

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
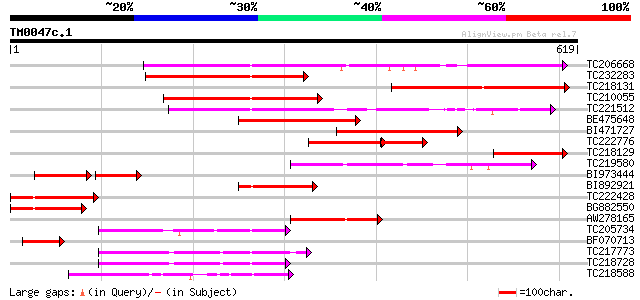
Query= TM0047c.1
(619 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC206668 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tran... 338 4e-93
TC232283 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tr... 242 3e-64
TC218131 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (8%) 241 9e-64
TC210055 similar to UP|Q9LRX7 (Q9LRX7) Phosphatidylinositol/phos... 234 1e-61
TC221512 similar to UP|Q94A34 (Q94A34) C7A10_870/C7A10_870 (At4g... 233 2e-61
BE475648 216 2e-56
BI471727 178 6e-45
TC222776 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tran... 119 2e-41
TC218129 weakly similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial ... 140 1e-33
TC219580 similar to UP|Q9FWS5 (Q9FWS5) F1B16.10 protein, partial... 125 7e-29
BI973444 homologue to GP|14486705|gb phosphatidylinositol transf... 77 6e-27
BI892921 similar to GP|15215780|gb| C7A10_870/C7A10_870 {Arabido... 98 9e-21
TC222428 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tr... 95 1e-19
BG882550 homologue to GP|14486705|gb| phosphatidylinositol trans... 93 4e-19
AW278165 similar to GP|14486705|gb phosphatidylinositol transfer... 89 8e-18
TC205734 similar to UP|O48939 (O48939) Polyphosphoinositide bind... 82 5e-16
BF070713 similar to GP|8778498|gb|A F20N2.11 {Arabidopsis thalia... 82 9e-16
TC217773 similar to UP|O48939 (O48939) Polyphosphoinositide bind... 79 5e-15
TC218728 UP|O48939 (O48939) Polyphosphoinositide binding protein... 76 4e-14
TC218588 UP|O48940 (O48940) Polyphosphoinositide binding protein... 71 1e-12
>TC206668 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (72%)
Length = 1782
Score = 338 bits (867), Expect = 4e-93
Identities = 195/484 (40%), Positives = 288/484 (59%), Gaps = 21/484 (4%)
Frame = +1
Query: 147 YYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIA 206
YYP G+HGVDKEGRPVYIERLGK P++LM++TT+DRY+KYHVQEFE+A + KFPAC+IA
Sbjct: 10 YYPHGHHGVDKEGRPVYIERLGKVDPNKLMQVTTMDRYVKYHVQEFEKAFKIKFPACTIA 189
Query: 207 AKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKM 266
AKR I S+TTILDVQG+G+KNFT++A +L+ + KID + YPETL QM+I+NAG GF ++
Sbjct: 190 AKRHIDSSTTILDVQGVGLKNFTKSARDLIMRLQKIDGDNYPETLCQMFIINAGPGF-RL 366
Query: 267 LWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGP 326
LW + F DPKT +KI +L +K KLLE+ID+S+LP+FLGG+CTC +GGCL++ KGP
Sbjct: 367 LWNTVKSFLDPKTTSKIHVLGNKYQSKLLEIIDASELPEFLGGTCTCADQGGCLRSDKGP 546
Query: 327 WNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF---QVHSLKGRCSDSSTAESGSDIND 383
W +P+I+K++ S EA R + + N + ++ Q +KG SD+STAESGS+ D
Sbjct: 547 WKNPEILKMILSGEARRARPVVKVLNSEGKVIAYARPQYPMVKG--SDTSTAESGSEAED 720
Query: 384 YSSPTRQRSSTHLRLAPVDEEVRVPDLNGY------YSCDDSAPTTEKVVE---NGQFHL 434
+SP +S +HLRL PV EE +V + Y D+ P +K V+ Q L
Sbjct: 721 IASPKAMKSYSHLRLTPVREEAKVVGKSSYAGGGNLAGYDEYVPMVDKAVDAAWKNQTSL 900
Query: 435 SQKQSLQ--------TNDMENVTHRTISEGT-SVNNLFRIIKEIVEKTNHLYVTRVLASF 485
+ Q+ + TN E + R + T LF + + +VT+ L +
Sbjct: 901 QRSQTSKGTPPLPDTTNTPEGIQARIVVALTVFFMTLFTLFHSVA-----CHVTKKLPAV 1065
Query: 486 MERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEK 545
+ T + T + + A E + + +RL LE
Sbjct: 1066SSN----------DDQGTSEPTLDATKTNYEDYRPPSPTPAYAEANLLSSMMKRLGELEV 1215
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
+ L +KP MP EKE++L ++ R+ ++E +L TK+ L+ A+ +Q E+ +++ +K
Sbjct: 1216KVDTLQSKPSEMPYEKEELLNAAVCRVDALEAELIATKKALYEALMRQEELLAYIDSQEK 1395
Query: 606 SNCR 609
+ R
Sbjct: 1396ARLR 1407
>TC232283 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (28%)
Length = 535
Score = 242 bits (618), Expect = 3e-64
Identities = 113/178 (63%), Positives = 143/178 (79%)
Frame = +1
Query: 149 PQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAK 208
P G+HG+DKEGRPVYIERLGK P++LM++TT+DRY+KYHVQEFE+A KFPACSIAAK
Sbjct: 4 PHGHHGIDKEGRPVYIERLGKVDPNKLMQVTTLDRYVKYHVQEFEKAFAIKFPACSIAAK 183
Query: 209 RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLW 268
R I S+TTILDV G+G+KNFT++A L+ + KID + YPETL QM+I+NAG GF ++LW
Sbjct: 184 RHIDSSTTILDVHGVGLKNFTKSARELITRLQKIDGDNYPETLCQMFIINAGPGF-RLLW 360
Query: 269 PAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGP 326
+ F DPKT +KI +L +K KLLEVID+S+LP+FLGG+CTC GGCL + KGP
Sbjct: 361 NTVKSFLDPKTTSKIHVLGNKYQSKLLEVIDASELPEFLGGTCTCEDHGGCLTSDKGP 534
>TC218131 similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (8%)
Length = 1077
Score = 241 bits (614), Expect = 9e-64
Identities = 124/195 (63%), Positives = 151/195 (76%)
Frame = +3
Query: 417 DDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHL 476
DDSAP EKV++ +FH++Q+QSLQ +D N+ S GTSVNN F +KE VEKTN L
Sbjct: 3 DDSAPAAEKVLDFDEFHITQEQSLQNDDTGNIACMENSTGTSVNNWFSFVKEKVEKTNLL 182
Query: 477 YVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPC 536
YV+RV+ FMERL+ F RSLR EFWRTQNN++PS EHN NN +AA E ERDHIL C
Sbjct: 183 YVSRVVIYFMERLVMFFRSLRLEFWRTQNNIYPSVAMEHN-NNPAAASEILSERDHILRC 359
Query: 537 AQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEI 596
QRL+RLEK F EL++KP G+PLEKE ML +S+DRIKSVEFDLEKTKRVLH+ V +QLEI
Sbjct: 360 MQRLERLEKTFGELSHKPAGIPLEKEHMLTNSLDRIKSVEFDLEKTKRVLHATVMKQLEI 539
Query: 597 ADLVENLQKSNCRVS 611
A+L+ENLQ S +V+
Sbjct: 540 AELLENLQASKSQVN 584
>TC210055 similar to UP|Q9LRX7 (Q9LRX7)
Phosphatidylinositol/phosphatidylcholine transfer
protein-like, partial (27%)
Length = 615
Score = 234 bits (596), Expect = 1e-61
Identities = 110/173 (63%), Positives = 140/173 (80%)
Frame = +3
Query: 169 KAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNF 228
+ PS+LM +TT+DR+LKYHVQ FE+ +EKFPACSIAAKR I TTTILDV G+ +F
Sbjct: 3 QVEPSKLMNVTTVDRFLKYHVQGFEKMFKEKFPACSIAAKRHIDKTTTILDVHGVNWVSF 182
Query: 229 TRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDS 288
++ A +L+ M KID + YPETL+QM+IVNAGSGF K+LW A+ F DP+T AKI +L +
Sbjct: 183 SKVAHDLVMRMQKIDGDNYPETLNQMFIVNAGSGF-KLLWNTAKGFLDPRTTAKIHVLGN 359
Query: 289 KSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEA 341
K +LLE+IDSSQLPDFLGGSC+CP +GGCL+++KGPWNDPDI+KL+HS EA
Sbjct: 360 KFQSRLLEIIDSSQLPDFLGGSCSCPNDGGCLRSNKGPWNDPDILKLLHSREA 518
>TC221512 similar to UP|Q94A34 (Q94A34) C7A10_870/C7A10_870
(At4g36490/C7A10_870), partial (43%)
Length = 1158
Score = 233 bits (594), Expect = 2e-61
Identities = 156/430 (36%), Positives = 231/430 (53%), Gaps = 7/430 (1%)
Frame = +1
Query: 174 RLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAA 233
+LM++TT++RYLKYHV+EFER K PACSIAAK+ I +TTILDVQG+G+K+ + A
Sbjct: 7 KLMQVTTMERYLKYHVKEFERTFAVKLPACSIAAKKHIDQSTTILDVQGVGLKSLNKAAR 186
Query: 234 NLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYK 293
+LL + KID + YPE+L++M+I+NAGSGF ++LW + F DPKT +KI +L +K K
Sbjct: 187 DLLQRLQKIDGDNYPESLNRMFIINAGSGF-RLLWNTIKSFLDPKTTSKIHVLGNKYQSK 363
Query: 294 LLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNE 353
LLE+ID+S+LP+FLGG+CTC +GGC+ + KGPWNDPDI+K+VH+ E R+ E
Sbjct: 364 LLEIIDASELPEFLGGTCTCADKGGCMLSDKGPWNDPDILKMVHNGEGKCKRKTLSGIEE 543
Query: 354 QQNFDSFQVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLN-G 412
+ R + STA N S P R VD + P
Sbjct: 544 K-------------RIIEDSTANQNLG-NKESFPERY---------DVDVQCLSPKKQCT 654
Query: 413 YYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEK 472
Y D P K V++ L+QK + + +G S
Sbjct: 655 VYKYDAFVPVLGKPVDSS*NTLTQKDKDALSKGADCFPSKTCDGYS-------------- 792
Query: 473 TNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVE------A 526
NH +V ++A M ++T IR R N+ P TE + S + A
Sbjct: 793 -NH-FVGGIMAIVM-GIVTMIRMTR--------NM-PRKITEAALYGSSGYYDGTMMKAA 936
Query: 527 AFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVL 586
F + + +R+ LE+ L+ KP +P EKE++L +++ R+ ++E DL TK+ L
Sbjct: 937 TFSCNDYMAMMKRMAELEEKVTILSMKPV-IPPEKEEVLNNALGRVTTIEQDLVATKKAL 1113
Query: 587 HSAVKQQLEI 596
A+ +Q+E+
Sbjct: 1114DDALARQVEL 1143
>BE475648
Length = 422
Score = 216 bits (551), Expect = 2e-56
Identities = 99/134 (73%), Positives = 119/134 (87%)
Frame = +1
Query: 250 TLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGG 309
+LH+MYI+NAG GFK+MLWPAAQKF D KTIAKIQ+L+ KSL KLL++IDSSQLPDFLGG
Sbjct: 4 SLHRMYIINAGPGFKRMLWPAAQKFLDAKTIAKIQVLEPKSLCKLLDIIDSSQLPDFLGG 183
Query: 310 SCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRC 369
+CTCP EGGCL++SKGPWNDPDI+K+VHS EAT RQ++R SNEQQN DSF + KG+C
Sbjct: 184 TCTCPGEGGCLRSSKGPWNDPDIMKMVHSVEATFERQIARMSNEQQNLDSFWICPQKGQC 363
Query: 370 SDSSTAESGSDIND 383
SD+STAESGSD++D
Sbjct: 364 SDTSTAESGSDLDD 405
>BI471727
Length = 421
Score = 178 bits (452), Expect = 6e-45
Identities = 87/138 (63%), Positives = 107/138 (77%)
Frame = +3
Query: 357 FDSFQVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSC 416
FDSFQ+H LK RCSD+STAESGSD+NDYSSP R RS + LAPV EEV+ PDLNGYYSC
Sbjct: 3 FDSFQMHPLKERCSDTSTAESGSDMNDYSSPNRHRSCPYPHLAPVHEEVKAPDLNGYYSC 182
Query: 417 DDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHL 476
DDSA EKV+E+ FHL+++Q LQTN++ N++ RT S GT VN+ F I+KE +EK N L
Sbjct: 183 DDSALAVEKVIESDHFHLNREQPLQTNNIGNISCRTDSGGTYVNSWFSIVKEKIEKINVL 362
Query: 477 YVTRVLASFMERLITFIR 494
YV RV+ FME+L+T R
Sbjct: 363 YVARVMTFFMEKLVTLFR 416
>TC222776 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (3%)
Length = 585
Score = 119 bits (297), Expect(2) = 2e-41
Identities = 57/85 (67%), Positives = 66/85 (77%)
Frame = +2
Query: 327 WNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAESGSDINDYSS 386
WNDPDI+KLVH+ EAT VRQ++R N Q FDSFQ+H LK RCSD+STAESGSD+NDYSS
Sbjct: 2 WNDPDIMKLVHNEEATFVRQITRMPNGQHTFDSFQMHPLKERCSDTSTAESGSDMNDYSS 181
Query: 387 PTRQRSSTHLRLAPVDEEVRVPDLN 411
P R RS + LAPV EEV + LN
Sbjct: 182 PNRHRSCPYPHLAPVHEEVNLEPLN 256
Score = 69.3 bits (168), Expect(2) = 2e-41
Identities = 31/52 (59%), Positives = 41/52 (78%)
Frame = +3
Query: 405 VRVPDLNGYYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEG 456
V+ PDLNGYYSCDDSA EKV+E+ FHL+++Q LQTN++ N++ RT S G
Sbjct: 360 VKAPDLNGYYSCDDSALAVEKVIESDHFHLNREQPLQTNNIGNISCRTDSGG 515
>TC218129 weakly similar to UP|Q9LFZ9 (Q9LFZ9) F20N2.11, partial (11%)
Length = 578
Score = 140 bits (354), Expect = 1e-33
Identities = 68/81 (83%), Positives = 76/81 (92%)
Frame = +2
Query: 529 ERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHS 588
ERD++LPC QRLQRLEKVFEELNNKP GMP EKE+MLMDSMDRIKSVEFDLEKTKRVLH+
Sbjct: 2 ERDYVLPCVQRLQRLEKVFEELNNKPDGMPQEKEQMLMDSMDRIKSVEFDLEKTKRVLHA 181
Query: 589 AVKQQLEIADLVENLQKSNCR 609
AV +QLEI +L+ENL+KSNCR
Sbjct: 182 AVMKQLEIVELLENLKKSNCR 244
>TC219580 similar to UP|Q9FWS5 (Q9FWS5) F1B16.10 protein, partial (16%)
Length = 798
Score = 125 bits (313), Expect = 7e-29
Identities = 86/281 (30%), Positives = 135/281 (47%), Gaps = 12/281 (4%)
Frame = +2
Query: 307 LGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLK 366
LGG+CTC GGC+++ KGPW DP+I+K+V S E RQ+ SN++
Sbjct: 11 LGGNCTCMDRGGCMRSDKGPWQDPNILKMVLSGEVQCSRQIVTVSNDEGTVIECDKPI-- 184
Query: 367 GRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVR-VPDLNGYYSCDDSAPTTEK 425
R SD+STAESGS++ D +SP + T+ RL PV EE R + +G+ D+ P +K
Sbjct: 185 -RSSDTSTAESGSEVEDITSPKASGNYTNPRLTPVHEEARLIGRASGFSEYDEYVPMVDK 361
Query: 426 VVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASF 485
V+ G +KQ N + + +S G S N Y+ V+ F
Sbjct: 362 AVDLG---WKEKQVATQNSYGSTENFLLSTGKSGGNC-------------AYILAVIVGF 493
Query: 486 MERLITFIRSLRFEFWR--------TQNNVHPSNTTEHNINNHS---AAVEAAFERDHIL 534
+ TF+RSL + + N+ P+ T + S + V + + I
Sbjct: 494 FVAIFTFVRSLALRVTKGIQDTKSDSAKNMLPNTTVDSITKEESRPPSPVPRLTKTELIS 673
Query: 535 PCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSV 575
+RL LE+ + L +KP MP EKE++L ++ R+ ++
Sbjct: 674 SALKRLGELEEKVDILQSKPNVMPYEKEELLNAAVYRVDAL 796
>BI973444 homologue to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (18%)
Length = 355
Score = 77.4 bits (189), Expect(2) = 6e-27
Identities = 33/51 (64%), Positives = 43/51 (83%)
Frame = +2
Query: 94 DDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEV 144
DDY+ +LRFLKAR F+IEK MW +ML WRKE+G DTI+++FEF+EL+EV
Sbjct: 203 DDYHMMLRFLKARKFDIEKAKHMWTDMLQWRKEFGADTIVQDFEFKELDEV 355
Score = 62.4 bits (150), Expect(2) = 6e-27
Identities = 37/63 (58%), Positives = 44/63 (69%), Gaps = 1/63 (1%)
Frame = +3
Query: 28 KIGTLRKKAMNASSKFTHSL-KKRGKRKIDYRVPAVSIEDVRDEGEETVVHELRQRLIER 86
+IG+L+KKA+NASSKF H+L KK +RK D RV +VSIEDVRD E V RQ LI
Sbjct: 3 RIGSLKKKALNASSKFKHTLRKKSSRRKSDGRVSSVSIEDVRDFEELQAVDAFRQSLIMD 182
Query: 87 GLL 89
LL
Sbjct: 183ELL 191
>BI892921 similar to GP|15215780|gb| C7A10_870/C7A10_870 {Arabidopsis
thaliana}, partial (14%)
Length = 352
Score = 98.2 bits (243), Expect = 9e-21
Identities = 44/87 (50%), Positives = 63/87 (71%)
Frame = +3
Query: 250 TLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGG 309
TL+Q++I+NAGSGF+ MLW A + F D +T+AKI +L L LLE IDSS LP FL
Sbjct: 3 TLNQLFIINAGSGFR-MLWKAVKTFLDVRTVAKIHVLGFNYLSVLLEAIDSSNLPTFLCV 179
Query: 310 SCTCPTEGGCLKTSKGPWNDPDIIKLV 336
+CTC GGCL + +GPW +P++++++
Sbjct: 180 NCTCSDYGGCLMSDRGPWKNPEVLEMI 260
>TC222428 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (18%)
Length = 627
Score = 94.7 bits (234), Expect = 1e-19
Identities = 55/98 (56%), Positives = 69/98 (70%), Gaps = 1/98 (1%)
Frame = +2
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRG-KRKIDYRV 59
+EG +DE +ERRSD E SEDERR+ +IG+L+KKA+NASSKF H+L+K+ +RK D RV
Sbjct: 332 FEGFSGSDEKKERRSDFEYSEDERRT-RIGSLKKKALNASSKFKHTLRKKSSRRKSDGRV 508
Query: 60 PAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYY 97
+VSIEDVRD E V RQ LI LLP DY+
Sbjct: 509 SSVSIEDVRDFEELQAVDAFRQSLIMDELLPEAFADYH 622
>BG882550 homologue to GP|14486705|gb| phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (15%)
Length = 419
Score = 92.8 bits (229), Expect = 4e-19
Identities = 52/85 (61%), Positives = 65/85 (76%), Gaps = 1/85 (1%)
Frame = +2
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKR-GKRKIDYRV 59
+EG +DE +ERRSD ENSEDERR+ +IG+L+KKA+NASSKF HSLKK+ +RK D RV
Sbjct: 161 FEGFSGSDEKKERRSDFENSEDERRT-RIGSLKKKALNASSKFKHSLKKKSSRRKSDGRV 337
Query: 60 PAVSIEDVRDEGEETVVHELRQRLI 84
+VSIEDVR+ E+ V RQ LI
Sbjct: 338 SSVSIEDVRNFEEQQAVDAFRQALI 412
>AW278165 similar to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (22%)
Length = 427
Score = 88.6 bits (218), Expect = 8e-18
Identities = 43/103 (41%), Positives = 66/103 (63%), Gaps = 2/103 (1%)
Frame = +2
Query: 307 LGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF--QVHS 364
LGG+CTC +GGCL++ KGPW +PDI K+V + A +Q+ + N ++ + V+
Sbjct: 2 LGGTCTCEDQGGCLRSDKGPWKNPDIFKMVLTGGAWRSKQVVKVLNNERKVIVYAKPVYP 181
Query: 365 LKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRV 407
+ +C+D+STAES S+ D SSP +S +HL L PV EE ++
Sbjct: 182 MV-KCNDTSTAESRSEAKDISSPKAMKSYSHLTLTPVHEEAKI 307
>TC205734 similar to UP|O48939 (O48939) Polyphosphoinositide binding protein
Ssh1p, partial (71%)
Length = 1491
Score = 82.4 bits (202), Expect = 5e-16
Identities = 68/215 (31%), Positives = 96/215 (44%), Gaps = 6/215 (2%)
Frame = +2
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE-FEFEELEEVLQYYPQGYHGVD 156
TL+RFLKARD+NI K +M + L WR E D +L + + + G G
Sbjct: 401 TLIRFLKARDWNIAKAHKMLIDCLNWRVENEIDNVLRKPIPMDLYRAIRDSQLIGMSGYS 580
Query: 157 KEGRPVYIERLGKAHPSRLMRITTIDR-----YLKYHVQEFERALQEKFPACSIAAKRRI 211
KEG PV +G ++T D+ Y++ H+Q E Q P + R I
Sbjct: 581 KEGLPVIAVGVG---------LSTYDKASDKYYIQSHIQLNEYRDQVILPTATRKHGRYI 733
Query: 212 FSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAA 271
+ +LD+ GL K LL A++ ID YPE YIVN F W
Sbjct: 734 GTCVKVLDMTGL--KFSALNQLRLLTAISTIDDLNYPEKTDTYYIVNVPYVF-SACWKVV 904
Query: 272 QKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
+ +T KIQ+L +LL+V+D + LP F
Sbjct: 905 KPLLQERTRRKIQVLQGCGKEELLKVMDYASLPHF 1009
>BF070713 similar to GP|8778498|gb|A F20N2.11 {Arabidopsis thaliana},
partial (7%)
Length = 424
Score = 81.6 bits (200), Expect = 9e-16
Identities = 39/45 (86%), Positives = 43/45 (94%)
Frame = +2
Query: 15 SDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRV 59
SDVENSEDERR S+IG L+KKAMNASS+FTHSLKKRGKRKIDY+V
Sbjct: 2 SDVENSEDERRPSRIGNLKKKAMNASSRFTHSLKKRGKRKIDYQV 136
>TC217773 similar to UP|O48939 (O48939) Polyphosphoinositide binding protein
Ssh1p, partial (92%)
Length = 1218
Score = 79.3 bits (194), Expect = 5e-15
Identities = 66/234 (28%), Positives = 104/234 (44%), Gaps = 2/234 (0%)
Frame = +1
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE--FEFEELEEVLQYYPQGYHGV 155
TL RFLKAR++N K +M + L WR + D IL + + + G G
Sbjct: 115 TLTRFLKAREWNATKAHKMIVDCLKWRVQNEIDNILSKPIIPTDLYRGIRDSQLIGLSGY 294
Query: 156 DKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTT 215
+EG PV+ +G + + ++ Y++ H+Q E + P+ S +R I +
Sbjct: 295 SREGLPVFAIGVGLSTFDK----ASVHYYVQSHIQINEYRDRVILPSASKKHERPITTCV 462
Query: 216 TILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFC 275
ILD+ GL + + LL ++ ID YPE + YIVNA F W +
Sbjct: 463 KILDMTGLKLSALNQ--IKLLTIISSIDDLNYPEKTNTYYIVNAPYIF-SACWKVVKPLL 633
Query: 276 DPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWND 329
+T K+Q+L +LL+++D + LP F C EG G N+
Sbjct: 634 QERTRRKVQVLQGCGRDELLKIMDYASLPHF------CRREGSGSSRHSGNGNE 777
>TC218728 UP|O48939 (O48939) Polyphosphoinositide binding protein Ssh1p,
complete
Length = 1416
Score = 76.3 bits (186), Expect = 4e-14
Identities = 63/211 (29%), Positives = 97/211 (45%), Gaps = 2/211 (0%)
Frame = +3
Query: 98 TLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE--FEFEELEEVLQYYPQGYHGV 155
TL+RFLKARD++ K +M + L WR + D IL + + V G G
Sbjct: 252 TLMRFLKARDWDPCKAHKMLVDCLNWRVQNEIDNILSKPIVPADLYRAVRDSQLIGLSGY 431
Query: 156 DKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTT 215
+EG PV+ +G + + ++ Y++ H+Q E + P+ S R I +
Sbjct: 432 SREGLPVFAIGVGLSTFDK----ASVHYYVQSHIQINEYRERIILPSASKKQGRPITTCI 599
Query: 216 TILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFC 275
+LD+ GL + + LL ++ ID YPE + YIVNA F W +
Sbjct: 600 KVLDMTGLKLSALNQ--IKLLTIISSIDDLNYPEKTNTYYIVNAPYIF-SACWKVVKPLL 770
Query: 276 DPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
+T KIQ+L +LL ++D S LP F
Sbjct: 771 QERTRRKIQVLPGCGRDELLTIMDYSSLPHF 863
>TC218588 UP|O48940 (O48940) Polyphosphoinositide binding protein Ssh2p,
complete
Length = 1030
Score = 71.2 bits (173), Expect = 1e-12
Identities = 68/252 (26%), Positives = 118/252 (45%), Gaps = 6/252 (2%)
Frame = +3
Query: 65 EDVRDEGEETVVHELR--QRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLL 122
+ V + ET + ++R + ++E + +D + + RFL+ARD ++EK M + L
Sbjct: 147 DGVAKDSTETELTKIRLLRAIVETRDPSSKEEDDFMIRRFLRARDLDVEKASAMLLKYLK 326
Query: 123 WRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTID 182
WR + + + + + + QG+ DK GRP+ + G H +
Sbjct: 327 WRNSFVPNGSVSVSDVPNELAQDKVFMQGH---DKIGRPILMV-FGGRHFQNKDGLDEFK 494
Query: 183 RYLKYHVQEFERAL---QEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAM 239
R++ Y + + ++ QEKF I +++G G N L+A+
Sbjct: 495 RFVVYVLDKVCASMPPGQEKF--------------VGIAELKGWGYSN--SDVRGYLSAL 626
Query: 240 TKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILD-SKSLYKLLEVI 298
+ I +YYPE L +++IVNA F K +W F D KT KI ++ +K LLE +
Sbjct: 627 S-ILQDYYPERLGKLFIVNAPYIFMK-VWQIVYPFIDNKTKKKIVFVEKNKVKSTLLEEM 800
Query: 299 DSSQLPDFLGGS 310
+ SQ+P+ GGS
Sbjct: 801 EESQVPEIFGGS 836
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,936,788
Number of Sequences: 63676
Number of extensions: 306627
Number of successful extensions: 1734
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 1685
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1697
length of query: 619
length of database: 12,639,632
effective HSP length: 103
effective length of query: 516
effective length of database: 6,081,004
effective search space: 3137798064
effective search space used: 3137798064
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0047c.1


