

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046b.4
(1706 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
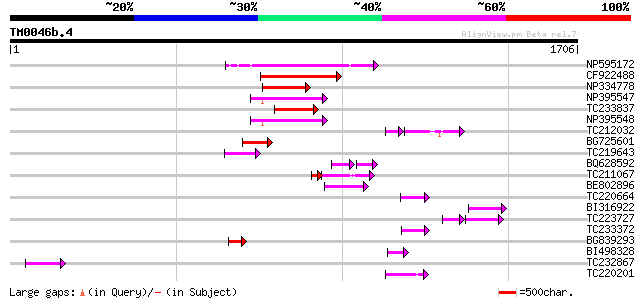
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 214 3e-55
CF922488 206 7e-53
NP334778 reverse transcriptase [Glycine max] 153 7e-37
NP395547 reverse transcriptase [Glycine max] 145 1e-34
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 140 4e-33
NP395548 reverse transcriptase [Glycine max] 132 1e-30
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 84 3e-22
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 101 2e-21
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 83 1e-15
BQ628592 55 7e-14
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 62 8e-12
BE802896 66 1e-10
TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 65 3e-10
BI316922 60 6e-09
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 43 2e-07
TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 55 3e-07
BG839293 55 3e-07
BI498328 52 2e-06
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 47 7e-05
TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance ... 47 9e-05
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 214 bits (544), Expect = 3e-55
Identities = 147/470 (31%), Positives = 237/470 (50%), Gaps = 9/470 (1%)
Frame = +1
Query: 650 QVQVKDKFLKIGSGLTTEQEDRLITLLGDNLDLFAWTINDVPGIDP--KVITHKLAIRPG 707
Q +V + LK L T + L LL +FA VP P + H + ++ G
Sbjct: 1627 QKEVPEDTLK---DLPTNIDPELAILLHTYAQVFA-----VPASLPPQREQDHAIPLKQG 1782
Query: 708 ATPV-IQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPT---WLANVVMVKKANGKWRM 763
+ PV ++P R +K+ + EK+I+ ++ + P+ + +++VKK +G WR
Sbjct: 1783 SGPVKVRPYRYPHTQKD-----QIEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRF 1947
Query: 764 CTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMT 823
CTDY +LN + KDS+P+P VD+L+D G + S +D SGY+QI++ P D E T F T
Sbjct: 1948 CTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRT 2127
Query: 824 NQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKE 883
+ +Y + MPFGL NA AT+Q LM+KIF + + + V+ DD+++ SA DH L+
Sbjct: 2128 HHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLES 2307
Query: 884 AFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQR 943
L+ +Q+ KCSFG +LG ++ G+ + K +A+L+ +P +VK+++
Sbjct: 2308 VLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRG 2487
Query: 944 LTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTP 1003
G RF+ + A P L+K+S F W E E AF KLK+ + PVLS P
Sbjct: 2488 FLGLTGYYRRFIKSYANIAGPLTDLLQKDS-FLWNNEAEAAFVKLKKAMTEAPVLSLPDF 2664
Query: 1004 SVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARR 1063
S P +L + V VL G+ I + S L + + LAI + +
Sbjct: 2665 SQPFILETDASGIGVGAVL----GQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALSK 2832
Query: 1064 LRPYFQSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELSEYD--IQYEP 1110
R Y + I+TD L+ ++ + + +W + YD I+Y+P
Sbjct: 2833 FRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIEYKP 2982
>CF922488
Length = 741
Score = 206 bits (524), Expect = 7e-53
Identities = 108/245 (44%), Positives = 148/245 (60%)
Frame = +3
Query: 754 VKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHP 813
V K +GK MC DY LN PKD +PLP+++ LVD + S MD +SGYNQI + P
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 814 SDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSAR 873
D E TTF+T +CYK M FGLKN GATYQR M +F + + +EVY+DDMIVKS
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 874 ASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMK 933
+H +L++ F +LR Y+++LNP KC F ++ K L F+ + RGIEV+ +K + ILEM
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 934 SPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLA 993
P + K+VQ GR+ + RF+ P F L KN +W +C AF ++K+ L
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 994 TLPVL 998
VL
Sbjct: 723 NPHVL 737
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 153 bits (386), Expect = 7e-37
Identities = 71/143 (49%), Positives = 93/143 (64%)
Frame = +3
Query: 762 RMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTF 821
RMC DY LN+ PKD++PLP++D L+ + L S MD +SGYNQI M P D E TTF
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 822 MTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDL 881
+T +CYK M FGLKN GATY R M +F + + +E YVD+MI KS +H +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 882 KEAFDQLRTYQMKLNPEKCSFGI 904
+ F QLR Y+++LNP KC FG+
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 145 bits (366), Expect = 1e-34
Identities = 83/248 (33%), Positives = 125/248 (49%), Gaps = 18/248 (7%)
Frame = +1
Query: 726 VQLETEKLIKARFIREVQYPTWLANVVMVKKANG------------------KWRMCTDY 767
V+ E KL++A I + +W++ V +V K G +WRMC DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 768 TSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQAN 827
LN+ KD YPLP +D+++ + +D YSGYNQI + P D+E T F +
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 828 YCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQ 887
+ Y+ MPFGL NA T+QR M IF V + +EV++DD A + +L++ +
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 888 LRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGR 947
+ LN EKC F +Q G LG ++ RGIEV +K I ++ P +VK + G
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 948 MAALSRFL 955
+ RF+
Sbjct: 721 VGFYRRFI 744
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 140 bits (354), Expect = 4e-33
Identities = 66/132 (50%), Positives = 93/132 (70%)
Frame = +2
Query: 798 SLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVG 857
S MD +SGYNQI M D E TTF+T + Y+ M FGLKN GATYQR M +F +
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMMH 184
Query: 858 RNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSR 917
+ +EVYVDDMI KS ++H +L + F +L+ YQ+KLNP KC+FG++ GK LGF+++ +
Sbjct: 185 KEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQK 364
Query: 918 GIEVNPDKGRAI 929
GIE++P+K +A+
Sbjct: 365 GIEIDPEKVKAL 400
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 132 bits (332), Expect = 1e-30
Identities = 76/248 (30%), Positives = 126/248 (50%), Gaps = 18/248 (7%)
Frame = +1
Query: 726 VQLETEKLIKARFIREVQYPTWLANVVMVKKANG------------------KWRMCTDY 767
V+ E KL++ I + W++ V++V K G W++C DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 768 TSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQAN 827
LN+ KD +PLP +D++++ +G+ +DAY GYNQI++ P D+E F
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 828 YCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQ 887
+ Y+ +PFGL NA T+Q M IF+ V +++EV++DD V L+ +
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 888 LRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGR 947
+ LN EKC F ++ G LG +++RGIEV+ K I ++ P++VK ++ G+
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 948 MAALSRFL 955
RF+
Sbjct: 721 ARFYRRFI 744
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 84.0 bits (206), Expect(2) = 3e-22
Identities = 59/190 (31%), Positives = 94/190 (49%), Gaps = 10/190 (5%)
Frame = +2
Query: 1189 LRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPR 1248
++ AI+ V+ L + GDS LV+ Q++GE + +DP L Y ++ L EI HV
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1249 GQNERADVLAKLASTGRLGNYQTVIQETLPRPSIDLVEIKLKA-------VKSVSEGEIS 1301
+N+ AD LA L S +L P + +E + + V+ +G+
Sbjct: 386 EENQMADALATLVSMFQL----------TPHGDLPYIEFRCRGRPAHCCLVEEERDGK-P 532
Query: 1302 WMESIKTFLES---PPKEDDLNTRTKRREASFYTLVGGELYRRGIMSPMLKCVDIKDALG 1358
W IK ++ES P + D + R KRR A+ + + G LY+R +L CV+ K+
Sbjct: 533 WYFDIKRYVESKEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVEN 712
Query: 1359 IMAEVHEGVF 1368
++ EVHEG F
Sbjct: 713 MLGEVHEGSF 742
Score = 41.6 bits (96), Expect(2) = 3e-22
Identities = 18/53 (33%), Positives = 30/53 (55%)
Frame = +1
Query: 1132 EKTQGEWVLSVDGSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEA 1184
++ + +W++ D +SN G G G + SPD I + + F +NN +EYEA
Sbjct: 34 DEDRDKWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEA 192
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 101 bits (252), Expect = 2e-21
Identities = 50/91 (54%), Positives = 62/91 (67%)
Frame = -3
Query: 700 HKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANG 759
HKLAI V Q +R++ EE+ + V E KL A FIR++ Y T L +VVMVKK NG
Sbjct: 277 HKLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNG 98
Query: 760 KWRMCTDYTSLNKVCPKDSYPLPNVDKLVDG 790
KWR+CTDY LN CPKD+YPLPN+D + DG
Sbjct: 97 KWRICTDYIDLN*ACPKDAYPLPNIDHMTDG 5
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 82.8 bits (203), Expect = 1e-15
Identities = 41/113 (36%), Positives = 66/113 (58%), Gaps = 4/113 (3%)
Frame = +3
Query: 646 EETKQVQVKD----KFLKIGSGLTTEQEDRLITLLGDNLDLFAWTINDVPGIDPKVITHK 701
EET+ V + + +KIG+G+T + LI LL D D+FAW+ D+PG+ ++ H+
Sbjct: 978 EETELVDLGSGSGKREVKIGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQHR 1157
Query: 702 LAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMV 754
L + P +PV Q RRM E + ++ E +K A F+ +YP W+AN+V +
Sbjct: 1158 LPLNPECSPVKQKLRRMKPETSLKIKEEVKK*FDAGFLAVARYPKWVANIVPI 1316
>BQ628592
Length = 423
Score = 55.1 bits (131), Expect(2) = 7e-14
Identities = 26/71 (36%), Positives = 43/71 (59%), Gaps = 2/71 (2%)
Frame = -1
Query: 969 LKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEE-- 1026
L KN W ++AF K+K++LA VL P P +LY+ + D+++ VL+Q +
Sbjct: 423 LPKNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
Query: 1027 GKKQKVIYFVS 1037
GKK++ IY++S
Sbjct: 243 GKKEQAIYYLS 211
Score = 42.0 bits (97), Expect(2) = 7e-14
Identities = 19/64 (29%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Frame = -3
Query: 1044 ELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELS 1102
++ Y +E+ ++ + RLR Y S + + + P++ + +KP L+GR+ W V LS
Sbjct: 193 KMNYSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLS 14
Query: 1103 EYDI 1106
E++I
Sbjct: 13 EFNI 2
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 62.4 bits (150), Expect(2) = 8e-12
Identities = 49/162 (30%), Positives = 71/162 (43%), Gaps = 1/162 (0%)
Frame = +1
Query: 937 SVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLP 996
SV +++ G + RF+P A+P +KKN F W E+ EQAF LKE L P
Sbjct: 109 SVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKAP 288
Query: 997 VLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALA 1056
VL+ P S L + V VLLQ + YF S L A L Y +K A
Sbjct: 289 VLALPDFSKTFELECDASGVGVRAVLLQ---GGHPIAYF-SEKLHSATLNYPTYDKELYA 456
Query: 1057 ILKTARRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSW 1097
+++ + + + I +D L+ + K L+ R W
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKW 582
Score = 27.7 bits (60), Expect(2) = 8e-12
Identities = 12/33 (36%), Positives = 21/33 (63%)
Frame = +2
Query: 909 FLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEV 941
F GF++ G++++P+K +AI E P V E+
Sbjct: 23 FSGFVVGRNGVQMDPEKIKAIQEWPPP*KVWEI 121
>BE802896
Length = 416
Score = 65.9 bits (159), Expect = 1e-10
Identities = 39/133 (29%), Positives = 61/133 (45%)
Frame = -2
Query: 946 GRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSV 1005
G RF+ A P L+K +F + ++C+ AF LK L T P++ P +
Sbjct: 406 GHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPDWTA 227
Query: 1006 PLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLR 1065
P L ++ A+ VL Q+ K +VIY+ S TL A+ Y EK LAI+ +
Sbjct: 226 PFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALEKFH 47
Query: 1066 PYFQSFQVKIKTD 1078
Y ++ + D
Sbjct: 46 SYLLGTRIIVYID 8
>TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (31%)
Length = 724
Score = 65.1 bits (157), Expect = 3e-10
Identities = 34/87 (39%), Positives = 51/87 (58%)
Frame = +1
Query: 1175 ASNNQSEYEALIAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRL 1234
A+NN +EY A+I G++ A++ G + I+GDS+LV Q+ G ++VK+ LS V + L
Sbjct: 61 ATNNAAEYRAMILGMKYALKKGFTGIRIQGDSKLVCMQIDGSWKVKNENLSTLYNVAKEL 240
Query: 1235 MMEVKEIKIEHVPRGQNERADVLAKLA 1261
+ +I HV R N AD A LA
Sbjct: 241 KDKFSSFQISHVLRNFNSDADAQANLA 321
>BI316922
Length = 405
Score = 60.5 bits (145), Expect = 6e-09
Identities = 37/113 (32%), Positives = 61/113 (53%)
Frame = +2
Query: 1382 KSRILLANDEERLFGVC*EM*KMSSLF*PTQGSTRRINYHDGALAVRHVGD*YLRTIPCG 1441
+SRILL N+E+ L +C EM*+M ++ + RR + +A+ H+ + +T P
Sbjct: 65 ESRILLVNNEKILHRICEEM*RMLEIWQHLSLACRRATQYSSTMALCHLRSRHTQTFPLI 244
Query: 1442 EGTNEVHHRGG*LLH*VDRS*GSGNYHGRQGQELPVAKDCL*IWGPQGISNG* 1494
+ T+E+ G * +H VDR * ++ Q E+ + + C+ IW PQ G*
Sbjct: 245 KETSEISPSGH*SVHQVDRD*AHRHHLNSQCSEVCLEEHCMLIWNPQHFDLG* 403
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 43.1 bits (100), Expect(2) = 2e-07
Identities = 26/70 (37%), Positives = 39/70 (55%), Gaps = 3/70 (4%)
Frame = +1
Query: 1302 WMESIKTFLESP---PKEDDLNTRTKRREASFYTLVGGELYRRGIMSPMLKCVDIKDALG 1358
W IK ++ S P+ D + RT RR A+ + + G LY+R L+CVD ++A
Sbjct: 139 WYFDIKRYVISKEYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANH 318
Query: 1359 IMAEVHEGVF 1368
++ EVHEG F
Sbjct: 319 MIEEVHEGSF 348
Score = 31.6 bits (70), Expect(2) = 2e-07
Identities = 38/115 (33%), Positives = 49/115 (42%), Gaps = 1/115 (0%)
Frame = +3
Query: 1372 RRAILGS*GDKSRILLANDEERLFGVC*EM*KMSSLF*PTQGSTRRINYHDGALAVRHVG 1431
+RA G KSR+LLA + L C EM +MSS+ Q T H LA HVG
Sbjct: 360 QRACYGQEDPKSRLLLAYHGK*LLCPCEEMPQMSSVRR*CQCPTASSECHVLPLAFLHVG 539
Query: 1432 D*YLRTI-PCGEGTNEVHHRGG*LLH*VDRS*GSGNYHGRQGQELPVAKDCL*IW 1485
+ R G + +H R L H V R H G ++ +D L IW
Sbjct: 540 NRCHRGH*AQGLEWSSLHPRSDRLFHQVGRGSFLYRRHEGCGGQVH*ERDHLPIW 704
>TC233372 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (12%)
Length = 561
Score = 55.1 bits (131), Expect = 3e-07
Identities = 27/82 (32%), Positives = 44/82 (52%)
Frame = +1
Query: 1180 SEYEALIAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVK 1239
+EY +LI GL + G + + ++GDS LV Q++G +++K+ + + L +
Sbjct: 1 AEYRSLILGLXHXXKKGYKXIIVQGDSLLVCNQIQGLWKIKNQNMGTLCXEAKELKDKFL 180
Query: 1240 EIKIEHVPRGQNERADVLAKLA 1261
KI H+PR N AD A LA
Sbjct: 181 SFKISHIPREYNSEADAQANLA 246
>BG839293
Length = 781
Score = 54.7 bits (130), Expect = 3e-07
Identities = 23/56 (41%), Positives = 36/56 (64%)
Frame = +1
Query: 658 LKIGSGLTTEQEDRLITLLGDNLDLFAWTINDVPGIDPKVITHKLAIRPGATPVIQ 713
+KIG+G+T + LI LL D D+FAW+ D+PG+ ++ H+L + P +PV Q
Sbjct: 562 VKIGTGITAPIREELIILLKDYQDIFAWSYQDMPGLSSDIVQHQLPLNPECSPVKQ 729
>BI498328
Length = 335
Score = 52.0 bits (123), Expect = 2e-06
Identities = 23/62 (37%), Positives = 36/62 (57%)
Frame = +1
Query: 1137 EWVLSVDGSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELG 1196
+W++ DG+SN G G G + SPD I + + F +NN +EYEA G++ AI+
Sbjct: 148 KWIVCFDGASNALGHGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFD 327
Query: 1197 VQ 1198
V+
Sbjct: 328 VK 333
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 47.0 bits (110), Expect = 7e-05
Identities = 36/126 (28%), Positives = 54/126 (42%), Gaps = 6/126 (4%)
Frame = -3
Query: 48 QPFVPAITRVDIPKHLQTMALDAFSG--ESDPMEHLRYF----NTKMVIGGATDAVKCRL 101
QP V A T P +Q + + F G DP HL + NT + G DA++ L
Sbjct: 851 QPEVQAQTITYPPSLIQLIQSNFFHGLPNEDPYAHLATYIEICNTIRLAGVPADAIRLSL 672
Query: 102 LPSTFKGMAMQWFIRQPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLK 161
L + G A +W S+ ++ ++ KFL ++ T + S HQ P E L
Sbjct: 671 LSFSLSGEAKRWLHSFKGNSLKSWDEVVEKFLKKYFPESKTAEGKAAISSFHQFPDESLS 492
Query: 162 TYMARF 167
+ RF
Sbjct: 491 EALERF 474
>TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance receptor
PtdSerR (Phosphatidylserine receptor), partial (5%)
Length = 1185
Score = 46.6 bits (109), Expect = 9e-05
Identities = 42/140 (30%), Positives = 73/140 (52%), Gaps = 12/140 (8%)
Frame = +1
Query: 1132 EKTQGEWV-LSVDGSS-----NNTGSGAGITIESPDKMI-IEQSLKFEFKASNNQSEYEA 1184
+K + WV L+VDGS + G G I E + +Q L + +E +A
Sbjct: 340 KKPESGWVKLNVDGSRIHEEPASAGCGGVIRDEWGTWCVGFDQKLDPNICRQAHYTELQA 519
Query: 1185 LIAGLRLAIE--LGVQKLFIKGDSQLVVKQVKGE--YQVKDPQLSKYLEVVRRLMMEVKE 1240
++ GL++A E + V+KL ++ DS+ V VK Y+ P+ K ++ + RL+ + K
Sbjct: 520 ILTGLKVAREDMINVEKLVVESDSEPAVNMVKSRLGYKYHRPEY-KVVQDINRLLKDPKW 696
Query: 1241 I-KIEHVPRGQNERADVLAK 1259
+ ++++VPR N A+ LAK
Sbjct: 697 LARLDYVPRKANRVANYLAK 756
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,592,769
Number of Sequences: 63676
Number of extensions: 838144
Number of successful extensions: 6443
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 6082
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6387
length of query: 1706
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1596
effective length of database: 5,635,272
effective search space: 8993894112
effective search space used: 8993894112
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0046b.4


