

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0044a.6
(617 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
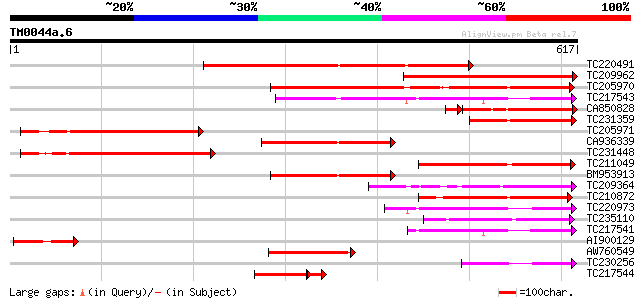
Score E
Sequences producing significant alignments: (bits) Value
TC220491 459 e-129
TC209962 345 4e-95
TC205970 291 8e-79
TC217543 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, parti... 245 4e-65
CA850828 weakly similar to GP|14209545|dbj P0481E12.26 {Oryza sa... 224 4e-62
TC231359 199 3e-51
TC205971 187 9e-48
CA936339 151 9e-37
TC231448 150 1e-36
TC211049 similar to UP|Q9LQV3 (Q9LQV3) F10B6.18, partial (19%) 145 4e-35
BM953913 135 4e-32
TC209364 133 3e-31
TC210872 130 2e-30
TC220973 128 7e-30
TC235110 similar to PIR|C86410|C86410 protein F3M18.18 [imported... 125 6e-29
TC217541 124 9e-29
AI900129 109 4e-24
AW760549 92 5e-19
TC230256 82 7e-16
TC217544 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, parti... 63 2e-14
>TC220491
Length = 875
Score = 459 bits (1182), Expect = e-129
Identities = 232/295 (78%), Positives = 257/295 (86%), Gaps = 2/295 (0%)
Frame = +2
Query: 212 RHLEDLGDFLFSDVRSPPLLQRKAA-DGKQKVPEVFNRVMQSNTMSFTSISETSSKDGLT 270
RHLE+LGDFLFSD++SP QR + +GKQKVPEVFNRVMQS+T F SISE SSKDGLT
Sbjct: 2 RHLENLGDFLFSDLKSPSQQQRNSTPEGKQKVPEVFNRVMQSSTTQFASISEASSKDGLT 181
Query: 271 IICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYK 330
IICSKRGGD+ K SHS+WLQTV SNPEAILFKFVPISSLL GIPGSGYLSHAINLYLRYK
Sbjct: 182 IICSKRGGDVFKQSHSNWLQTVASNPEAILFKFVPISSLLTGIPGSGYLSHAINLYLRYK 361
Query: 331 PPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQ 390
PP +DLQ FLEFQIPRQWAPMF +LPLR HQR+K SS SLQF F+ PKLHVS QV+SEQ
Sbjct: 362 PPPDDLQCFLEFQIPRQWAPMFYDLPLR-HQRKKCSSPSLQFGFMFPKLHVSCAQVVSEQ 538
Query: 391 KPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTPSMWRGSDDNESSNQFLEPV 450
KPVVGLRLYLEGRK DRLA+H+HHLSSLPN MI SS TS+ WRGSDDNESS+ FLEP+
Sbjct: 539 KPVVGLRLYLEGRKSDRLAIHVHHLSSLPNTMIYSSGTSS---WRGSDDNESSDIFLEPI 709
Query: 451 RWKRFSNVCTAVVKHDPNWL-NSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNI 504
RWK F+NVCTAVVKHDP WL +S GVYIVTGAQL+S+ +WPKNVLHLRLL+T+I
Sbjct: 710 RWKGFANVCTAVVKHDPTWLQETSGGVYIVTGAQLISKGTWPKNVLHLRLLYTHI 874
>TC209962
Length = 659
Score = 345 bits (884), Expect = 4e-95
Identities = 164/189 (86%), Positives = 175/189 (91%)
Frame = +3
Query: 429 STPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRC 488
STPSMWRGSDDNES +QFLE +RWKRFSNVCTAVVKHDPNWLN+S GVYIVTGAQL+S+
Sbjct: 6 STPSMWRGSDDNESRDQFLERIRWKRFSNVCTAVVKHDPNWLNNS*GVYIVTGAQLLSKG 185
Query: 489 SWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQ 548
SWP+NVLHLRLLF + PNCSIRK+EW AAPEASRKSSFLTNLSTTFSFTQ T GPPKQ
Sbjct: 186 SWPRNVLHLRLLFAHTPNCSIRKSEWTAAPEASRKSSFLTNLSTTFSFTQHGNT-GPPKQ 362
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
AP +LNSG+YPDGPPVPVRAGKLLKYVE AEVVRGPHD PGHWLVTAAKLVT+GGKIGLQ
Sbjct: 363 APTVLNSGVYPDGPPVPVRAGKLLKYVETAEVVRGPHDAPGHWLVTAAKLVTDGGKIGLQ 542
Query: 609 VKFALLDYW 617
VKFALLDYW
Sbjct: 543 VKFALLDYW 569
>TC205970
Length = 1317
Score = 291 bits (744), Expect = 8e-79
Identities = 160/334 (47%), Positives = 222/334 (65%), Gaps = 3/334 (0%)
Frame = +3
Query: 284 SHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQ 343
SHS W QTV S P+ I FVPI+SLL GI GSGYL+HA+NLYLRYKP E+L FLEFQ
Sbjct: 24 SHSEWCQTVLSQPDVISMSFVPITSLLGGINGSGYLTHAMNLYLRYKPQIEELHQFLEFQ 203
Query: 344 IPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGR 403
+PRQWAP+F EL L +R+ ++ SLQF+F+GPKL+V++T V +KPV GLRLYLEG+
Sbjct: 204 LPRQWAPVFGELAL-GPERKPQNTASLQFSFMGPKLYVNTTPVDVGKKPVTGLRLYLEGK 380
Query: 404 KCDRLALHIHHLSSLPNKMILSSETSTPSMWRGSDDNESS-NQFLEPVRWKRFSNVCTAV 462
+ + LA+H+ HLSSLP L E + G+ N+SS ++ E V+WK FS+VCTA
Sbjct: 381 RSNCLAIHLQHLSSLPKTFQLQDEPN------GNASNDSSERKYYEKVQWKSFSHVCTAP 542
Query: 463 VKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCS-IRKTEWAAAPEAS 521
V D + N+ +VTGA + K VL LRL F + + ++ EW +P +
Sbjct: 543 VDSDDD--NA-----VVTGAHFEVGDTGLKKVLFLRLHFCKVVGATRVKVPEWEGSPGLT 701
Query: 522 RKSSFLTNL-STTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEV 580
+KS ++ L STTFS Q+ PP+ + +NS +YP GPPVP ++ KLL++V+ E+
Sbjct: 702 QKSGIISTLISTTFSGPQKPP---PPRPSDVNINSALYPGGPPVPTQSPKLLRFVDTTEM 872
Query: 581 VRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALL 614
RGP D+PG+W+V+ A+L+ E KI L+VK++LL
Sbjct: 873 TRGPQDSPGYWVVSGARLLVEKAKISLKVKYSLL 974
>TC217543 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, partial (29%)
Length = 1093
Score = 245 bits (626), Expect = 4e-65
Identities = 142/333 (42%), Positives = 194/333 (57%), Gaps = 6/333 (1%)
Frame = +1
Query: 290 QTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWA 349
+TV P+ I F PI SLL G+PG YL+ AI+LYL YKPP EDLQYFL+FQI R WA
Sbjct: 4 ETVKLAPDIINMNFTPIVSLLEGVPGVKYLARAIDLYLEYKPPIEDLQYFLDFQITRVWA 183
Query: 350 PMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLA 409
P L QR++ SLQF+ +GPKL VS QV +KPV GLRL LEG K +RLA
Sbjct: 184 PEQNNL-----QRKEPVCQSLQFSLMGPKLFVSPDQVTVGRKPVTGLRLSLEGSKQNRLA 348
Query: 410 LHIHHLSSLPNKMILSSETST---PSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHD 466
+H+ HL SLP + + W G ++ +S ++ EP++WK FS+V TA +++
Sbjct: 349 IHLQHLVSLPKNLQPHWDAHMAIGAPKWHGPEEQDS--RWFEPIKWKNFSHVSTAPIEYT 522
Query: 467 PNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEW---AAAPEASRK 523
+ +GV+IVTGAQL KNVLHL+LLF+ +P C+IR++ W + P A R
Sbjct: 523 ETSIGDLSGVHIVTGAQLGVWDFGAKNVLHLKLLFSKVPGCTIRRSVWDHNPSTPAAQRS 702
Query: 524 SSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRG 583
++L+ S ++ +S + GKL K V+ E+ +G
Sbjct: 703 DGASSSLTKKTSEDKKEDSS----------------------IHIGKLAKIVDMTEMSKG 816
Query: 584 PHDTPGHWLVTAAKLVTEGGKIGLQVKFALLDY 616
P D PGHWLVT AKL E GKI L++K++LL+Y
Sbjct: 817 PQDIPGHWLVTGAKLGVEKGKIVLRIKYSLLNY 915
>CA850828 weakly similar to GP|14209545|dbj P0481E12.26 {Oryza sativa
(japonica cultivar-group)}, partial (8%)
Length = 584
Score = 224 bits (572), Expect(2) = 4e-62
Identities = 110/125 (88%), Positives = 115/125 (92%)
Frame = +2
Query: 493 NVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAM 552
NVLHLRLLFT+IPNCSIRK+EW AAPEASRKS FLTNLSTTFSFTQ T+GPPKQAP
Sbjct: 68 NVLHLRLLFTHIPNCSIRKSEWTAAPEASRKS-FLTNLSTTFSFTQHG-TTGPPKQAPTA 241
Query: 553 LNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFA 612
LNSG+YPDGPPVPVRAGKLLKYVE AEVVRGPHD PGHWLVTAAKLVT+GGKIGLQ KFA
Sbjct: 242 LNSGVYPDGPPVPVRAGKLLKYVETAEVVRGPHDAPGHWLVTAAKLVTDGGKIGLQXKFA 421
Query: 613 LLDYW 617
LLDYW
Sbjct: 422 LLDYW 436
Score = 32.7 bits (73), Expect(2) = 4e-62
Identities = 13/18 (72%), Positives = 16/18 (88%)
Frame = +1
Query: 475 GVYIVTGAQLVSRCSWPK 492
GVYIVTGAQL+ + SWP+
Sbjct: 13 GVYIVTGAQLLXKGSWPR 66
>TC231359
Length = 683
Score = 199 bits (506), Expect = 3e-51
Identities = 95/116 (81%), Positives = 105/116 (89%)
Frame = +2
Query: 501 FTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPD 560
+T+IPNCSIRK+EW APEASRKSSFLTNLSTTFSFTQQS P KQAP +L+SG+YPD
Sbjct: 2 YTHIPNCSIRKSEWGGAPEASRKSSFLTNLSTTFSFTQQSP---PQKQAPTVLDSGVYPD 172
Query: 561 GPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLDY 616
GPPVPVR+GK+LKYV+ +E VRGPHD PGHWLVTAAKLVTEGGKIGLQVKFALLDY
Sbjct: 173 GPPVPVRSGKMLKYVDTSEFVRGPHDAPGHWLVTAAKLVTEGGKIGLQVKFALLDY 340
>TC205971
Length = 673
Score = 187 bits (476), Expect = 9e-48
Identities = 90/200 (45%), Positives = 136/200 (68%)
Frame = +1
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ ++G G+DL +D +L+F K + RL+ +D+ N R + +P ++I V
Sbjct: 109 AIRAIGLGYDLTNDLKLKFCK----------NHSRLIAIDDDNLRTVELPPR--ISIPNV 252
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
++I+CDKGD +R SDVL F QMSE NQ ++ GKIP+G+FN F +G W +DAA+T
Sbjct: 253 PKSIKCDKGDRMRLCSDVLSFQQMSEQFNQDLSLSGKIPTGHFNTAFGFTGVWQKDAANT 432
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LAFDG I+LY + + +VL + VK++VP+ WDPAAL+RFI+ YGTH+IVG+ +GG
Sbjct: 433 KTLAFDGVSITLYDIAFEKTQVVLHDHVKQAVPSSWDPAALTRFIEKYGTHVIVGVKMGG 612
Query: 192 QDVICVKQKHSSKIPPGDLR 211
D+I KQ++SS +PP +++
Sbjct: 613 TDIIYAKQQYSSTVPPAEVQ 672
>CA936339
Length = 431
Score = 151 bits (381), Expect = 9e-37
Identities = 75/145 (51%), Positives = 100/145 (68%)
Frame = +1
Query: 275 KRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAE 334
+RGGD L+ HS WL+TV S+P+ I F PI+ L+ +PG L+HAI LYL YKPP E
Sbjct: 1 RRGGDDLEQDHSMWLRTVWSSPDVIQMTFCPITDLIDEVPGKEQLTHAIGLYLEYKPPIE 180
Query: 335 DLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVV 394
+L+YFLEFQI WAP+ +P QR++ SLQF+ +G KL+VS Q+ + PV
Sbjct: 181 ELRYFLEFQIAHVWAPLHERIP--GQQRKEPICPSLQFSIMGQKLYVSQEQITVGRLPVT 354
Query: 395 GLRLYLEGRKCDRLALHIHHLSSLP 419
GLRL+LEG K +RL++H+ HLSSLP
Sbjct: 355 GLRLFLEGSKQNRLSVHLQHLSSLP 429
>TC231448
Length = 813
Score = 150 bits (380), Expect = 1e-36
Identities = 82/213 (38%), Positives = 131/213 (61%)
Frame = +3
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ S+G GFD+ D K +GS RL+ ++E+ R + IPG GV+I V
Sbjct: 114 AINSIGLGFDITQDISFDNCK-----KGS-----RLIFVNEEQCRHLEIPG--GVSIPNV 257
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
+I+C +G+ +RF+SDVL +QM E NQ+ + G + SG+ A F LS +D A
Sbjct: 258 PNSIKCVRGESIRFESDVLTRDQMMEHFNQQMLLSGNLASGHLCASFGLSDRSIKDLASI 437
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA+DG+FI Y + L + ++V+++VP+ WDP AL+RFIQ +GTH+IVG+++GG
Sbjct: 438 KSLAYDGWFIKRYTIELERHHCKILDQVEEAVPSSWDPEALARFIQRFGTHVIVGVSMGG 617
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSD 224
+DV+ ++Q+ +S + P +++ L+D F D
Sbjct: 618 KDVLYLRQEDTSYLGPTSIQKLLKDTASRKFKD 716
>TC211049 similar to UP|Q9LQV3 (Q9LQV3) F10B6.18, partial (19%)
Length = 668
Score = 145 bits (367), Expect = 4e-35
Identities = 71/172 (41%), Positives = 111/172 (64%), Gaps = 1/172 (0%)
Frame = +1
Query: 445 QFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNI 504
+F E + K+F +VCTA VK+DP W + +IVTGAQL + ++VLHLRLLF+ +
Sbjct: 7 RFFEAINGKKFYHVCTAPVKYDPRWSSDKDVAFIVTGAQLHVKKHESRSVLHLRLLFSKV 186
Query: 505 PNCSIRKTEWAAAPEA-SRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPP 563
NC++ K+ W S++S + +ST+ S Q+ K+ +L+S ++P GPP
Sbjct: 187 SNCAVVKSSWTQGSSGLSQRSGIFSVISTSISGKDQNQ-----KKPVVLLDSSVFPTGPP 351
Query: 564 VPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLD 615
VPV+ KLLK+++ +++ +GP D+PGHWL+T A+LV + KI L KF+LL+
Sbjct: 352 VPVQTQKLLKFIDTSQLCKGPQDSPGHWLITGARLVLDKRKICLWAKFSLLN 507
>BM953913
Length = 421
Score = 135 bits (341), Expect = 4e-32
Identities = 68/136 (50%), Positives = 92/136 (67%)
Frame = +2
Query: 284 SHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQ 343
+HS WL T+ S+P+ I F PI+ LL IP +L+ AI LYL YKPP E+L+YFLEFQ
Sbjct: 2 NHSKWLSTIKSSPDIIEMTFCPITDLLDEIPAKEHLTRAIGLYLEYKPPIEELRYFLEFQ 181
Query: 344 IPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGR 403
IP WAP+ ++P QR++ SLQF+ +G KL+VS Q+ ++PV GL L LEG
Sbjct: 182 IPCVWAPLQDKIP--GQQRKEPVCPSLQFSIMGQKLYVSQEQITVGRRPVTGLHLCLEGS 355
Query: 404 KCDRLALHIHHLSSLP 419
K +RL++H+ HL SLP
Sbjct: 356 KQNRLSVHVQHLVSLP 403
>TC209364
Length = 725
Score = 133 bits (334), Expect = 3e-31
Identities = 86/227 (37%), Positives = 123/227 (53%), Gaps = 1/227 (0%)
Frame = +2
Query: 391 KPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTPSMWRGSDDNESSNQFLEPV 450
+PVVGLRL LEGR +RLA+H+ HL+SLP + +S ++ S D+ + N + V
Sbjct: 29 RPVVGLRLQLEGRSSNRLAIHLQHLASLPKSLSVSDNSNAYL----SCDSYNCNLH-KKV 193
Query: 451 RWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIR 510
+W S VCTA V+ D + V IVTGAQL L LRL F+ + ++R
Sbjct: 194 KWNSLSYVCTAPVESDDS-------VSIVTGAQLQVE----NKCLFLRLCFSKVIGVTLR 340
Query: 511 KT-EWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAG 569
K EW +P + S + TTF ++ PK + S IY PVR
Sbjct: 341 KAPEWDQSPSLGQFSIKGGGILTTF--ISKAEQRDHPKPGDVTIGSSIYSAARLAPVRTP 514
Query: 570 KLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALLDY 616
KLL++V+ E++RGP DTPGHW+V+ A+L + KI L VK++L +
Sbjct: 515 KLLRFVDTTEIMRGPVDTPGHWVVSGARLHVQDAKIYLHVKYSLFSF 655
>TC210872
Length = 484
Score = 130 bits (326), Expect = 2e-30
Identities = 72/170 (42%), Positives = 105/170 (61%), Gaps = 2/170 (1%)
Frame = +3
Query: 445 QFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNI 504
+F E V+WK FS+VCTA V+ S + IVTGAQL KN+L LRL F+ +
Sbjct: 3 RFYEKVQWKNFSHVCTAPVE-------SEEDLSIVTGAQLQVENYGIKNILFLRLRFSTV 161
Query: 505 PNC-SIRKTEWAAAPEASRKSSFLTNL-STTFSFTQQSATSGPPKQAPAMLNSGIYPDGP 562
+++ EW + + KS ++ L S F+ T Q PP+ A +NS +YP GP
Sbjct: 162 LGAKAVKHPEWEGSLKLGAKSGLISTLISQHFTSTFQKP---PPRPADVNINSAVYPGGP 332
Query: 563 PVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFA 612
PVPV+A KLLK+V+ E+ RGP ++PG+W+++ AKLV + GKI L+VK++
Sbjct: 333 PVPVQAPKLLKFVDTTEMTRGPQESPGYWVISGAKLVVDKGKISLRVKYS 482
>TC220973
Length = 986
Score = 128 bits (322), Expect = 7e-30
Identities = 74/213 (34%), Positives = 120/213 (55%), Gaps = 4/213 (1%)
Frame = +2
Query: 408 LALHIHHLSSLPNKMILSSETSTP---SMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVK 464
L++H+ HL SLP + ++ W+G ++ +S ++ EPV+WK FS+V TA ++
Sbjct: 2 LSVHVQHLVSLPKILHPYWDSHVAIGAPKWQGPEEQDS--RWFEPVKWKNFSHVSTAPIE 175
Query: 465 HDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKS 524
+ ++ +GVYIVTGAQL +NVL+++LL++ +P C+IR++ W P K+
Sbjct: 176 NPETFIGDFSGVYIVTGAQLGVWDFGSRNVLYMKLLYSRLPGCTIRRSLWDHVPNKPPKT 355
Query: 525 SFLTNLSTTFSFT-QQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRG 583
N S + T +++AT A KL+KYV+ +++ +G
Sbjct: 356 VNAENTSNPDNSTLRENAT-------------------------ANKLVKYVDLSKMTKG 460
Query: 584 PHDTPGHWLVTAAKLVTEGGKIGLQVKFALLDY 616
P D PGHWLVT KL E GK+ L+VK++LL+Y
Sbjct: 461 PQDPPGHWLVTGGKLGVEKGKVVLRVKYSLLNY 559
>TC235110 similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(17%)
Length = 754
Score = 125 bits (314), Expect = 6e-29
Identities = 72/165 (43%), Positives = 99/165 (59%), Gaps = 1/165 (0%)
Frame = +3
Query: 451 RWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIR 510
+W FS+V TA V++ + ++ ST IVT A + K VL LRL F+ + + +IR
Sbjct: 3 KWSMFSHVYTAPVQYSSSSMDESTA--IVTKAWFEVKLVGMKKVLFLRLGFSTVASATIR 176
Query: 511 KTEWAAAPEASRKSSFLTNL-STTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPVPVRAG 569
++EW SRKS F + L ST S QS P K +NS IY GPPVP R
Sbjct: 177 RSEWDGPSTTSRKSGFFSALMSTRLSKELQSPDQKPNK---VDINSAIYNVGPPVPTRVP 347
Query: 570 KLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKFALL 614
K+L +V+ E+VRGP DTPG+W+VT AKL EGG+I ++ K++LL
Sbjct: 348 KMLSFVDTKEMVRGPEDTPGYWVVTGAKLCVEGGRISIKAKYSLL 482
>TC217541
Length = 746
Score = 124 bits (312), Expect = 9e-29
Identities = 70/186 (37%), Positives = 105/186 (55%), Gaps = 3/186 (1%)
Frame = +3
Query: 434 WRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSWPKN 493
W G ++ +S ++ EP++WK FS+V TA +++ + +GV+IVTGAQL KN
Sbjct: 51 WHGPEEQDS--RWFEPIKWKNFSHVSTAPIEYTETSIGDLSGVHIVTGAQLGVWDFGAKN 224
Query: 494 VLHLRLLFTNIPNCSIRKTEW---AAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAP 550
VLHL+LLF+ +P C+IR++ W +AP A R ++L S ++ +S
Sbjct: 225 VLHLKLLFSKVPGCTIRRSVWDHNPSAPVAQRPDGASSSLMKKTSEDKKEDSS------- 383
Query: 551 AMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVK 610
+ GKL K V+ E+ +GP D PGHWLVT AKL E GKI L++K
Sbjct: 384 ---------------IHIGKLAKIVDMTEMSKGPQDIPGHWLVTGAKLGVEKGKIVLRIK 518
Query: 611 FALLDY 616
++LL+Y
Sbjct: 519 YSLLNY 536
>AI900129
Length = 431
Score = 109 bits (272), Expect = 4e-24
Identities = 58/71 (81%), Positives = 61/71 (85%)
Frame = +2
Query: 5 EEGIEMVALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAG 64
E+G+EMVALE LGKGFDLASDFRLRFAKGIRE +RLVVLDEQNKRDILIPG G
Sbjct: 242 EKGVEMVALECLGKGFDLASDFRLRFAKGIRE--------ERLVVLDEQNKRDILIPGTG 397
Query: 65 GVTIKGVSENI 75
GVTIKGVSENI
Sbjct: 398 GVTIKGVSENI 430
>AW760549
Length = 296
Score = 92.4 bits (228), Expect = 5e-19
Identities = 44/95 (46%), Positives = 62/95 (64%)
Frame = +3
Query: 282 KHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLE 341
K HS WL T+ S P+ I +P++SL +G++SHAINLY RYKPP EDL FLE
Sbjct: 15 KMYHSEWLDTIDSEPDVISMLLLPLTSLWNRSGRNGFVSHAINLYHRYKPPIEDLHQFLE 194
Query: 342 FQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNFLG 376
FQ+PR WAP+ E+ L H + + ++ ++F+ LG
Sbjct: 195 FQLPRHWAPVASEISLGSHHKHQVNTW-IRFSILG 296
>TC230256
Length = 639
Score = 82.0 bits (201), Expect = 7e-16
Identities = 44/125 (35%), Positives = 67/125 (53%)
Frame = +1
Query: 492 KNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPA 551
+NVL+++LL++ +P C+IR++ W P K+ N S + T + +G
Sbjct: 10 RNVLYMKLLYSRLPGCTIRRSLWDHIPNKPPKTVNAGNTSNLDNSTLKENATG------- 168
Query: 552 MLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQVKF 611
KL+KYV+ +E+ +GP D PGHWLVT KL E GKI L+VK+
Sbjct: 169 -----------------NKLVKYVDLSEMTKGPQDPPGHWLVTGGKLGVEKGKIVLRVKY 297
Query: 612 ALLDY 616
+LL+Y
Sbjct: 298 SLLNY 312
>TC217544 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, partial (30%)
Length = 520
Score = 63.2 bits (152), Expect(2) = 2e-14
Identities = 29/62 (46%), Positives = 42/62 (66%)
Frame = +2
Query: 267 DGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSGYLSHAINLY 326
+ +T+I +RGGD L+ SH+ W +TV P+ I F PI SLL G+PG +L+ AI+LY
Sbjct: 287 EDVTVIFRRRGGDDLEQSHTKWAETVKLAPDIINMNFTPIVSLLEGVPGIKHLARAIDLY 466
Query: 327 LR 328
L+
Sbjct: 467 LQ 472
Score = 35.8 bits (81), Expect = 0.058
Identities = 23/69 (33%), Positives = 38/69 (54%)
Frame = +2
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
++++LG+GFD+ SD RL + KG + RLV LDE + ++ +P + + I V
Sbjct: 71 SIQALGRGFDVTSDIRLLYCKG--------APGSRLVHLDEDHTKN--LPLSHDLVIPNV 220
Query: 72 SENIRCDKG 80
S +I G
Sbjct: 221 SVDIDWSPG 247
Score = 34.7 bits (78), Expect(2) = 2e-14
Identities = 14/20 (70%), Positives = 17/20 (85%)
Frame = +3
Query: 325 LYLRYKPPAEDLQYFLEFQI 344
+Y+ KPP EDLQYFL+FQI
Sbjct: 459 IYICNKPPIEDLQYFLDFQI 518
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,405,713
Number of Sequences: 63676
Number of extensions: 466048
Number of successful extensions: 2413
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 2367
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2382
length of query: 617
length of database: 12,639,632
effective HSP length: 103
effective length of query: 514
effective length of database: 6,081,004
effective search space: 3125636056
effective search space used: 3125636056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0044a.6


