

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.7
(189 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
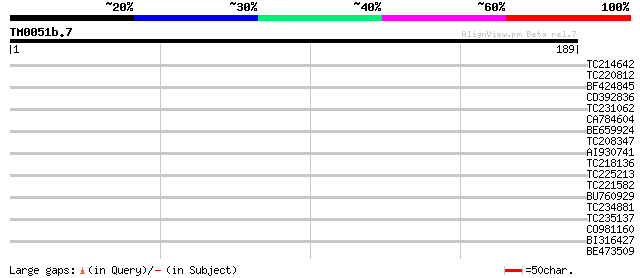
Score E
Sequences producing significant alignments: (bits) Value
TC214642 similar to UP|Q6S9Z3 (Q6S9Z3) Allantoin permease, parti... 32 0.18
TC220812 similar to UP|Q9LRM9 (Q9LRM9) Gb|AAD27719.1, partial (57%) 32 0.18
BF424845 similar to SP|Q41706|A3_V A3 protein. [Cowpea] {Vigna u... 31 0.39
CD392836 28 3.3
TC231062 weakly similar to UP|Q6N8I3 (Q6N8I3) CycJ cytochrome-c ... 27 5.6
CA784604 similar to GP|3193328|gb| F6N15.13 gene product {Arabid... 27 5.6
BE659924 27 5.6
TC208347 similar to UP|Q9XF63 (Q9XF63) RING-H2 zinc finger prote... 27 5.6
AI930741 27 7.4
TC218136 27 7.4
TC225213 UP|O64970 (O64970) Cationic peroxidase 2 , partial (64%) 27 7.4
TC221582 similar to UP|Q93YZ7 (Q93YZ7) At2g31810/F20M17.15, part... 27 7.4
BU760929 26 9.6
TC234881 26 9.6
TC235137 weakly similar to UP|Q7PV56 (Q7PV56) ENSANGP00000014031... 26 9.6
CO981160 26 9.6
BI316427 similar to GP|2674203|gb|A CLP protease regulatory subu... 26 9.6
BE473509 similar to GP|9758500|dbj| contains similarity to AT-ho... 26 9.6
>TC214642 similar to UP|Q6S9Z3 (Q6S9Z3) Allantoin permease, partial (69%)
Length = 955
Score = 32.0 bits (71), Expect = 0.18
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +2
Query: 119 VGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDISLEYISF 169
VG + +M+ + N+ I G P F QW V+ S+ LGI ++ L + +
Sbjct: 293 VG*ARALMISQIS*FNLLRIIGPPFFLQWGVVCSLALGICPANMHLLLLGY 445
>TC220812 similar to UP|Q9LRM9 (Q9LRM9) Gb|AAD27719.1, partial (57%)
Length = 1088
Score = 32.0 bits (71), Expect = 0.18
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +3
Query: 125 VMVKENVHINIFDIQGAPLFYQWIVITSVVLG 156
+++K NVH N+F Q LFY W++ +++ G
Sbjct: 198 ILMKMNVHGNVFYQQVQLLFYPWVISLAILFG 293
>BF424845 similar to SP|Q41706|A3_V A3 protein. [Cowpea] {Vigna unguiculata},
partial (29%)
Length = 325
Score = 30.8 bits (68), Expect = 0.39
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = +3
Query: 119 VGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDI 162
VG +M+++ +IN+ I G F QW V+ + LGI ++
Sbjct: 141 VG*ALPLMIRQISYINLLRIMGPSFFLQWGVVCDLSLGICQANV 272
>CD392836
Length = 586
Score = 27.7 bits (60), Expect = 3.3
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -1
Query: 98 PSLNPPSDPSEPSGPVFGRGPVGFVQEVMVKENVHINI 135
PSL P PS GPV R V V+V++N I++
Sbjct: 295 PSLAPAPSPSPSYGPVISREKVRDALLVLVQDNQFIDM 182
>TC231062 weakly similar to UP|Q6N8I3 (Q6N8I3) CycJ cytochrome-c biosynthesis
protein precursor, partial (41%)
Length = 837
Score = 26.9 bits (58), Expect = 5.6
Identities = 18/49 (36%), Positives = 23/49 (46%)
Frame = +1
Query: 3 CVGLDRFRVGRVTGGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRC 51
C G R G V GG R HR+ LR + I + +G+ LRRC
Sbjct: 217 CPGRQRGVPGVVAGGGVRRHRSHLRHRGTVRGIV---AGFVQGRALRRC 354
>CA784604 similar to GP|3193328|gb| F6N15.13 gene product {Arabidopsis
thaliana}, partial (47%)
Length = 420
Score = 26.9 bits (58), Expect = 5.6
Identities = 17/36 (47%), Positives = 20/36 (55%), Gaps = 1/36 (2%)
Frame = +2
Query: 84 NRNRPIQLIWSR-FFPSLNPPSDPSEPSGPVFGRGP 118
+R RP W R FFP+L+PPS P PS G P
Sbjct: 2 HRPRP----WPRSFFPTLSPPSSP--PSATRTGASP 91
>BE659924
Length = 588
Score = 26.9 bits (58), Expect = 5.6
Identities = 27/104 (25%), Positives = 47/104 (44%), Gaps = 4/104 (3%)
Frame = -3
Query: 70 RRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSDPSE----PSGPVFGRGPVGFVQEV 125
RRR + +R+ N N + P + P SD P P+ RG G V
Sbjct: 529 RRRRIESLRATATANANHRL---CPHRRPEIRPGSDLGARFHIPLHPIIIRGTRG---TV 368
Query: 126 MVKENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDISLEYISF 169
M+ +N+F + + L ++ + I+ ++LG+ + S+E I F
Sbjct: 367 MIPPLTTLNLFPLSFSILLWKMMRISFLILGL---EFSVEEIVF 245
>TC208347 similar to UP|Q9XF63 (Q9XF63) RING-H2 zinc finger protein ATL3
(At1g72310) (RING-H2 zinc finger protein (ATL3);
86824-85850) (RING-H2 zinc finger protein ATL3;
90350-91324), partial (24%)
Length = 1229
Score = 26.9 bits (58), Expect = 5.6
Identities = 15/47 (31%), Positives = 21/47 (43%)
Frame = +1
Query: 52 SGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNRPIQLIWSRFFP 98
SGG A + AA + R V V +DW P P + +S+ P
Sbjct: 331 SGGAALLPNLAATAAAALFSRPVQTVEADWIPRC*APYRCWYSKAMP 471
>AI930741
Length = 458
Score = 26.6 bits (57), Expect = 7.4
Identities = 12/52 (23%), Positives = 27/52 (51%)
Frame = -3
Query: 86 NRPIQLIWSRFFPSLNPPSDPSEPSGPVFGRGPVGFVQEVMVKENVHINIFD 137
+RP ++ W P ++PP +P+ G + + +++VK + +N+ D
Sbjct: 426 HRPTEVHWRAPAPKVSPPLHCDDPA----MLGKLTLLLDILVKAEMAVNVID 283
>TC218136
Length = 1097
Score = 26.6 bits (57), Expect = 7.4
Identities = 16/55 (29%), Positives = 26/55 (47%), Gaps = 1/55 (1%)
Frame = +1
Query: 62 QRMAAQVMRRRVVAV-VRSDWAPNRNRPIQLIWSRFFPSLNPPSDPSEPSGPVFG 115
Q+ Q + +++ + VR D AP + + PS PP PS+P +FG
Sbjct: 778 QQPGVQYLEQQMYGLSVRDDGAPKNSYQVSS------PSYVPPGKPSKPEDKLFG 924
>TC225213 UP|O64970 (O64970) Cationic peroxidase 2 , partial (64%)
Length = 859
Score = 26.6 bits (57), Expect = 7.4
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 11/43 (25%)
Frame = -3
Query: 117 GPVGF-----------VQEVMVKENVHINIFDIQGAPLFYQWI 148
GP+GF V+ V+ V IN++ GAP F QW+
Sbjct: 836 GPLGFWNGHPGHFFKNVRNVVKGSKVGINLWVQTGAPTFTQWV 708
>TC221582 similar to UP|Q93YZ7 (Q93YZ7) At2g31810/F20M17.15, partial (19%)
Length = 505
Score = 26.6 bits (57), Expect = 7.4
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = -2
Query: 10 RVGRVTGGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRR 50
R + +G S R HR R R++ + +S+Q E EG+ RR
Sbjct: 309 RTRQQSGLSFRSHRRRTRKSCVAEPLSEQGREEREGRWWRR 187
>BU760929
Length = 446
Score = 26.2 bits (56), Expect = 9.6
Identities = 16/59 (27%), Positives = 27/59 (45%)
Frame = -1
Query: 23 RNRLRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDW 81
R Q E++ +R EEE + GGV R + A+ + Q R++V++ W
Sbjct: 188 REEPNQRMQERSRCFERYSEEEEEGEGDGGGGVGRYIPARSLRHQHRRKKVISGTGMVW 12
>TC234881
Length = 690
Score = 26.2 bits (56), Expect = 9.6
Identities = 11/19 (57%), Positives = 11/19 (57%)
Frame = +1
Query: 96 FFPSLNPPSDPSEPSGPVF 114
FF NPPSDPS G F
Sbjct: 196 FFTPRNPPSDPSGSGGRAF 252
>TC235137 weakly similar to UP|Q7PV56 (Q7PV56) ENSANGP00000014031 (Fragment),
partial (6%)
Length = 577
Score = 26.2 bits (56), Expect = 9.6
Identities = 15/48 (31%), Positives = 21/48 (43%)
Frame = -1
Query: 59 VAAQRMAAQVMRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSDP 106
+ Q V+RR VR RN+P+QL++ L PP P
Sbjct: 313 IELQPQLQDVLRRLDEEPVRVHSGAARNQPLQLLYPPLHTLLPPPPPP 170
>CO981160
Length = 703
Score = 26.2 bits (56), Expect = 9.6
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = +3
Query: 85 RNRPIQLIWSRFFPSLNPPS 104
RN P+ +W FF +PPS
Sbjct: 456 RNLPLDFLWDAFFQVSHPPS 515
>BI316427 similar to GP|2674203|gb|A CLP protease regulatory subunit CLPX
{Arabidopsis thaliana}, partial (4%)
Length = 378
Score = 26.2 bits (56), Expect = 9.6
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 2/33 (6%)
Frame = -1
Query: 88 PIQLIWS--RFFPSLNPPSDPSEPSGPVFGRGP 118
P+Q+ + + FP L PP PS S P FG P
Sbjct: 375 PLQISFGVGKLFPKLEPPQHPSFES-PEFGAAP 280
>BE473509 similar to GP|9758500|dbj| contains similarity to AT-hook
DNA-binding protein~gene_id:MZA15.3 {Arabidopsis
thaliana}, partial (12%)
Length = 483
Score = 26.2 bits (56), Expect = 9.6
Identities = 12/37 (32%), Positives = 19/37 (50%), Gaps = 3/37 (8%)
Frame = +1
Query: 78 RSDWAPNRNRPIQLI---WSRFFPSLNPPSDPSEPSG 111
RS P RP+ + W++ P L PP+ P + +G
Sbjct: 328 RSGAGPENTRPMATLRWAWAQLTPPLPPPTPPPKNTG 438
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,000,561
Number of Sequences: 63676
Number of extensions: 105591
Number of successful extensions: 950
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 941
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 947
length of query: 189
length of database: 12,639,632
effective HSP length: 92
effective length of query: 97
effective length of database: 6,781,440
effective search space: 657799680
effective search space used: 657799680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0051b.7


