFV 298
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
ITGSC DAH N W++DLL+L+ LDS+NKKILGICFGHQILGR LGGKV RS TGWD+G
Sbjct: 299 ITGSCSDAHGNDTWVNDLLNLLRKLDSMNKKILGICFGHQILGRALGGKVTRSPTGWDLG 478
Query: 150 VTTMKLASS---SSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHM 206
V T+ L+ S + I+L+LPS LSI +CHRDE+R LP KAE+I WS+KTGIEMFRYGDH+
Sbjct: 479 VRTITLSPSLPFALISLDLPSRLSIIECHRDEIRELPAKAEVIAWSDKTGIEMFRYGDHI 658
Query: 207 LGIQGHPEFT 216
+GIQGHPE++
Sbjct: 659 MGIQGHPEYS 688
>AW704150
Length = 511
Score = 177 bits (450), Expect = 3e-45
Identities = 91/170 (53%), Positives = 121/170 (70%), Gaps = 3/170 (1%)
Frame = +2
Query: 41 SEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHAN 100
SEY+ K +GG G F RMLAEEGE WD+YKV G+FP + L LYDGF+IT SC DAH+N
Sbjct: 2 SEYVKKVYGGYSGVFMRMLAEEGETWDMYKVACGDFPNKNNLDLYDGFIITESCSDAHSN 181
Query: 101 HPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASS-- 158
+ W+++LL+L+H L+SINKKIL I F HQIL L V S T ++ + T+ L+ S
Sbjct: 182 NTWVNNLLNLLHKLNSINKKILEIYFNHQILRHTLERNVTHSPTK*NLNIRTITLSPSLP 361
Query: 159 -SSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHML 207
+ I+LNLPS+LSI + H++E+R LP KAEII S KT I++F Y DH++
Sbjct: 362 FTLISLNLPSKLSIIEYHKNEIR*LPTKAEIITLSNKTRIKIFEYRDHII 511
>BI967353
Length = 448
Score = 80.1 bits (196), Expect = 1e-15
Identities = 36/62 (58%), Positives = 46/62 (74%)
Frame = -1
Query: 206 MLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLKD 265
++GIQGHPE++ DILFH IDRL RN + + F + +AAL EP+ EAWK+LCV FLK
Sbjct: 448 IMGIQGHPEYSKDILFHLIDRLIQRNYIIDAFGKQARDRAALWEPDREAWKKLCVTFLKG 269
Query: 266 RL 267
RL
Sbjct: 268 RL 263
>TC220856 similar to GB|AAS20952.1|42565013|AY531115 calmodulin binding
protein 25 {Arabidopsis thaliana;} , partial (11%)
Length = 428
Score = 39.3 bits (90), Expect = 0.002
Identities = 25/67 (37%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Frame = -3
Query: 15 ESGCGGGGGGGGG---KGKRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEE--GERWDLY 69
E+G GGGGGGGGG K ++ +L E LLK G G + L E G+RW +
Sbjct: 294 EAGHGGGGGGGGGLVQKRRKVVVLRQNSAGERLLKSPG*GIGVASEELCEP*IGDRWAKH 115
Query: 70 KVVNGEF 76
+N +
Sbjct: 114 PWLNAGY 94
>BQ094105 homologue to GP|13591696|gb| histone acetyltransferase GCN5
{Arabidopsis thaliana}, partial (12%)
Length = 341
Score = 37.0 bits (84), Expect = 0.009
Identities = 20/47 (42%), Positives = 26/47 (54%)
Frame = -1
Query: 2 KGRTMLLERELEMESGCGGGGGGGGGKGKRYALLMCGEDSEYLLKKH 48
+GR ++L REL + G GGGGGGGG G R L G + + H
Sbjct: 224 EGRRVVLRRELALVDG--GGGGGGGGVGGRRGLGAAGTEGSSAVGLH 90
>BG510825
Length = 440
Score = 35.0 bits (79), Expect = 0.035
Identities = 21/47 (44%), Positives = 25/47 (52%), Gaps = 4/47 (8%)
Frame = -2
Query: 9 ERELEMESGCGGGGGGGGGKGKRYALLMC----GEDSEYLLKKHGGC 51
+R G GGGGGGGGG G LL C G +YLL++ GC
Sbjct: 265 DRVRRFRGGGGGGGGGGGGGGGGGGLLCCLRGTG*HLDYLLER--GC 131
>TC215235 similar to UP|O24447 (O24447) Carbamoyl phosphate synthetase small
subunit , partial (36%)
Length = 826
Score = 34.7 bits (78), Expect = 0.046
Identities = 17/37 (45%), Positives = 23/37 (61%), Gaps = 2/37 (5%)
Frame = +1
Query: 111 VHTLDSINKKI--LGICFGHQILGRVLGGKVGRSSTG 145
V T+ +I K+ GIC GHQ+LG+ LGGK + G
Sbjct: 103 VETVKNILGKVPVFGICMGHQLLGQALGGKTFKMKFG 213
>BE806249 similar to GP|2462781|gb| carbamoyl phosphate synthetase small
subunit {Arabidopsis thaliana}, partial (23%)
Length = 414
Score = 34.3 bits (77), Expect = 0.060
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +2
Query: 121 ILGICFGHQILGRVLGGKVGRSSTG 145
+ GIC GHQ+LG+ LGGK + G
Sbjct: 137 VFGICMGHQLLGQALGGKTFKMKFG 211
>TC230740 similar to UP|Q9FZA1 (Q9FZA1) F3H9.7 protein, partial (28%)
Length = 566
Score = 34.3 bits (77), Expect = 0.060
Identities = 19/43 (44%), Positives = 23/43 (53%)
Frame = -2
Query: 9 ERELEMESGCGGGGGGGGGKGKRYALLMCGEDSEYLLKKHGGC 51
+R G GGGGGGGGG G L G +YLL++ GC
Sbjct: 280 DRVRRFRGGGGGGGGGGGGGGLLCCLRGTG*HLDYLLER--GC 158
>TC233861 weakly similar to UP|Q41192 (Q41192) NaPRP3, partial (21%)
Length = 438
Score = 34.3 bits (77), Expect = 0.060
Identities = 18/43 (41%), Positives = 23/43 (52%)
Frame = -1
Query: 17 GCGGGGGGGGGKGKRYALLMCGEDSEYLLKKHGGCFGFFTRML 59
G GGGGGG GG+G R L + E ++ HG C G +L
Sbjct: 177 GGGGGGGGLGGEGGREGFL*LEAEGEVEVQFHGLCGGELAALL 49
>TC215234 similar to UP|O24447 (O24447) Carbamoyl phosphate synthetase small
subunit , partial (85%)
Length = 1578
Score = 34.3 bits (77), Expect = 0.060
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +2
Query: 121 ILGICFGHQILGRVLGGKVGRSSTG 145
+ GIC GHQ+LG+ LGGK + G
Sbjct: 962 VFGICMGHQLLGQALGGKTFKMKFG 1036
>TC227610 similar to UP|Q8L9I8 (Q8L9I8) Anthranilate synthase beta chain,
partial (59%)
Length = 695
Score = 34.3 bits (77), Expect = 0.060
Identities = 29/105 (27%), Positives = 45/105 (42%), Gaps = 8/105 (7%)
Frame = +1
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTM-----KLASSSSIALNLPSELSIFKCH 175
+ G+C G Q +G GGK+ R+ G G ++M K L+ P +
Sbjct: 118 LFGVCMGLQCIGEAFGGKIVRAPHGVMHGKSSMVYYDEKGEDGVLAGLSNPFLAGRYHSL 297
Query: 176 RDEVRNLP-PKAEIIGWSEKTGIEMFRYG--DHMLGIQGHPEFTI 217
E + P + E+ W+E I R+ H+ G+Q HPE I
Sbjct: 298 VIEKGSFPDDELEVTAWTEDGLIMAARHKKYKHLQGVQFHPESII 432
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 33.9 bits (76), Expect = 0.078
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = +2
Query: 8 LERELEMESGCGGGGGGGGGKGKRYALLMCGEDSEY 43
L R+ ME G GGGGG GG G + CG+ +
Sbjct: 344 LARDCGMEGGRFGGGGGSGGGGGKSTCFNCGKPGHF 451
Score = 27.3 bits (59), Expect = 7.3
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = +2
Query: 8 LERELEMESGCGGGGGGGGG 27
L R+ S GGGGGG GG
Sbjct: 71 LARDCNRSSNSGGGGGGSGG 130
>TC216622
Length = 455
Score = 32.7 bits (73), Expect = 0.17
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = -2
Query: 17 GCGGGGGGGGGKG 29
GCGGGGGGGGG G
Sbjct: 271 GCGGGGGGGGGYG 233
>TC206804 similar to UP|CNG2_ARATH (O65718) Cyclic nucleotide-gated ion
channel 2 (AtCNGC2) (Cyclic nucleotide-and
calmodulin-regulated ion channel 2) (DEFENSE NO DEATH
1), partial (16%)
Length = 701
Score = 32.7 bits (73), Expect = 0.17
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = -2
Query: 19 GGGGGGGGGKGKRYALLMCGEDSEYLLKKHGGC 51
GGGG GG G+G R+ LL+ ++++ G C
Sbjct: 700 GGGGXGGEGRGWRFLLLLNQSSNKHITASSGDC 602
>TC227988 similar to UP|Q9SWE2 (Q9SWE2) Alanine:glyoxylate aminotransferase 2
homolog, partial (67%)
Length = 1096
Score = 32.7 bits (73), Expect = 0.17
Identities = 18/33 (54%), Positives = 22/33 (66%), Gaps = 3/33 (9%)
Frame = -3
Query: 6 MLLER-ELEMESGCGGGGGGGGG--KGKRYALL 35
+++ER EL G GGGGG GGG +GK ALL
Sbjct: 149 LVIERGELRRRDGIGGGGGSGGG*ERGKARALL 51
>TC206177 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (22%)
Length = 406
Score = 32.3 bits (72), Expect = 0.23
Identities = 14/25 (56%), Positives = 16/25 (64%)
Frame = +1
Query: 19 GGGGGGGGGKGKRYALLMCGEDSEY 43
GGGGGGGGG G Y+ CGE +
Sbjct: 70 GGGGGGGGGGGSCYS---CGESGHF 135
>BQ628716 GP|23093168|g CG4059-PB {Drosophila melanogaster}, partial (1%)
Length = 374
Score = 32.3 bits (72), Expect = 0.23
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = -3
Query: 3 GRTMLLERELEMESGCGGGGGGGGGK 28
G T L R +SG GGGGGGGGG+
Sbjct: 78 GLTGLRWRHFRGQSGGGGGGGGGGGR 1
Score = 27.7 bits (60), Expect = 5.6
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = -3
Query: 19 GGGGGGGGGKGK 30
GGGGGGGGG G+
Sbjct: 36 GGGGGGGGGGGR 1
Score = 26.9 bits (58), Expect = 9.6
Identities = 11/15 (73%), Positives = 11/15 (73%)
Frame = -3
Query: 17 GCGGGGGGGGGKGKR 31
G GGGGGGGG G R
Sbjct: 45 GQSGGGGGGGGGGGR 1
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,246,628
Number of Sequences: 63676
Number of extensions: 230843
Number of successful extensions: 6030
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 2484
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4426
length of query: 267
length of database: 12,639,632
effective HSP length: 96
effective length of query: 171
effective length of database: 6,526,736
effective search space: 1116071856
effective search space used: 1116071856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0055.14


