

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.5
(638 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
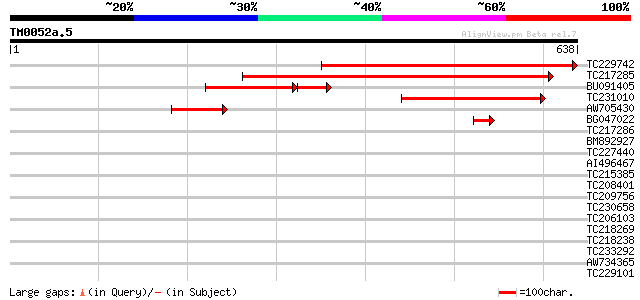
Sequences producing significant alignments: (bits) Value
TC229742 weakly similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol... 535 e-152
TC217285 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol 3-O-gl... 473 e-134
BU091405 183 4e-64
TC231010 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol 3-O-gl... 222 3e-58
AW705430 homologue to GP|24459977|dbj UDP-glucose:sterol 3-O-glu... 105 5e-23
BG047022 52 6e-07
TC217286 weakly similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol... 41 0.001
BM892927 33 0.39
TC227440 UP|Q8LKV1 (Q8LKV1) GAGA-binding protein, complete 33 0.51
AI496467 32 0.66
TC215385 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, par... 32 0.87
TC208401 similar to UP|Q8LFQ5 (Q8LFQ5) RNA binding protein-like,... 32 0.87
TC209756 32 1.1
TC230658 similar to UP|Q9FT43 (Q9FT43) Sterol glucosyltransferas... 31 1.5
TC206103 similar to GB|AAM19988.1|20466133|AY098978 AT3g61870/F2... 30 3.3
TC218269 30 4.3
TC218238 weakly similar to UP|Q8H0F2 (Q8H0F2) Anthocyanin 3'-glu... 30 4.3
TC233292 similar to UP|Q8S9A0 (Q8S9A0) Glucosyltransferase-9, pa... 29 5.6
AW734365 29 5.6
TC229101 disease resistance-like protein 29 5.6
>TC229742 weakly similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol
3-O-glucosyltransferase, partial (34%)
Length = 1133
Score = 535 bits (1377), Expect = e-152
Identities = 243/288 (84%), Positives = 267/288 (92%)
Frame = +2
Query: 351 MTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGPLVDVV 410
+ WWG+RGIINDFRK+KLKL PIAYFSMY GSISHLPTGYMWSPHVVPKPSDWGPLVDVV
Sbjct: 14 LIWWGIRGIINDFRKRKLKLAPIAYFSMYSGSISHLPTGYMWSPHVVPKPSDWGPLVDVV 193
Query: 411 GYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGII 470
GYCFL+L SKY+P+EDFV+WI+KGP PLYFGFGSMPL+DP RTTDVI+EALKDT QRGII
Sbjct: 194 GYCFLNLGSKYQPQEDFVRWIQKGPKPLYFGFGSMPLDDPKRTTDVIVEALKDTGQRGII 373
Query: 471 DRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFF 530
DRGWGNLG+LAEVPDNVF+LEECPHDWLFPQCSA+VHHGGAGTTATGLKAGCPTTIVPFF
Sbjct: 374 DRGWGNLGNLAEVPDNVFVLEECPHDWLFPQCSALVHHGGAGTTATGLKAGCPTTIVPFF 553
Query: 531 GDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIAKLVENEDGVA 590
GDQFFWGDRI++K LGPAPIPISQL++ENLSN+IKFMLQPEVKSR M +AKL+ENEDGV
Sbjct: 554 GDQFFWGDRINQKGLGPAPIPISQLSLENLSNSIKFMLQPEVKSRAMEVAKLIENEDGVT 733
Query: 591 AAVDAFHRHLPDELPLPTPSPQEEDQINPLQWFFLQIGKWCCAPCGGV 638
AAVD+FHRHLP ELPLP PS +D NPLQWFFLQ+ K+C PCGGV
Sbjct: 734 AAVDSFHRHLPPELPLPIPSAVVDDHANPLQWFFLQLAKFCTLPCGGV 877
>TC217285 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol
3-O-glucosyltransferase, partial (59%)
Length = 1302
Score = 473 bits (1218), Expect = e-134
Identities = 211/350 (60%), Positives = 272/350 (77%)
Frame = +2
Query: 263 QLKQLKDIIDSLLPACTAPDLETGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPW 322
Q Q+K+II+SLLPAC PD+++GVPF+A AIIANPPAYGH HVAEAL +P+HIFFTMPW
Sbjct: 2 QRNQMKEIINSLLPACKEPDIDSGVPFKADAIIANPPAYGHTHVAEALKIPIHIFFTMPW 181
Query: 323 TPTYDFPHPLARVPQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGS 382
TPT +FPHPL+RV Q AGY +SY IVD + W G+R +IND RKKKLKL P+ Y S +GS
Sbjct: 182 TPTTEFPHPLSRVKQQAGYRLSYQIVDSLIWLGIRDMINDLRKKKLKLRPVTYLSGSQGS 361
Query: 383 ISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPPLYFGF 442
+ +P Y+WSPH+VPKP DWGP +DVVG+CFL LA Y+P E V+W+++G P+Y GF
Sbjct: 362 ETDVPHAYIWSPHLVPKPKDWGPKIDVVGFCFLDLALNYEPPESLVKWLEEGDKPIYIGF 541
Query: 443 GSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQC 502
GS+P+++P + T +I++AL+ T QRGII++GWG LG+LAE D+++LL+ CPHDWLF +C
Sbjct: 542 GSLPVQEPKKMTQIIVDALEITGQRGIINKGWGGLGNLAEPKDSIYLLDNCPHDWLFLRC 721
Query: 503 SAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSN 562
AVVHHGGAGTTA GLKA CPTTIVPFFGDQ FWG+R+H + +GP PIP+ + ++ L +
Sbjct: 722 KAVVHHGGAGTTAAGLKAACPTTIVPFFGDQPFWGERVHARGVGPPPIPVDEFSLPKLVD 901
Query: 563 AIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSPQ 612
AIK ML P+VK R + +AK +ENEDGV AV AF + LP + PQ
Sbjct: 902 AIKLMLDPKVKERAIELAKAMENEDGVTGAVKAFFKQLPQKKSESDADPQ 1051
>BU091405
Length = 431
Score = 183 bits (464), Expect(2) = 4e-64
Identities = 87/103 (84%), Positives = 94/103 (90%)
Frame = +1
Query: 221 FNTFVKSAGIDFYPLGGDPRILAGYMARNKGLIPSGPGEVSIQLKQLKDIIDSLLPACTA 280
F+TFVKSAG++FYPLGGDPR LA YMARNKG+IPSGP E+SIQ KQLK IIDSLLPAC +
Sbjct: 1 FDTFVKSAGVNFYPLGGDPRALAEYMARNKGIIPSGPTEISIQRKQLKAIIDSLLPACIS 180
Query: 281 PDLETGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPWT 323
PDLETGVPFRAQAII+NP A GH HVAEALGVPLHIFFTMPWT
Sbjct: 181 PDLETGVPFRAQAIISNPTACGHTHVAEALGVPLHIFFTMPWT 309
Score = 80.9 bits (198), Expect(2) = 4e-64
Identities = 33/39 (84%), Positives = 37/39 (94%)
Frame = +2
Query: 324 PTYDFPHPLARVPQSAGYWMSYIIVDLMTWWGMRGIIND 362
PTY+F HPLARVPQSAGYW+SYIIVDL+ WWG+RGIIND
Sbjct: 314 PTYEFSHPLARVPQSAGYWLSYIIVDLLIWWGIRGIIND 430
>TC231010 similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol
3-O-glucosyltransferase, partial (28%)
Length = 893
Score = 222 bits (566), Expect = 3e-58
Identities = 98/163 (60%), Positives = 130/163 (79%)
Frame = +1
Query: 441 GFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFP 500
GFGS+PL+ P + T +I++AL++T QRGII++GWG LGSLAE +V+LL+ CPHDWLFP
Sbjct: 4 GFGSLPLQQPEKITQIIIQALEETGQRGIINKGWGGLGSLAEQNKSVYLLDNCPHDWLFP 183
Query: 501 QCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENL 560
+C+AVVHHGGAGTTA GL+A CPTTIVPFFGDQ FWGDR+ + +GPAPIP+ + + + L
Sbjct: 184 RCTAVVHHGGAGTTAAGLRAECPTTIVPFFGDQPFWGDRVRARGVGPAPIPVDEFSFDRL 363
Query: 561 SNAIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHLPDE 603
+AI ML+PEVK R + +A ++NE+GV AV AF++H P E
Sbjct: 364 VDAIHLMLKPEVKKRAVELANAMKNENGVLGAVKAFYKHYPAE 492
>AW705430 homologue to GP|24459977|dbj UDP-glucose:sterol
3-O-glucosyltransferase {Panax ginseng}, partial (10%)
Length = 433
Score = 105 bits (263), Expect = 5e-23
Identities = 46/63 (73%), Positives = 57/63 (90%)
Frame = +2
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRIL 242
L I +L+VGTRGDVQPF+AI KR+Q+YGHRVRLATHSNF FV +AG++FYPLGGDP++L
Sbjct: 5 LNIVMLIVGTRGDVQPFIAIGKRMQDYGHRVRLATHSNFKEFVLTAGLEFYPLGGDPKVL 184
Query: 243 AGY 245
AG+
Sbjct: 185 AGW 193
>BG047022
Length = 405
Score = 52.4 bits (124), Expect = 6e-07
Identities = 21/24 (87%), Positives = 23/24 (95%)
Frame = +1
Query: 522 CPTTIVPFFGDQFFWGDRIHEKEL 545
CPTTIVPFFGDQFFW DRI++KEL
Sbjct: 334 CPTTIVPFFGDQFFWRDRIYDKEL 405
>TC217286 weakly similar to UP|Q8H9B4 (Q8H9B4) UDP-glucose:sterol
3-O-glucosyltransferase, partial (7%)
Length = 436
Score = 41.2 bits (95), Expect = 0.001
Identities = 19/38 (50%), Positives = 24/38 (63%)
Frame = +2
Query: 575 RVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSPQ 612
R + +AK +ENEDGV AV AF + LP + P P PQ
Sbjct: 5 RAIELAKAMENEDGVTGAVKAFFKQLPQKKPEPDADPQ 118
>BM892927
Length = 428
Score = 33.1 bits (74), Expect = 0.39
Identities = 22/53 (41%), Positives = 30/53 (56%), Gaps = 2/53 (3%)
Frame = -2
Query: 225 VKSAGIDFYPLGGDPRILAGYMARNKGLIPSGPGEVSIQLKQLKDI--IDSLL 275
V + GIDF+ LGGD R G R +G I PG ++ L+ L D +DSL+
Sbjct: 190 VDADGIDFFELGGDERFGVGVQNRGEG-IKDEPGVLNGDLQ*LGDCRRVDSLV 35
>TC227440 UP|Q8LKV1 (Q8LKV1) GAGA-binding protein, complete
Length = 1305
Score = 32.7 bits (73), Expect = 0.51
Identities = 20/65 (30%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Frame = -1
Query: 437 PLYFGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRG-WGNLGSLAEVPDNVFLLEE-CP 494
P+Y+ F S T L + ++ G +GNLG+L+ +P ++FLL++ C
Sbjct: 780 PIYYNFSSFLRNPCTLNCRSTLILWRTRPPLWLLSLGRFGNLGTLSFLPSSIFLLDDRCF 601
Query: 495 HDWLF 499
WLF
Sbjct: 600 FHWLF 586
>AI496467
Length = 400
Score = 32.3 bits (72), Expect = 0.66
Identities = 12/13 (92%), Positives = 13/13 (99%)
Frame = +3
Query: 433 KGPPPLYFGFGSM 445
+GPPPLYFGFGSM
Sbjct: 3 QGPPPLYFGFGSM 41
>TC215385 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, partial (66%)
Length = 1271
Score = 32.0 bits (71), Expect = 0.87
Identities = 37/112 (33%), Positives = 51/112 (45%), Gaps = 2/112 (1%)
Frame = -2
Query: 166 PTVSGSLSSALKRSVPRLQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFV 225
PTV+ + LKR R Q+AI G R DV + R +E N +
Sbjct: 529 PTVT---AHKLKRRESR-QVAIEGGGGRPDVGDGGGVLVRSREEPLVGLNVEEQNQIMDI 362
Query: 226 KSAGIDFYPLGGDPRILAGYMARNKGLIPSGPGEVSIQLKQLKDI--IDSLL 275
+ GIDF+ LGGD R G R + I PG ++ L+ L D +DSL+
Sbjct: 361 DADGIDFFELGGDERFGVGVQNRGES-IEYEPGVLNGDLQ*LGDCRRVDSLI 209
>TC208401 similar to UP|Q8LFQ5 (Q8LFQ5) RNA binding protein-like, partial
(36%)
Length = 1073
Score = 32.0 bits (71), Expect = 0.87
Identities = 15/42 (35%), Positives = 20/42 (46%), Gaps = 2/42 (4%)
Frame = +2
Query: 299 PAYGHAHVAEALGVPLH--IFFTMPWTPTYDFPHPLARVPQS 338
P Y + H E +GVP H + T PW P P + VP +
Sbjct: 719 PLYQYHHPTETIGVPAHHNFYQTAPWVPMTIMSKPSSIVPHT 844
>TC209756
Length = 1356
Score = 31.6 bits (70), Expect = 1.1
Identities = 42/211 (19%), Positives = 81/211 (37%), Gaps = 19/211 (9%)
Frame = +1
Query: 343 MSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPHVVPKPSD 402
M I + W G I D +++K + + + + P + SP +P
Sbjct: 76 MPLIDTENFPWRGFNKIFFDHLVQEMKTLELGEWWLCNTTYDLEPGAFSVSPKFLPI--- 246
Query: 403 WGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPP--LYFGFGSMPLEDPTRTTDVILEA 460
GPL++ S ++ ++ ++W+ + PP +Y FGS+ + DP + ++ L
Sbjct: 247 -GPLMESDN----SKSAFWEEDTTCLEWLDQQPPQSVIYVSFGSLAVMDPNQFKELALAL 411
Query: 461 LKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHD----------WL-------FPQCS 503
++D+ + + + DN + HD W P +
Sbjct: 412 -------DLLDKPF--IWVVRPCNDNKENVNAYAHDFHGSKGKIVGWAPQKKILNHPALA 564
Query: 504 AVVHHGGAGTTATGLKAGCPTTIVPFFGDQF 534
+ + H G +T G+ AG P P DQ+
Sbjct: 565 SFISHCGWNSTLEGICAGVPFLCWPCATDQY 657
>TC230658 similar to UP|Q9FT43 (Q9FT43) Sterol glucosyltransferase-like
protein, partial (6%)
Length = 883
Score = 31.2 bits (69), Expect = 1.5
Identities = 13/29 (44%), Positives = 20/29 (68%)
Frame = +3
Query: 526 IVPFFGDQFFWGDRIHEKELGPAPIPISQ 554
+ PF DQF+W +R+H LG +P P+S+
Sbjct: 732 VCPFILDQFYWAERMH--WLGVSPEPLSR 812
>TC206103 similar to GB|AAM19988.1|20466133|AY098978 AT3g61870/F21F14_40
{Arabidopsis thaliana;} , partial (53%)
Length = 726
Score = 30.0 bits (66), Expect = 3.3
Identities = 16/56 (28%), Positives = 31/56 (54%), Gaps = 1/56 (1%)
Frame = +1
Query: 24 KGLWQMDGGDDKLVRVDIKSQDGKFDLASSKEV-EDWREKLGHKSSSLVTSSPRKG 78
+G W +GG+ L R+ + + K +L S ++ W++ H+ +S ++S RKG
Sbjct: 379 QGTWGGNGGELGLSRMILMTSRNKSNLPQSPQMGNQWKKHHHHRLNSKYSNSQRKG 546
>TC218269
Length = 532
Score = 29.6 bits (65), Expect = 4.3
Identities = 27/86 (31%), Positives = 36/86 (41%), Gaps = 11/86 (12%)
Frame = -1
Query: 263 QLKQLKD--IIDS---LLPA-CTAPDLETGVP-----FRAQAIIANPPAYGHAHVAEALG 311
Q KQ+KD I S LLP C P T P + A + I P +G V +
Sbjct: 430 QNKQIKDSRIFTSTMILLPIPCVFPAYITNSPPSQFLYSASSFIRQVPLFGKIIVITGIK 251
Query: 312 VPLHIFFTMPWTPTYDFPHPLARVPQ 337
+ H+ W PT+ PHP + Q
Sbjct: 250 MLFHL-----WLPTWRAPHPSLAIVQ 188
>TC218238 weakly similar to UP|Q8H0F2 (Q8H0F2) Anthocyanin
3'-glucosyltransferase, partial (26%)
Length = 936
Score = 29.6 bits (65), Expect = 4.3
Identities = 35/115 (30%), Positives = 47/115 (40%), Gaps = 17/115 (14%)
Frame = +3
Query: 438 LYFGFGSMPLEDPTRTTDVILEALKDTE--------QRGIIDRGWGNLGSLAEVPDNVFL 489
LY FGSM + PT I AL+D++ ++G + G GN L E V
Sbjct: 27 LYVSFGSMN-KFPTPQLVEIAHALEDSDHDFIWVVRKKGESEDGEGN-DFLQEFDKRVKA 200
Query: 490 LEECPHDWLF-PQC--------SAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFF 535
+ W + PQ AVV H G T + AG P P F +QF+
Sbjct: 201 SNKGYLIWGWAPQLLILEHHAIGAVVTHCGWNTIIESVNAGLPMATWPLFAEQFY 365
>TC233292 similar to UP|Q8S9A0 (Q8S9A0) Glucosyltransferase-9, partial (46%)
Length = 993
Score = 29.3 bits (64), Expect = 5.6
Identities = 22/71 (30%), Positives = 28/71 (38%)
Frame = +1
Query: 464 TEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCP 523
T+ RG+I RGW P + L +L H G +T G+ AG P
Sbjct: 451 TKGRGLIIRGWA--------PQVLILSHHAIGGFLT--------HCGWNSTLEGIGAGLP 582
Query: 524 TTIVPFFGDQF 534
P F DQF
Sbjct: 583 MITWPLFADQF 615
>AW734365
Length = 409
Score = 29.3 bits (64), Expect = 5.6
Identities = 18/43 (41%), Positives = 23/43 (52%), Gaps = 1/43 (2%)
Frame = +3
Query: 326 YDFPHPLARVPQSAGYWMSYIIVDLMTW-WGMRGIINDFRKKK 367
Y HPLA+ W IV L+TW W +R I N FR++K
Sbjct: 273 YYSTHPLAQYQ-----WWRSRIVTLLTWAWSVRLIHNYFRREK 386
>TC229101 disease resistance-like protein
Length = 2755
Score = 29.3 bits (64), Expect = 5.6
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Frame = +2
Query: 294 IIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYD--FPHPLARVPQSAGYW 342
I+ NPP+Y H+ + LGV I T ++D PH QS G W
Sbjct: 2603 ILINPPSYKEKHILDDLGVSKTICVTQKELASFDSLLPH-----*QSGGDW 2740
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,142,306
Number of Sequences: 63676
Number of extensions: 522943
Number of successful extensions: 2556
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 2543
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2555
length of query: 638
length of database: 12,639,632
effective HSP length: 103
effective length of query: 535
effective length of database: 6,081,004
effective search space: 3253337140
effective search space used: 3253337140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0052a.5


