

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.1
(387 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
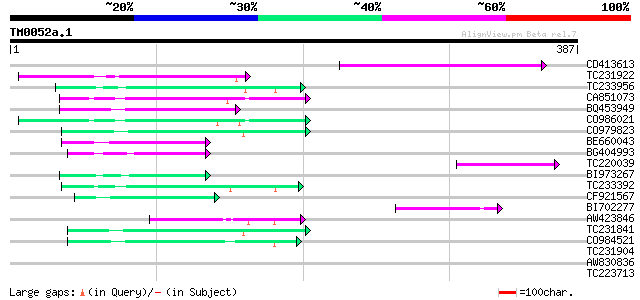
Score E
Sequences producing significant alignments: (bits) Value
CD413613 65 4e-11
TC231922 62 4e-10
TC233956 57 2e-08
CA851073 55 7e-08
BQ453949 51 8e-07
CO986021 50 2e-06
CO979823 49 5e-06
BE660043 49 5e-06
BG404993 48 7e-06
TC220039 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase h... 46 2e-05
BI973267 46 3e-05
TC233392 45 6e-05
CF921567 45 7e-05
BI702277 44 9e-05
AW423846 44 9e-05
TC231841 44 1e-04
CO984521 44 2e-04
TC231904 similar to GB|AAL25568.1|16648753|AY058152 AT5g16150/T2... 41 0.001
AW830836 40 0.001
TC223713 39 0.004
>CD413613
Length = 623
Score = 65.5 bits (158), Expect = 4e-11
Identities = 46/142 (32%), Positives = 68/142 (47%), Gaps = 1/142 (0%)
Frame = -1
Query: 226 SRPARWRRPQPGTIKCNFDAS-FRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAE 284
S +W RP IKCN DAS FR GLG RNS+G +A + + P+ AE
Sbjct: 497 SSTPKWSRPSLEFIKCNLDASLFRNYLLHGLGACNRNSQGTFVADLSCWFKGLLPPMEAE 318
Query: 285 AMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVS 344
A+ + ++ F ++LE DC L + T ++ + +++ C S+FE S
Sbjct: 317 DRALSQVITWLLNNNFTKVILEIDCQQLVDTINTSYFQKNEVGDILKLCVSKCSSFENCS 138
Query: 345 LCFVRRTGNAVADFLARNANTY 366
+ FV R N V LAR + Y
Sbjct: 137 I*FVSRQTNQVIHSLARTSRFY 72
>TC231922
Length = 803
Score = 62.0 bits (149), Expect = 4e-10
Identities = 51/169 (30%), Positives = 74/169 (43%), Gaps = 11/169 (6%)
Frame = -3
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSL-NGGRDILFWPESSNGHYTSKAGYVFLRKLASIEA 65
WRR L ++ ++ + + +S+ N D +FW S+G Y++K+ Y L
Sbjct: 534 WRRRLFDYEYAMAVDFMNEISGISIQNQAHDSMFWKADSSGVYSTKSAYRLLM------- 376
Query: 66 PSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDV-DPGCCLCS 124
PS S A S R ++ W PR WR LP R L RR + + D C LC
Sbjct: 375 PSISPAP--SRRIFQILWHLKIPPRAAVFSWRLFLDRLPTRGNLSRRSIPIQDIMCPLCG 202
Query: 125 GENETAEHLFMLCPAAKQV*YASSLSLRV---------DSFIAFADFWA 164
++E A HLF C K + + S +RV FI F D +A
Sbjct: 201 CQHEEAGHLFFHCKMTKGLWWESMRWIRVIGALPADPASHFIQFCDGFA 55
>TC233956
Length = 757
Score = 56.6 bits (135), Expect = 2e-08
Identities = 43/179 (24%), Positives = 70/179 (39%), Gaps = 8/179 (4%)
Frame = +2
Query: 32 NGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRC 91
N D W S G ++K+ Y ++ E +GF K W P+
Sbjct: 47 NNFNDTWVWRAESTGIISTKSAYQVIKS----ELDDEGQHLGF-----KKLWEIKVPPKA 199
Query: 92 KETCWRAISGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLCPAAKQV*YASSLS 150
WR + LP +D L +R + V+ C C ++ETA HLF C + +
Sbjct: 200 LSFVWRLLWDRLPTKDNLIKRQIQVEDDLCPFCHSQSETASHLFFTCGKIMPLWWEFLSW 379
Query: 151 LRVDSFIAF--ADFWAWVVEQGDSEFISTVQT-----IIYAIWEARNQVQFQQRTFSVT 202
++ D D + S+ +T +T + +IW RN + FQ +T +T
Sbjct: 380 VKEDKVFHCRPMDNFLQHYSSAASKVSNTRRTMWWIAVTNSIWRLRNDIIFQNQTVDIT 556
>CA851073
Length = 633
Score = 54.7 bits (130), Expect = 7e-08
Identities = 48/181 (26%), Positives = 74/181 (40%), Gaps = 10/181 (5%)
Frame = +1
Query: 35 RDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKET 94
+D + W G Y++K+ Y R L + S + +K W PR
Sbjct: 25 QDTMVWKADPCGVYSTKSAY---RILMTCNRHVSEANI------FKTIWKLKIPPRAAVF 177
Query: 95 CWRAISGYLPLRDRLFRRHVDVDPG-CCLCSGENETAEHLFMLCPAAKQV*YAS------ 147
WR I LP R L RR+V + C LC E E A+HLF C + + + S
Sbjct: 178 SWRLIKDRLPTRHNLLRRNVSIQENECPLCGYEQEEADHLFFNCKMTRGLWWESMRWNQI 357
Query: 148 --SLSLRVDS-FIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGV 204
LS+ S F+ F D + + + + + +IW+ RN + FQ F + V
Sbjct: 358 VGPLSVSPASHFVQFCD--GFGAGRNHTRWCGWWIALTSSIWKHRNLLIFQGNQFEPSKV 531
Query: 205 L 205
+
Sbjct: 532 M 534
>BQ453949
Length = 450
Score = 51.2 bits (121), Expect = 8e-07
Identities = 35/124 (28%), Positives = 53/124 (42%), Gaps = 1/124 (0%)
Frame = +3
Query: 35 RDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKET 94
RD+L W +S+G Y++K+ Y FL+ S S+ +K W PR
Sbjct: 27 RDLLLWGSNSDGSYSTKSAYNFLKNEDSQTIEDSA---------FKNIWNLKLPPRACAF 179
Query: 95 CWRAISGYLPLRDRLFRRHVDV-DPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRV 153
R +P + L RRHV + C LC E ET H+ C + + + + L
Sbjct: 180 S*RIF*NRIPTKVNLRRRHVKLPSSNCPLCDEEEETVGHVMYSCIKTRHLWWETELGQ*S 359
Query: 154 DSFI 157
SF+
Sbjct: 360 GSFL 371
>CO986021
Length = 900
Score = 49.7 bits (117), Expect = 2e-06
Identities = 51/210 (24%), Positives = 84/210 (39%), Gaps = 11/210 (5%)
Frame = -1
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGG-RDILFWPESSNGHYTSKAGYVFLRKLASIEA 65
WRR+L + + A + +S++ +D + W G Y++K+ Y R L
Sbjct: 873 WRRNLFDHENEQAIAFMDDISAISIHQQLQDSMLWKADPTGIYSTKSAY---RLLLPNNR 703
Query: 66 PSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDV-DPGCCLCS 124
P ++ +K W PR + WR L +R L RRHV + D C LC
Sbjct: 702 PGQHSSN------FKILWKLKIPPRAELFSWRLFRDRLSIRANLLRRHVALQDIMCPLCG 541
Query: 125 GENETAEHLFMLCPAA-----KQV*YASSLSLRVDS----FIAFADFWAWVVEQGDSEFI 175
E A HLF C + + + +L +D FI F+D + ++ +
Sbjct: 540 NHQEEAGHLFFHCRMTIGLWWESMNWTRTLGAFLDDPAAHFIQFSD--GFGAQRNHNRRC 367
Query: 176 STVQTIIYAIWEARNQVQFQQRTFSVTGVL 205
+ IW+ RN + F+ F V+
Sbjct: 366 LWWIALTSTIWQHRNSLLFKGTPFQPPKVM 277
>CO979823
Length = 853
Score = 48.5 bits (114), Expect = 5e-06
Identities = 45/178 (25%), Positives = 69/178 (38%), Gaps = 8/178 (4%)
Frame = -2
Query: 36 DILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETC 95
D W GHY++K+GY L + E + A W +
Sbjct: 702 DKWLWKPEPGGHYSTKSGYHVLWGELTEEIQDADFA---------EIWKLKIPTKAAVFA 550
Query: 96 WRAISGYLPLRDRLFRRHVDV-DPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVD 154
WR + LP + L RR V V D C LC+ E A HLF C + + S + +
Sbjct: 549 WRLVRDRLPTKSNLRRRQVMVQDMVCPLCNNIEEGAAHLFFNCTKTLPLWWESMSWVNLK 370
Query: 155 SFIA------FADFWAWVVEQGDSE-FISTVQTIIYAIWEARNQVQFQQRTFSVTGVL 205
+ + F + + + S+ + + + IW+ RN+V FQ TF VL
Sbjct: 369 TAMPQTPRQHFLQYGTDIADGLKSKRWKCWWIALTWTIWQHRNKVVFQNATFHGNKVL 196
>BE660043
Length = 714
Score = 48.5 bits (114), Expect = 5e-06
Identities = 28/103 (27%), Positives = 44/103 (42%), Gaps = 1/103 (0%)
Frame = -2
Query: 36 DILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETC 95
D W SNG+Y+S+ Y+ L+ ++ G +K W +
Sbjct: 626 DSWVWKPESNGYYSSRNAYILLQ---------GNSEEGNMDDIFKDLWKLKIPAKASIFA 474
Query: 96 WRAISGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLC 137
R I LP + + +R +D++ C CS + ETA HLF C
Sbjct: 473 GRLIRDRLPTKSNMRKRQIDINDSLCPFCSIKEETASHLFFDC 345
>BG404993
Length = 412
Score = 48.1 bits (113), Expect = 7e-06
Identities = 30/99 (30%), Positives = 42/99 (42%), Gaps = 1/99 (1%)
Frame = -3
Query: 40 WPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAI 99
W +G+Y++K Y L+ EA G + WK P+ WR I
Sbjct: 320 WKADPSGNYSTKTAYELLQ-----EAIRGDNEDGIFVQLWK----LKIPPKAPVFTWRLI 168
Query: 100 SGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLC 137
+ LP + L RR V++ C LC+ E A HLF C
Sbjct: 167 NDRLPTKVNLRRRQVEISDSLCPLCNNSEEDAAHLFFNC 51
>TC220039 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase homolog COK-4,
partial (27%)
Length = 1636
Score = 46.2 bits (108), Expect = 2e-05
Identities = 21/71 (29%), Positives = 42/71 (58%), Gaps = 1/71 (1%)
Frame = +2
Query: 306 ETDCLNLFNAW-RTIEEDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNAN 364
ETDC +L N W + ++ +Y + +++DCR +++ + + + ++ N VA L+R A
Sbjct: 1220 ETDCQDLTNTWNKPKNQNSTYYATILQDCRQMLANSQHILIVHSKKEANCVAHRLSRLAF 1399
Query: 365 TYANMVWVEEV 375
T + W+E+V
Sbjct: 1400 TTSERYWMEDV 1432
>BI973267
Length = 406
Score = 45.8 bits (107), Expect = 3e-05
Identities = 29/104 (27%), Positives = 42/104 (39%), Gaps = 1/104 (0%)
Frame = +1
Query: 35 RDILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKET 94
+D W +NG +T+K+ Y L+ E GF P+
Sbjct: 58 KDTWTWRAEANGEFTTKSTYQVLQS----ELDDQGQYEGFQQ-----LCNIKIPPKALSF 210
Query: 95 CWRAISGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLC 137
WR + LP +D L +R + V+ C C + ETA HLF C
Sbjct: 211 AWRLL*DRLPTKDNLVKRQIQVENDLCPFCHNQPETASHLFFKC 342
>TC233392
Length = 1145
Score = 45.1 bits (105), Expect = 6e-05
Identities = 41/173 (23%), Positives = 67/173 (38%), Gaps = 8/173 (4%)
Frame = -1
Query: 36 DILFWPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETC 95
D W N Y++++ Y L+ +E A+ + W +
Sbjct: 656 DTWVWKHEPNELYSTRSAYKLLQ--GHLEGEDQDGAL-------QDLWKLKIPAKASIFA 504
Query: 96 WRAISGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLCPAAKQV*YASSL---SL 151
WR I LP + L RR V ++ C C E A H+F+ C + + + S SL
Sbjct: 503 WRLIRDRLPTKSNLHRRQVVLEDSLCPFCRIREEDASHIFLECNKIRPLWWESQTWVRSL 324
Query: 152 RVDSFIAFADFWAWVVEQGDSEFISTVQT----IIYAIWEARNQVQFQQRTFS 200
V F V S+ + +T + +++W+ RNQV FQ F+
Sbjct: 323 GVSPINKRQHFLQHVHGTPGSKRYNRWKTWWIALTWSLWKHRNQVIFQNAQFN 165
>CF921567
Length = 686
Score = 44.7 bits (104), Expect = 7e-05
Identities = 29/100 (29%), Positives = 39/100 (39%), Gaps = 1/100 (1%)
Frame = -1
Query: 45 NGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLP 104
NG Y++K+ Y L + A R W+ PR WR LP
Sbjct: 677 NGLYSTKSAYKVL*E---------GHASAIEDRVLNIMWSLKIPPRASAFSWRLFKNRLP 525
Query: 105 LRDRLFRRHVDVDP-GCCLCSGENETAEHLFMLCPAAKQV 143
R+ L RR V + C LC E E+ HLF C + +
Sbjct: 524 TRENLKRRQVSLHTYSCPLCDLEEESVNHLFFNCSKTRSL 405
>BI702277
Length = 420
Score = 44.3 bits (103), Expect = 9e-05
Identities = 23/73 (31%), Positives = 40/73 (54%)
Frame = +2
Query: 264 GEAMAAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDR 323
GE +A+A SP +AE +A +W++Q+A L + ETDC + +AWR +
Sbjct: 2 GEVLASATRLEDQCYSPQIAEYLAFKWSIQMANQLLLQNVFFETDCREIASAWRNASQ-- 175
Query: 324 SYLSILIRDCRLL 336
+ I ++ CR++
Sbjct: 176 RIVHI*MQFCRIV 214
Score = 37.4 bits (85), Expect = 0.012
Identities = 19/56 (33%), Positives = 32/56 (56%)
Frame = +1
Query: 321 EDRSYLSILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNANTYANMVWVEEVP 376
+D +YL +++DCR + + + V+L ++ GN VAD LA A + VE+ P
Sbjct: 175 KDCTYLDAILQDCRDMTTHIDTVALVHTKKIGNIVADSLA*LAFVLVDQSGVEDAP 342
>AW423846
Length = 385
Score = 44.3 bits (103), Expect = 9e-05
Identities = 33/117 (28%), Positives = 54/117 (45%), Gaps = 10/117 (8%)
Frame = +3
Query: 96 WRAISGYLPLRDRLFRRHVDV-DPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVD 154
WR I LP + L RRHV++ D C C + E A HLF C + +A SLS +
Sbjct: 39 WRLIRDRLPTKTNLHRRHVEIHDLTCPFCKNKEEDAAHLFFTCSKIMPLWWA-SLSW-TN 212
Query: 155 SFIAFAD-----FWAWVVEQGDSEFISTVQ----TIIYAIWEARNQVQFQQRTFSVT 202
+ F + F V+ G Q ++ ++IW+ RN++ ++ F+ +
Sbjct: 213 TTAPFPENPHMHFLQQVLRNGKDTSYQKWQCWWTSLTWSIWQHRNKIVXEEERFNAS 383
>TC231841
Length = 791
Score = 43.9 bits (102), Expect = 1e-04
Identities = 39/174 (22%), Positives = 60/174 (34%), Gaps = 8/174 (4%)
Frame = +3
Query: 40 WPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAI 99
W +G YT+K+ Y L E ++ W + WR I
Sbjct: 75 WKADPSGQYTAKSAYGVLWGEMFEEQQDG---------VFEELWKLKLPSKITIFAWRLI 227
Query: 100 SGYLPLRDRLFRRHVDVD-PGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVDSFIA 158
LP R L R+ ++VD P C C E+A HLF C V + S + +
Sbjct: 228 RDRLPTRSNLRRKQIEVDDPRCPFCRSAEESAAHLFFHCSRIAPVWWESLSWVNLLGVFP 407
Query: 159 -------FADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVL 205
+ S + + + IW+ RN + F TF+ +L
Sbjct: 408 NHPRQHFLQHIYGVTAGMRASRWKWWWLALTWTIWKQRNNMIFSNGTFNANKIL 569
>CO984521
Length = 716
Score = 43.5 bits (101), Expect = 2e-04
Identities = 38/174 (21%), Positives = 62/174 (34%), Gaps = 14/174 (8%)
Frame = -2
Query: 40 WPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAI 99
W +G Y++K+ Y L + E +K W + WR I
Sbjct: 634 WAAEPSGCYSTKSAYKALHHVTVGEEQDGK---------FKELWKLRVPLKVAIFAWRLI 482
Query: 100 SGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLCPAAKQV*YASSLSLRVDSFIA 158
LP + L ++ V++ C LC ETA HLF C + + S S++
Sbjct: 481 QDKLPTKANLRKKRVELQEYLCPLCRSVEETASHLFFHCSKVSPLWWES------QSWVN 320
Query: 159 FADFWAWVVEQGDSEFISTVQ-------------TIIYAIWEARNQVQFQQRTF 199
+ + +Q S+ I + Y+IW+ RN + F F
Sbjct: 319 MMGVFPYQPDQHFSQHIFGASVGLQGKRWQWWWFALTYSIWKHRNSIIFSNANF 158
>TC231904 similar to GB|AAL25568.1|16648753|AY058152 AT5g16150/T21H19_70
{Arabidopsis thaliana;} , partial (17%)
Length = 903
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/43 (44%), Positives = 24/43 (55%), Gaps = 1/43 (2%)
Frame = +1
Query: 96 WRAISGYLPLRDRLFRRHVDVDPGCC-LCSGENETAEHLFMLC 137
WR I LP + L RR +D+ C CS ++ETA HLF C
Sbjct: 283 WRLIRDRLPTKSNLRRRQIDISDSLCPFCSIKDETASHLFFEC 411
>AW830836
Length = 294
Score = 40.4 bits (93), Expect = 0.001
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Frame = -2
Query: 96 WRAISGYLPLRDRLFRRHVDV---DPGCCLCSGENETAEHLFMLCPAAKQV 143
W+ I + R +L +R + + D C LC GE ET +HL CP A +
Sbjct: 164 WKVIRDKIQTRGKLKKRQIKLQAEDYSCALCHGEEETIQHLLFTCPFASTI 12
>TC223713
Length = 420
Score = 38.9 bits (89), Expect = 0.004
Identities = 25/99 (25%), Positives = 39/99 (39%), Gaps = 1/99 (1%)
Frame = +1
Query: 40 WPESSNGHYTSKAGYVFLRKLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAI 99
W +G YT++ Y +R++ G R ++ + WR +
Sbjct: 10 WEADQSGQYTAQTAYNLMREVE---------VGGIQDRAFEEL*KLKVPIKFAVFAWRLL 162
Query: 100 SGYLPLRDRLFRRHVDV-DPGCCLCSGENETAEHLFMLC 137
LP + L RR ++V + C CS E A HLF C
Sbjct: 163 RDRLPTKANLHRRQIEVTNRSCPFCSNMEEEAGHLFFHC 279
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.138 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,782,315
Number of Sequences: 63676
Number of extensions: 292726
Number of successful extensions: 2145
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 2130
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2140
length of query: 387
length of database: 12,639,632
effective HSP length: 99
effective length of query: 288
effective length of database: 6,335,708
effective search space: 1824683904
effective search space used: 1824683904
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0052a.1


