

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046a.5
(327 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
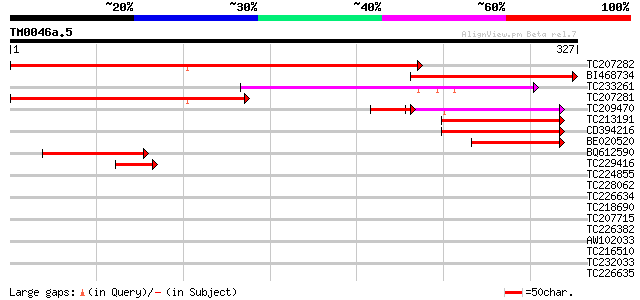
Score E
Sequences producing significant alignments: (bits) Value
TC207282 similar to UP|Q8IML8 (Q8IML8) CG31047-PA (RH18724p), pa... 263 1e-70
BI468734 similar to GP|21928139|gb AT4g18060/F15J5_30 {Arabidops... 172 1e-43
TC233261 weakly similar to UP|Q8L7W0 (Q8L7W0) AT4g18060/F15J5_30... 147 5e-36
TC207281 127 9e-30
TC209470 homologue to UP|TMG3_HUMAN (Q9BZD7) Transmembrane gamma... 93 5e-26
TC213191 87 8e-18
CD394216 similar to GP|16974678|gb SH3 domain-containing protein... 87 8e-18
BE020520 similar to GP|16974678|gb| SH3 domain-containing protei... 77 1e-14
BQ612590 homologue to GP|16974678|gb| SH3 domain-containing prot... 62 3e-10
TC229416 similar to UP|Q9LSP4 (Q9LSP4) Arabidopsis thaliana geno... 50 1e-06
TC224855 similar to UP|KHL1_HUMAN (Q9NR64) Kelch-like protein 1,... 33 0.13
TC228062 weakly similar to UP|Q39221 (Q39221) St12p protein, par... 31 0.87
TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 30 1.5
TC218690 similar to UP|O81812 (O81812) Auxilin-like protein, par... 29 2.5
TC207715 weakly similar to UP|Q9LHY8 (Q9LHY8) ESTs D15336(C0474)... 29 3.3
TC226382 similar to UP|GS11_ARATH (Q9LMP7) Golgi SNARE 11 protei... 29 3.3
AW102033 28 4.3
TC216510 weakly similar to UP|Q6T3R2 (Q6T3R2) NDR1-like protein,... 28 4.3
TC232033 28 4.3
TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 28 5.6
>TC207282 similar to UP|Q8IML8 (Q8IML8) CG31047-PA (RH18724p), partial (26%)
Length = 986
Score = 263 bits (671), Expect = 1e-70
Identities = 140/241 (58%), Positives = 176/241 (72%), Gaps = 3/241 (1%)
Frame = +3
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA+RKQASKLREQV++QQQAV+KQF GY SD VV D E+Q+HQ+LEKLY +TR
Sbjct: 51 MDAIRKQASKLREQVARQQQAVLKQFGGGGYGGSDNVVTDGVELQLHQRLEKLYISTRAG 230
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI--DNILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYGAEN N L++AA A
Sbjct: 231 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDSRKYGAENTCTSGNTLSRAALSFAQAHAQ 410
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE L + L +QV +PLR M+ G PLEDARHLAQRY RMRQEAE E+SKRQ +V
Sbjct: 411 IEKERGNLLKALGTQVAEPLRAMVVGAPLEDARHLAQRYDRMRQEAEAQAIEVSKRQAKV 590
Query: 179 RESPTS-EQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSD 237
RE P S E KL AAEA++++LK NM +LGKEAAAALA+V+AQQQRLT QR++AM+ ++
Sbjct: 591 REMPPSAENTMKLEAAEAKLQDLKTNMTILGKEAAAALAAVEAQQQRLTLQRIIAMVEAE 770
Query: 238 R 238
R
Sbjct: 771 R 773
>BI468734 similar to GP|21928139|gb AT4g18060/F15J5_30 {Arabidopsis
thaliana}, partial (30%)
Length = 376
Score = 172 bits (437), Expect = 1e-43
Identities = 81/96 (84%), Positives = 89/96 (92%)
Frame = +3
Query: 232 AMMVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTP 291
A MVSDRQKKESAPPV +SENG+ KTMYFLAEATHPF G+SEKELSFSKGDF+VVRKV+
Sbjct: 30 AEMVSDRQKKESAPPVGISENGTEKTMYFLAEATHPFSGESEKELSFSKGDFVVVRKVSQ 209
Query: 292 TGWSEGECNGKAGWFPSAYVEKRQRVPSSNFSSEVY 327
+GWSEGECNGKAGWFPSAYVEKRQR+PSSN + EVY
Sbjct: 210 SGWSEGECNGKAGWFPSAYVEKRQRIPSSNMAGEVY 317
>TC233261 weakly similar to UP|Q8L7W0 (Q8L7W0) AT4g18060/F15J5_30, partial
(24%)
Length = 648
Score = 147 bits (372), Expect = 5e-36
Identities = 86/212 (40%), Positives = 114/212 (53%), Gaps = 40/212 (18%)
Frame = +3
Query: 134 VLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTS-EQVAKLHA 192
+ +PLR I G PLEDARHL +RY ++ QE E E+ +R+ ++R S S E A+L
Sbjct: 12 ISEPLRAQITGAPLEDARHLTRRYDKLHQEVEAQAAEVLRRRSKLRNSSVSAESSARLQN 191
Query: 193 AEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMM------------------ 234
AE R+KELK+ +A LG+EA +A+ SV+ QQQ++T Q L M+
Sbjct: 192 AETRLKELKSALAALGREATSAMLSVEEQQQQMTLQSLRTMVDAERSYHQHVLVILEKLY 371
Query: 235 ---VSDRQKKESAP----------PVVVSENGSG--------KTMYFLAEATHPFYGDSE 273
+ DRQ KE+ P + N SG YF A+ HPF +E
Sbjct: 372 TEIIEDRQPKEATSFTLPKDGYNQPADENANSSGIDYKHNSQTATYFFAKVVHPFDAQAE 551
Query: 274 KELSFSKGDFIVVRKVTPTGWSEGECNGKAGW 305
ELS S DF+VVR+V P GWSEGEC G AGW
Sbjct: 552 GELSLSVDDFVVVRQVGPNGWSEGECKGNAGW 647
>TC207281
Length = 581
Score = 127 bits (318), Expect = 9e-30
Identities = 70/140 (50%), Positives = 93/140 (66%), Gaps = 2/140 (1%)
Frame = +1
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+A+RKQASKLREQV++QQQAV+KQF GY SD +V DE E+Q HQ+LEKLY +TR
Sbjct: 160 MEAIRKQASKLREQVARQQQAVLKQFGAGGYGGSDNMVTDEVELQQHQKLEKLYISTRAG 339
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI--DNILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYGA+N + L++AA AR
Sbjct: 340 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDNRKYGADNTCTSGSTLSRAALNYARARAQ 519
Query: 119 VEKEHEELNRLLASQVLDPL 138
+EKE L + L + V L
Sbjct: 520 MEKERGNLLKALGTXVCKAL 579
>TC209470 homologue to UP|TMG3_HUMAN (Q9BZD7) Transmembrane
gamma-carboxyglutamic acid protein 3 precursor, partial
(5%)
Length = 760
Score = 92.8 bits (229), Expect(2) = 5e-26
Identities = 49/113 (43%), Positives = 67/113 (58%), Gaps = 21/113 (18%)
Frame = +3
Query: 229 RLVAMMVSDRQKKESAPPVVV--------------------SENGSGKTM-YFLAEATHP 267
+L M+S+RQ+ E+ P V + NGS +M YFL E P
Sbjct: 153 QLEGEMISERQRIEAPPTPSVDSSMTPPPSYEEVNGVCASQAHNGSTDSMGYFLGEVLFP 332
Query: 268 FYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSS 320
++ +SE EL+ S GD+IV+RKVT GW+EGEC GKAGWFP Y+E+R+RV +S
Sbjct: 333 YHAESEVELNLSVGDYIVIRKVTNNGWAEGECKGKAGWFPFGYIERRERVLAS 491
Score = 42.7 bits (99), Expect(2) = 5e-26
Identities = 20/26 (76%), Positives = 25/26 (95%)
Frame = +1
Query: 209 KEAAAALASVDAQQQRLTFQRLVAMM 234
KEAAAA+A+V+AQQQRLT QRL+AM+
Sbjct: 31 KEAAAAMAAVEAQQQRLTLQRLIAMV 108
>TC213191
Length = 556
Score = 87.4 bits (215), Expect = 8e-18
Identities = 40/72 (55%), Positives = 54/72 (74%), Gaps = 1/72 (1%)
Frame = +3
Query: 250 SENGSGKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPS 308
+ NGS +M YFL E P+ SE EL+ S GD++VVRKVT +GW+EGEC G+AGWFP
Sbjct: 180 AHNGSTDSMGYFLGEVLFPYSAVSEVELNLSVGDYVVVRKVTNSGWAEGECKGRAGWFPF 359
Query: 309 AYVEKRQRVPSS 320
+Y+E+R+RV +S
Sbjct: 360 SYIERRERVLAS 395
>CD394216 similar to GP|16974678|gb SH3 domain-containing protein 2
{Arabidopsis thaliana}, partial (30%)
Length = 640
Score = 87.4 bits (215), Expect = 8e-18
Identities = 40/72 (55%), Positives = 54/72 (74%), Gaps = 1/72 (1%)
Frame = -2
Query: 250 SENGSGKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPS 308
+ NGS +M YFL E P+ SE EL+ S GD++VVRKVT +GW+EGEC G+AGWFP
Sbjct: 405 AHNGSTDSMGYFLGEVLFPYSAVSEVELNLSVGDYVVVRKVTNSGWAEGECKGRAGWFPF 226
Query: 309 AYVEKRQRVPSS 320
+Y+E+R+RV +S
Sbjct: 225 SYIERRERVLAS 190
>BE020520 similar to GP|16974678|gb| SH3 domain-containing protein 2
{Arabidopsis thaliana}, partial (17%)
Length = 410
Score = 76.6 bits (187), Expect = 1e-14
Identities = 32/54 (59%), Positives = 44/54 (81%)
Frame = -3
Query: 267 PFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSS 320
P+ SE EL+ S GD++VVRKVT +GW+EGEC G+AGWFP +Y+E+R+RV +S
Sbjct: 297 PYSAVSEVELNLSVGDYVVVRKVTNSGWAEGECKGRAGWFPFSYIERRERVLAS 136
>BQ612590 homologue to GP|16974678|gb| SH3 domain-containing protein 2
{Arabidopsis thaliana}, partial (11%)
Length = 428
Score = 62.4 bits (150), Expect = 3e-10
Identities = 30/61 (49%), Positives = 41/61 (67%)
Frame = +2
Query: 20 QAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKEIVKAAETFTAIGY 79
QAV+KQF GY SD +V DE E+Q HQ+LEKLY +TR + +Q++IV+ E + G
Sbjct: 242 QAVLKQFGAGGYGGSDNMVTDEVELQQHQKLEKLYISTRAGKHYQRDIVRGVEGYIVTGS 421
Query: 80 K 80
K
Sbjct: 422 K 424
>TC229416 similar to UP|Q9LSP4 (Q9LSP4) Arabidopsis thaliana genomic DNA,
chromosome 3, TAC clone:K14A17, partial (7%)
Length = 627
Score = 50.4 bits (119), Expect = 1e-06
Identities = 23/24 (95%), Positives = 24/24 (99%)
Frame = +3
Query: 62 DFQKEIVKAAETFTAIGYKHIETG 85
DFQKEIVKAA+TFTAIGYKHIETG
Sbjct: 300 DFQKEIVKAAKTFTAIGYKHIETG 371
>TC224855 similar to UP|KHL1_HUMAN (Q9NR64) Kelch-like protein 1, partial
(3%)
Length = 885
Score = 33.5 bits (75), Expect = 0.13
Identities = 26/72 (36%), Positives = 35/72 (48%), Gaps = 1/72 (1%)
Frame = +3
Query: 195 ARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDRQKKESAPPVVVSENGS 254
+R+ LKA +L K A + QQ LT A+ S++ APPVVV + S
Sbjct: 249 SRISSLKAQTILLQKMKRVVKAVEEETQQELT-----AVDESEQPSTSEAPPVVVPVSPS 413
Query: 255 GK-TMYFLAEAT 265
TM+F AE T
Sbjct: 414 DTLTMFFQAEGT 449
>TC228062 weakly similar to UP|Q39221 (Q39221) St12p protein, partial (54%)
Length = 1534
Score = 30.8 bits (68), Expect = 0.87
Identities = 18/63 (28%), Positives = 31/63 (48%), Gaps = 2/63 (3%)
Frame = +3
Query: 6 KQASK--LREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDF 63
+ ASK +R+ +S + +F G S D+VV++ MQIH ++K + T F
Sbjct: 867 RMASKHIIRDPISAFNVSADGKFLACGTPSGDIVVVNSTNMQIHTMIKKAHLGIVTALAF 1046
Query: 64 QKE 66
+
Sbjct: 1047SPD 1055
>TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (51%)
Length = 1024
Score = 30.0 bits (66), Expect = 1.5
Identities = 24/83 (28%), Positives = 41/83 (48%), Gaps = 2/83 (2%)
Frame = +2
Query: 162 QEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE--AAAALASVD 219
+EA L+ + K+++++ E E++ +L AEA M +LKAN A E ALA D
Sbjct: 377 REAGELKMDRQKKKLQIEEL---ERIVRLKNAEADMFQLKANEAKREAEQLQRIALAKSD 547
Query: 220 AQQQRLTFQRLVAMMVSDRQKKE 242
++ T L + +K+
Sbjct: 548 KSEEEYTSNYLKQKLSEAEAEKQ 616
>TC218690 similar to UP|O81812 (O81812) Auxilin-like protein, partial (48%)
Length = 1043
Score = 29.3 bits (64), Expect = 2.5
Identities = 24/78 (30%), Positives = 37/78 (46%), Gaps = 3/78 (3%)
Frame = +3
Query: 148 EDARHLAQRYSRMRQ-EAETLREEISKRQVRVRESPTSEQVAKLH--AAEARMKELKANM 204
E + LA+ + R+ E E L E + R+ R R +++ A L AAEAR K +
Sbjct: 9 EQLKKLAEAKEKKREREKEKLAVERAIREARERAFADAKERATLERAAAEARQKNISDGR 188
Query: 205 AVLGKEAAAALASVDAQQ 222
LGK + A A++
Sbjct: 189 ERLGKTTSQANEKTPAEK 242
>TC207715 weakly similar to UP|Q9LHY8 (Q9LHY8) ESTs D15336(C0474), partial
(57%)
Length = 975
Score = 28.9 bits (63), Expect = 3.3
Identities = 38/137 (27%), Positives = 57/137 (40%), Gaps = 7/137 (5%)
Frame = +3
Query: 181 SPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDRQ- 239
SPT++ + AE M + A +LGK+A + D + Q LT + DR+
Sbjct: 183 SPTNQSPDAVKKAEDVMSTMLAKGFILGKDAINKAKAFDERHQ-LTSNASSTVASIDRKI 359
Query: 240 ---KKESAPPVVVSENGSGKTM---YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTG 293
K S +V NG + M Y L+E T +E++ S S G I+ +G
Sbjct: 360 GLSDKLSIGTTIV--NGKVREMDERYQLSEMTKSAMAAAEQKAS-SAGSAIMSNPYVKSG 530
Query: 294 WSEGECNGKAGWFPSAY 310
A WF SA+
Sbjct: 531 ---------ASWFSSAF 554
>TC226382 similar to UP|GS11_ARATH (Q9LMP7) Golgi SNARE 11 protein (AtGOS11)
(Golgi SNAP receptor complex member 1-1), complete
Length = 1040
Score = 28.9 bits (63), Expect = 3.3
Identities = 20/60 (33%), Positives = 30/60 (49%)
Frame = +3
Query: 2 DALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMR 61
DALRKQA KL Q+ +Q + K S + +D D E I + L++L + M+
Sbjct: 222 DALRKQARKLEAQLDEQMNSYRKLVSNNVSTKADAAESDL-ESWIERLLKQLQQVNTQMQ 398
>AW102033
Length = 420
Score = 28.5 bits (62), Expect = 4.3
Identities = 25/87 (28%), Positives = 41/87 (46%), Gaps = 10/87 (11%)
Frame = +3
Query: 148 EDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEA--RMKELKANMA 205
E+ +RY+ + E + ++ +SK + S +++ A AAEA KE +
Sbjct: 78 EELEAAVKRYASVMTELDAAKQALSKTRQEYDSSLDAKKSAFKLAAEAGDASKENTERAS 257
Query: 206 VLGKEAAAA--------LASVDAQQQR 224
L KE +A LAS+ AQQQ+
Sbjct: 258 ELSKEISAVKESIEQAKLASIVAQQQQ 338
>TC216510 weakly similar to UP|Q6T3R2 (Q6T3R2) NDR1-like protein, partial
(35%)
Length = 1117
Score = 28.5 bits (62), Expect = 4.3
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = -3
Query: 269 YGDSEKELSFSKGDFIVVRKVTP 291
YG EKE S++ DF V+R+V P
Sbjct: 89 YGGYEKEFSYTLVDFFVLREVNP 21
>TC232033
Length = 865
Score = 28.5 bits (62), Expect = 4.3
Identities = 14/61 (22%), Positives = 30/61 (48%)
Frame = +2
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
++ R +L++Q K + AVI S Y ++ + +G +Q+ EKL + + +
Sbjct: 383 LEEKRLHVKELQKQSKKLEDAVINAKKNSSYRQMQLMKLHKGFLQVKDCTEKLKTSEKEL 562
Query: 61 R 61
+
Sbjct: 563 Q 565
>TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), complete
Length = 2039
Score = 28.1 bits (61), Expect = 5.6
Identities = 21/67 (31%), Positives = 35/67 (51%), Gaps = 2/67 (2%)
Frame = +3
Query: 162 QEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKE--AAAALASVD 219
+E L+ E K+++++ E E++ +L AEA M +LKA+ A E ALA D
Sbjct: 1434 REVTELKMERQKKKLQIEEL---EKIVRLKNAEADMFQLKADEAKREAERLQMIALAKQD 1604
Query: 220 AQQQRLT 226
++ T
Sbjct: 1605 KSEEEFT 1625
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.127 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,712,143
Number of Sequences: 63676
Number of extensions: 96900
Number of successful extensions: 434
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 425
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 430
length of query: 327
length of database: 12,639,632
effective HSP length: 98
effective length of query: 229
effective length of database: 6,399,384
effective search space: 1465458936
effective search space used: 1465458936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0046a.5


