

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047b.8
(171 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
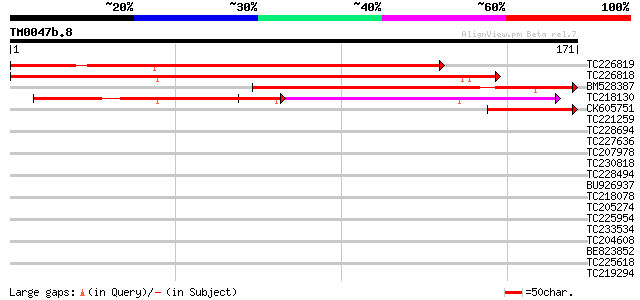
Score E
Sequences producing significant alignments: (bits) Value
TC226819 homologue to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, p... 222 8e-59
TC226818 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, par... 213 5e-56
BM528387 110 3e-25
TC218130 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, par... 98 2e-21
CK605751 44 3e-05
TC221259 similar to UP|Q8W4J7 (Q8W4J7) Peptidylprolyl isomerase ... 31 0.25
TC228694 UP|Q6S9W0 (Q6S9W0) Endo-1,3-beta-glucanase, complete 31 0.33
TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, p... 29 0.97
TC207978 weakly similar to UP|Q6YU70 (Q6YU70) Cyclin G-associate... 29 1.3
TC230818 similar to UP|Q9LDH9 (Q9LDH9) Gb|AAF34826.1 (MJM20.1 pr... 28 1.6
TC228494 similar to UP|Q6NQ48 (Q6NQ48) At1g34320, partial (13%) 28 2.2
BU926937 28 2.2
TC218078 similar to UP|Q71VM2 (Q71VM2) Plakoglobin/armadillo/bet... 28 2.2
TC205274 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T1... 28 2.2
TC225954 similar to UP|HS21_PEA (P19242) 17.1 kDa class II heat ... 28 2.2
TC233534 homologue to UP|Q6IID7 (Q6IID7) HDC18670, partial (16%) 28 2.8
TC204608 similar to UP|Q9LVI4 (Q9LVI4) Gb|AAF09073.1 (AT3g17860/... 28 2.8
BE823852 similar to GP|13605793|gb| At1g50300/F14I3_23 {Arabidop... 27 4.8
TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductas... 27 4.8
TC219294 weakly similar to UP|KI82_YEAST (P25341) Probable serin... 27 4.8
>TC226819 homologue to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (67%)
Length = 646
Score = 222 bits (565), Expect = 8e-59
Identities = 109/132 (82%), Positives = 116/132 (87%), Gaps = 1/132 (0%)
Frame = +1
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQ-AHTGRNLLQAKKGCSVN 59
M FS NQRF +SSSLLLL + +S SS + TFLSD VF SQ AHTGRNLLQAKKGCSVN
Sbjct: 256 MVFSFNQRFWVSSSLLLLAV---LSVSSLTPTFLSDTVFDSQPAHTGRNLLQAKKGCSVN 426
Query: 60 FEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYP 119
FEFLNYT+ITSKCKGP YPPK+CCGAFKEFACPY DVLNDLTN+CASTMFSYINLYGKYP
Sbjct: 427 FEFLNYTVITSKCKGPRYPPKECCGAFKEFACPYADVLNDLTNECASTMFSYINLYGKYP 606
Query: 120 PGLFAHECREGK 131
PGLFA ECREGK
Sbjct: 607 PGLFASECREGK 642
>TC226818 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (65%)
Length = 729
Score = 213 bits (541), Expect = 5e-56
Identities = 106/154 (68%), Positives = 119/154 (76%), Gaps = 6/154 (3%)
Frame = +3
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQA-HTGRNLLQAKKGCSVN 59
MAFS QRF LS SLLLL+L +++S+SS + T SD+VF SQA H GRNLLQAKK C VN
Sbjct: 30 MAFSFTQRFFLSPSLLLLLLLIALSASSLTPTCFSDSVFDSQAAHAGRNLLQAKKSCPVN 209
Query: 60 FEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYP 119
FEFLNYT+ITSKCKGP YPPK CC AFKEFACPY + LND TNDCA+ MFSYINLYGKYP
Sbjct: 210 FEFLNYTVITSKCKGPQYPPKQCCAAFKEFACPYANALNDSTNDCAAIMFSYINLYGKYP 389
Query: 120 PGLFAHECREGKEGLA--CP---ALPPSALADDT 148
G FA CR+GK+G+ CP A PPS DDT
Sbjct: 390 FGFFASVCRQGKKGITINCPASAASPPSVSEDDT 491
>BM528387
Length = 319
Score = 110 bits (275), Expect = 3e-25
Identities = 52/102 (50%), Positives = 69/102 (66%), Gaps = 4/102 (3%)
Frame = +1
Query: 74 GPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECREGKEG 133
GP YPPK CC AFK+FACP+ + +ND+T DCA+ MFSYIN+YGKYPPGLFA++C+EG +G
Sbjct: 1 GPQYPPKVCCEAFKQFACPFSNEVNDMTTDCATVMFSYINIYGKYPPGLFANQCKEGNQG 180
Query: 134 LACPALPPSALADDTSSQVVHFPS----LVLVLTACFLILLF 171
L + P+ +TSS +H + L + FL LF
Sbjct: 181 LDFFLVTPT----NTSSNTIHVAAPPHFFSLATISAFLGFLF 294
>TC218130 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (54%)
Length = 595
Score = 97.8 bits (242), Expect = 2e-21
Identities = 50/103 (48%), Positives = 60/103 (57%), Gaps = 6/103 (5%)
Frame = +1
Query: 70 SKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECRE 129
+ K PS CG + ACPY +VLNDLTNDCA+ MFS+INLYGKYPPG FA C E
Sbjct: 178 ANAKEPSTLQATLCGIERVVACPYANVLNDLTNDCATIMFSFINLYGKYPPGFFASMCHE 357
Query: 130 GKEGL------ACPALPPSALADDTSSQVVHFPSLVLVLTACF 166
GK+G+ A A PPS ADDT + V V+ +
Sbjct: 358 GKKGITINCLPAAAASPPSVSADDTGMKTHQMRVSVTVVVVVY 486
Score = 95.5 bits (236), Expect = 1e-20
Identities = 53/84 (63%), Positives = 60/84 (71%), Gaps = 8/84 (9%)
Frame = +3
Query: 8 RFLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQA-HTGRNLLQAKKGCSVNFEFLNYT 66
RF LS SLLLL+L +++SSSS L+++ F SQA HTGRNLLQAKK C VNFEFLNYT
Sbjct: 3 RFFLSPSLLLLLLLIALSSSS-----LTESSFNSQAEHTGRNLLQAKKSCPVNFEFLNYT 167
Query: 67 IITSKCKGPSYPP-------KDCC 83
IIT KCKG YPP K CC
Sbjct: 168IITCKCKGAQYPPSNAVRH*KSCC 239
>CK605751
Length = 724
Score = 44.3 bits (103), Expect = 3e-05
Identities = 20/27 (74%), Positives = 24/27 (88%)
Frame = -3
Query: 145 ADDTSSQVVHFPSLVLVLTACFLILLF 171
ADDT++Q VH PSL+L+L ACFLILLF
Sbjct: 722 ADDTANQAVHCPSLLLMLIACFLILLF 642
>TC221259 similar to UP|Q8W4J7 (Q8W4J7) Peptidylprolyl isomerase
(Cyclophilin)-like, partial (21%)
Length = 547
Score = 31.2 bits (69), Expect = 0.25
Identities = 30/83 (36%), Positives = 38/83 (45%)
Frame = -2
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITS 70
LSSS LALS SS SS S V G N+L + +G S + ++I
Sbjct: 399 LSSSSAFFSLALSCSSPSSGS------V*GLTNIVTLNILISSRGRS-SSSTGTFSITDR 241
Query: 71 KCKGPSYPPKDCCGAFKEFACPY 93
K P+ PPK C F+ FA Y
Sbjct: 240 VVKPPTTPPKTVCLKFR*FADLY 172
>TC228694 UP|Q6S9W0 (Q6S9W0) Endo-1,3-beta-glucanase, complete
Length = 1251
Score = 30.8 bits (68), Expect = 0.33
Identities = 24/85 (28%), Positives = 39/85 (45%), Gaps = 7/85 (8%)
Frame = +3
Query: 36 DAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYV- 94
D++ GS N+ +A ++ + T I + G SYPPKD G F A Y+
Sbjct: 447 DSLAGSVLPALENIQKAISAANLQGQMKVSTAIDTTLLGNSYPPKD--GVFSSSASSYIR 620
Query: 95 DVLNDLTNDCASTM------FSYIN 113
++N L + A + F+Y+N
Sbjct: 621 PIVNFLARNGAPLLANVYPYFAYVN 695
>TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, partial (8%)
Length = 1200
Score = 29.3 bits (64), Expect = 0.97
Identities = 34/104 (32%), Positives = 45/104 (42%), Gaps = 10/104 (9%)
Frame = -3
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSS-----SSTFLSDAVFGSQAHTGRNLLQAKKG 55
+ FS N SSS L A S+SSSSS SS+FLS + S + R +K
Sbjct: 499 LRFSFN---FASSSSFLAFSAFSISSSSSFVLTGSSSFLSLSSSSSDSSPQRATCFSKGA 329
Query: 56 CSVNFEFLNYTIITSKC-----KGPSYPPKDCCGAFKEFACPYV 94
V F L +++ C G S+ D G F F+C V
Sbjct: 328 GLVGFLALVV*LVSVLCATTFESGSSFTLTDSLGDF--FSCSIV 203
>TC207978 weakly similar to UP|Q6YU70 (Q6YU70) Cyclin G-associated
kinase-like protein, partial (8%)
Length = 1099
Score = 28.9 bits (63), Expect = 1.3
Identities = 19/44 (43%), Positives = 23/44 (52%), Gaps = 4/44 (9%)
Frame = -2
Query: 127 CREGKEGLACPALPPSALADDTSSQVVHFPSL----VLVLTACF 166
C G G AC L A ADD ++ V PSL VL ++ACF
Sbjct: 465 CC*GSSGKACHDLFADADADDVTAGEVFKPSLIGEGVLAVSACF 334
>TC230818 similar to UP|Q9LDH9 (Q9LDH9) Gb|AAF34826.1 (MJM20.1 protein),
partial (94%)
Length = 782
Score = 28.5 bits (62), Expect = 1.6
Identities = 19/55 (34%), Positives = 26/55 (46%)
Frame = -2
Query: 4 SLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSV 58
SL FL S L +LAL +S+S S + + F S H N L + CS+
Sbjct: 556 SLEDHFLTSLFSTLKLLALLNASTSFSKSNSNPFFFNSSTHFPSNPLSSFSRCSI 392
>TC228494 similar to UP|Q6NQ48 (Q6NQ48) At1g34320, partial (13%)
Length = 631
Score = 28.1 bits (61), Expect = 2.2
Identities = 17/38 (44%), Positives = 21/38 (54%)
Frame = -1
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAV 38
+ FSLN F LS S LLLI + + +S S F S V
Sbjct: 616 LQFSLNLIFKLSKSKLLLIRLIGIFLFTSCSMFCSSIV 503
>BU926937
Length = 451
Score = 28.1 bits (61), Expect = 2.2
Identities = 14/44 (31%), Positives = 21/44 (46%), Gaps = 4/44 (9%)
Frame = -3
Query: 106 STMFSYINLYGKY----PPGLFAHECREGKEGLACPALPPSALA 145
STM IN+ G + ++ +C G L C PPS++A
Sbjct: 431 STMMQPINILGNH*VDSASSIYCSQCSMGSRWLCCCKSPPSSIA 300
>TC218078 similar to UP|Q71VM2 (Q71VM2)
Plakoglobin/armadillo/beta-catenin-like protein
(Fragment), partial (88%)
Length = 1310
Score = 28.1 bits (61), Expect = 2.2
Identities = 15/34 (44%), Positives = 21/34 (61%)
Frame = +3
Query: 21 ALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKK 54
+LS SSSSSSS++ SD V G + R L ++
Sbjct: 72 SLSPSSSSSSSSYSSDGVAGFETEEPRTPLAVRR 173
>TC205274 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T18N14_50
{Arabidopsis thaliana;} , partial (79%)
Length = 1470
Score = 28.1 bits (61), Expect = 2.2
Identities = 17/38 (44%), Positives = 24/38 (62%)
Frame = +1
Query: 13 SSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLL 50
+SL++LILA ++S S F S+AVF + T NLL
Sbjct: 58 TSLMILILA----AASYFSNFKSNAVFSKTSRTNSNLL 159
>TC225954 similar to UP|HS21_PEA (P19242) 17.1 kDa class II heat shock
protein, partial (64%)
Length = 419
Score = 28.1 bits (61), Expect = 2.2
Identities = 19/54 (35%), Positives = 33/54 (60%), Gaps = 1/54 (1%)
Frame = -2
Query: 1 MAFSLNQRFLLSSSLLLLILA-LSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAK 53
+AFS + FL++ LL IL L+ S S SSS FLS + + + + R++++ +
Sbjct: 241 LAFSGSVNFLMNLPTLLSILKYLAPSFSLSSSLFLSPLIRSTLSSSVRSMIRCR 80
>TC233534 homologue to UP|Q6IID7 (Q6IID7) HDC18670, partial (16%)
Length = 657
Score = 27.7 bits (60), Expect = 2.8
Identities = 16/29 (55%), Positives = 20/29 (68%)
Frame = -2
Query: 4 SLNQRFLLSSSLLLLILALSVSSSSSSST 32
S+NQ FLL LLLL+L L + SSSS+
Sbjct: 92 SINQLFLLLLLLLLLLLLLLLLPPSSSSS 6
>TC204608 similar to UP|Q9LVI4 (Q9LVI4) Gb|AAF09073.1 (AT3g17860/MEB5_8),
partial (22%)
Length = 1439
Score = 27.7 bits (60), Expect = 2.8
Identities = 11/37 (29%), Positives = 15/37 (39%)
Frame = +3
Query: 74 GPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFS 110
G S P + C + E+ C Y + N C FS
Sbjct: 360 GTSIPKESFCNCWSEYECCYCEATTTRRNTCYGASFS 470
>BE823852 similar to GP|13605793|gb| At1g50300/F14I3_23 {Arabidopsis
thaliana}, partial (13%)
Length = 674
Score = 26.9 bits (58), Expect = 4.8
Identities = 17/36 (47%), Positives = 23/36 (63%), Gaps = 6/36 (16%)
Frame = +3
Query: 1 MAFSLNQRFLLSSSL------LLLILALSVSSSSSS 30
++FSL FLLS SL LL + +LS+ SSS+S
Sbjct: 366 LSFSLRSLFLLSPSLRSLPLSLLSLASLSIPSSSTS 473
>TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductase-like; Trp
repressor binding protein-like (1,4-benzoquinone
reductase-like protein), partial (75%)
Length = 1864
Score = 26.9 bits (58), Expect = 4.8
Identities = 18/37 (48%), Positives = 22/37 (58%)
Frame = +2
Query: 3 FSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAVF 39
FS + FLL LLLL+L L SSS SS+ S + F
Sbjct: 596 FSSSFFFLLLLLLLLLLLLLDEPSSSPSSSSSSSSSF 706
>TC219294 weakly similar to UP|KI82_YEAST (P25341) Probable
serine/threonine-protein kinase KIN82 , partial (5%)
Length = 711
Score = 26.9 bits (58), Expect = 4.8
Identities = 21/86 (24%), Positives = 35/86 (40%), Gaps = 8/86 (9%)
Frame = -1
Query: 9 FLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQ--------AKKGCSVNF 60
F+LS +LL++ + + S+ +TF + F T +L +KKGC +
Sbjct: 435 FMLSVPAILLLVLPTPTGSTGPTTFPAGTSFSIGLGTSGGVLHLINAQFTPSKKGCCLI- 259
Query: 61 EFLNYTIITSKCKGPSYPPKDCCGAF 86
P P+ CCG+F
Sbjct: 258 -----------SVAPLLTPRRCCGSF 214
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.137 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,331,528
Number of Sequences: 63676
Number of extensions: 179498
Number of successful extensions: 1220
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1190
length of query: 171
length of database: 12,639,632
effective HSP length: 90
effective length of query: 81
effective length of database: 6,908,792
effective search space: 559612152
effective search space used: 559612152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0047b.8


