

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
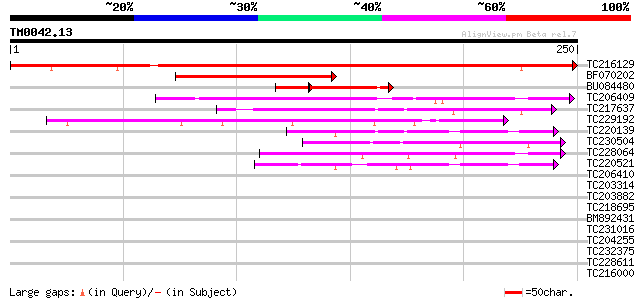
Score E
Sequences producing significant alignments: (bits) Value
TC216129 400 e-112
BF070202 81 5e-16
BU084480 similar to GP|8671768|gb|A T10O22.14 {Arabidopsis thali... 62 2e-13
TC206409 similar to UP|Q6Z4E9 (Q6Z4E9) Immunophilin / FKBP-type ... 69 2e-12
TC217637 similar to GB|CAD35362.1|21535744|ATH490171 FK506 bindi... 60 9e-10
TC229192 similar to UP|Q6ZLL2 (Q6ZLL2) Immunophilin/FKBP-type pe... 54 7e-08
TC220139 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl... 46 2e-05
TC230504 similar to UP|Q9SCY3 (Q9SCY3) FKBP like protein , part... 45 4e-05
TC228064 similar to UP|FKB1_ARATH (Q9LM71) Probable FKBP-type pe... 44 7e-05
TC220521 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl... 42 3e-04
TC206410 similar to UP|Q6Z4E9 (Q6Z4E9) Immunophilin / FKBP-type ... 40 0.001
TC203314 homologue to UP|FKB2_VICFA (Q41649) FK506-binding prote... 38 0.004
TC203882 similar to UP|FKB2_VICFA (Q41649) FK506-binding protein... 38 0.005
TC218695 similar to UP|Q9LDC0 (Q9LDC0) FKBP-like protein (Arabid... 38 0.005
BM892431 similar to GP|20453225|gb| At5g26752 {Arabidopsis thali... 37 0.006
TC231016 37 0.006
TC204255 PDB|1OD5_A.0|33357661|1OD5_A Chain A, Crystal Structure... 37 0.008
TC232375 similar to UP|Q9FJL3 (Q9FJL3) Peptidylprolyl isomerase,... 37 0.008
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 37 0.011
TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light rec... 36 0.014
>TC216129
Length = 1001
Score = 400 bits (1029), Expect = e-112
Identities = 210/262 (80%), Positives = 226/262 (86%), Gaps = 12/262 (4%)
Frame = +3
Query: 1 MATFFGSPPVFSLPLTR-NSHHIASSSPTPPPIPPQQSQSPTASSSQ---------SVQG 50
MA FFGSPP+FSLP T +HHI+SSS TPPP PP QSQ PT+S Q S+Q
Sbjct: 93 MAAFFGSPPIFSLPPTIIRTHHISSSSQTPPPTPPPQSQPPTSSPQQLRTTNLNDESMQV 272
Query: 51 TAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEV 110
+A+ Q PIKP STKV+STDWIATSLTRRFG+GAGLAWVGFLAFGV+SEQIKTRLEV
Sbjct: 273 CTEAKQQKPIKP---STKVESTDWIATSLTRRFGIGAGLAWVGFLAFGVISEQIKTRLEV 443
Query: 111 SQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTF 170
SQQEANTR+VEK EEV LPNGIRY ELKVGGGASPRPGDLVVIDI GK+ES+GEVFVNTF
Sbjct: 444 SQQEANTRNVEKVEEVVLPNGIRYYELKVGGGASPRPGDLVVIDITGKIESSGEVFVNTF 623
Query: 171 EGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADL--GSGV 228
EGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGF ENGAD G+GV
Sbjct: 624 EGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFGENGADFDSGTGV 803
Query: 229 QIPPLATLEYIVEVDKVSIAPA 250
QIPPLATLEYI+EV+KVSIAPA
Sbjct: 804 QIPPLATLEYILEVEKVSIAPA 869
>BF070202
Length = 217
Score = 80.9 bits (198), Expect = 5e-16
Identities = 42/71 (59%), Positives = 51/71 (71%)
Frame = +2
Query: 74 WIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIR 133
WIA+ LTR FG+ +GLAWVG LAFGV+S+ +KT L VSQ++ TR V E+V L I
Sbjct: 5 WIASFLTRLFGIRSGLAWVGCLAFGVISQHLKTLLGVSQRDVYTRTVVSVEDVVLSYVIT 184
Query: 134 YCELKVGGGAS 144
Y EL VGGGAS
Sbjct: 185 YYELYVGGGAS 217
>BU084480 similar to GP|8671768|gb|A T10O22.14 {Arabidopsis thaliana},
partial (17%)
Length = 421
Score = 62.4 bits (150), Expect(2) = 2e-13
Identities = 31/37 (83%), Positives = 35/37 (93%)
Frame = +2
Query: 133 RYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNT 169
RY ELK+GGGASPRPGDLVVIDIMGK+ES+ EVFV+T
Sbjct: 314 RYYELKLGGGASPRPGDLVVIDIMGKIESS-EVFVDT 421
Score = 30.4 bits (67), Expect(2) = 2e-13
Identities = 13/17 (76%), Positives = 15/17 (87%)
Frame = +3
Query: 118 RDVEKEEEVTLPNGIRY 134
R+VEK EEV LPNGIR+
Sbjct: 168 RNVEKVEEVVLPNGIRF 218
>TC206409 similar to UP|Q6Z4E9 (Q6Z4E9) Immunophilin / FKBP-type
peptidyl-prolyl cis-trans isomerase-like, partial (57%)
Length = 983
Score = 68.9 bits (167), Expect = 2e-12
Identities = 54/193 (27%), Positives = 95/193 (48%), Gaps = 8/193 (4%)
Frame = +1
Query: 65 SSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEE 124
SST + + + SL+ G A L+ + GV + + + S++ + + +
Sbjct: 181 SSTNKIAAEPVTVSLSIE-GRRALLSCLLTTVVGVYACDVAGAVSTSRRALRGAKIPESD 357
Query: 125 EVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSR 184
TLPNG++Y +LKVG GA + G V I + K +S TF ++ + V G
Sbjct: 358 YTTLPNGLKYYDLKVGNGAEAKKGSRVAIHYVAKWKSI------TFMTSRQGMG-VGGGT 516
Query: 185 PY-------SKG-VCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATL 236
PY +G V +G++ ++ M+ GG+R +IVPP+L + G +IPP +T+
Sbjct: 517 PYGFDVGQSERGTVLKGLDLGVQGMRVGGQRLLIVPPELAYGSKGVQ-----EIPPNSTI 681
Query: 237 EYIVEVDKVSIAP 249
E +E+ + +P
Sbjct: 682 ELDIELLSIKQSP 720
>TC217637 similar to GB|CAD35362.1|21535744|ATH490171 FK506 binding protein 1
{Arabidopsis thaliana;} , partial (60%)
Length = 870
Score = 60.1 bits (144), Expect = 9e-10
Identities = 50/153 (32%), Positives = 73/153 (47%), Gaps = 3/153 (1%)
Frame = +1
Query: 92 VGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
+G LAF V L S +A E P+G+ +C+ VG G G L+
Sbjct: 214 IGLLAFDAV-------LAYSSLQAAPAAENPCEFQVAPSGLAFCDKLVGAGPQAVKGQLI 372
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGI--EYVLKTMKAGGKRKV 209
+G++E+ G+VF +++ K PL +G KG EGI + M AGGKR +
Sbjct: 373 KAHYVGRLEN-GKVFDSSYNRGK-PLTFRVGVGEVIKGWDEGIIGGDGVPPMLAGGKRTL 546
Query: 210 IVPPQLGFRENGADL-GSGVQIPPLATLEYIVE 241
+PP+LG+ GA G IPP + L + VE
Sbjct: 547 KIPPELGYGSRGAGCRGGSCIIPPDSVLLFDVE 645
>TC229192 similar to UP|Q6ZLL2 (Q6ZLL2) Immunophilin/FKBP-type
peptidyl-prolyl cis-trans isomerase-like protein,
partial (65%)
Length = 1206
Score = 53.9 bits (128), Expect = 7e-08
Identities = 63/226 (27%), Positives = 94/226 (40%), Gaps = 22/226 (9%)
Frame = +1
Query: 17 RNSHHIAS----SSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDST 72
RN I S S+P P P P + + +S S + K+Q +P K SS+ S
Sbjct: 19 RNERRILSPPLTSNPNPNPNSPVFAMASISSLSLTSVSLPKSQSLDPKKISDSSSSSGSR 198
Query: 73 DW---IATSLTRRFGLGAGLAWV-GFLAFGVVSEQIKTRLEVSQQEANTRDVEKE----- 123
A S RR L + A + G L VS +D K
Sbjct: 199 SQSCCCAPSFQRRKMLLSSAAIIAGTLCSNSVSGVSLAAEFADMPALRGKDYGKTKMRYP 378
Query: 124 EEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKV--------ESTGEVFVNTFEGDKK 175
+ +G++Y +L+ G G P+ G+ VV+D G E+ + +FEGD K
Sbjct: 379 DYTETESGLQYKDLRPGNGPKPKMGETVVVDWDGYTIGYYGRIFEARNKTKGGSFEGDDK 558
Query: 176 PL-ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFREN 220
+G Y++ V E + M GG R++IVPP+LG+ EN
Sbjct: 559 DFFKFKIG---YNE-VIPAFEEAVSGMALGGIRRIIVPPELGYPEN 684
>TC220139 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl isomerase
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) , partial (29%)
Length = 722
Score = 45.8 bits (107), Expect = 2e-05
Identities = 41/121 (33%), Positives = 58/121 (47%), Gaps = 1/121 (0%)
Frame = +1
Query: 123 EEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVM 181
EE G++ LK G G +P GD V + G + G F ++ + D P + +
Sbjct: 166 EEREIGSRGLKKKLLKEGQGWETPEVGDEVQVHYTGTLLD-GTKFDSSRDRDS-PFSFTL 339
Query: 182 GSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVE 241
G KG EGI KTMK G +PP+L + E+ GS IPP ATL++ VE
Sbjct: 340 GQGQVIKGWDEGI----KTMKKGENAIFTIPPELAYGES----GSPPTIPPNATLQFDVE 495
Query: 242 V 242
+
Sbjct: 496 L 498
>TC230504 similar to UP|Q9SCY3 (Q9SCY3) FKBP like protein , partial (68%)
Length = 988
Score = 44.7 bits (104), Expect = 4e-05
Identities = 35/119 (29%), Positives = 60/119 (50%), Gaps = 3/119 (2%)
Frame = +1
Query: 130 NGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKG 189
+G+ YC++ G G G+L+ + + + G VF ++++ +PL + +G KG
Sbjct: 265 SGLGYCDIAEGFGDEAPLGELINVHYTARF-ADGIVFDSSYKR-ARPLTMRIGVGKVIKG 438
Query: 190 VCEGIEYV--LKTMKAGGKRKVIVPPQLGFRENGADLGSG-VQIPPLATLEYIVEVDKV 245
+ +GI M+ GGKR++ +PP L + A SG IP ATL Y + +V
Sbjct: 439 LDQGILGAEGXPPMRIGGKRQLQIPPHLAYGPEPAGCFSGDCNIPANATLLYDINFVEV 615
>TC228064 similar to UP|FKB1_ARATH (Q9LM71) Probable FKBP-type
peptidyl-prolyl cis-trans isomerase 1, chloroplast
precursor (PPIase) (Rotamase) , partial (66%)
Length = 1043
Score = 43.9 bits (102), Expect = 7e-05
Identities = 36/160 (22%), Positives = 71/160 (43%), Gaps = 25/160 (15%)
Frame = +3
Query: 111 SQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVID---IMGKVESTGEVFV 167
++ N + + +++ +T P+G++Y +L G G G V + + + +
Sbjct: 270 AEARRNKKAIPEDQYITSPDGLKYYDLVEGKGPVAEKGTTVQVHFDCLYRGITAVSSRES 449
Query: 168 NTFEGDK---KPLALVMGSRPYSKGVCEGIE-------------------YVLKTMKAGG 205
G++ +P +G+ P + E ++ +++ M+ GG
Sbjct: 450 KLLAGNRIIAQPYEFKVGAPPGKERKREFVDNPNGLFSAQAAPKPPPAVYIIVEGMRVGG 629
Query: 206 KRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
KR VIVPP+ G+ + G + +IPP AT E VE+ +V
Sbjct: 630 KRTVIVPPENGYGQKGMN-----EIPPGATFELNVELLQV 734
>TC220521 similar to UP|FKB7_WHEAT (Q43207) 70 kDa peptidylprolyl isomerase
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) , partial (40%)
Length = 890
Score = 42.0 bits (97), Expect = 3e-04
Identities = 45/139 (32%), Positives = 65/139 (46%), Gaps = 5/139 (3%)
Frame = +1
Query: 109 EVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFV 167
E + E+ V +E+E+ G++ LK G G +P GD V +V TG +
Sbjct: 199 EEEEFESPILKVGEEKEIG-KMGLKKKLLKEGEGWDTPDSGDQV------EVHYTGTLLD 357
Query: 168 NT-FEGDKK---PLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGAD 223
T F+ + P +G KG EGI KTMK G +PP+L + E+
Sbjct: 358 GTKFDSSRDRGTPFKFKLGQGQVIKGWDEGI----KTMKKGENALFTIPPELAYGES--- 516
Query: 224 LGSGVQIPPLATLEYIVEV 242
GS IPP ATL++ VE+
Sbjct: 517 -GSPPTIPPNATLQFDVEL 570
>TC206410 similar to UP|Q6Z4E9 (Q6Z4E9) Immunophilin / FKBP-type
peptidyl-prolyl cis-trans isomerase-like, partial (36%)
Length = 539
Score = 40.0 bits (92), Expect = 0.001
Identities = 19/60 (31%), Positives = 36/60 (59%)
Frame = +2
Query: 190 VCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
V +G++ ++ M+ GG+R +IVPP+L + G +IPP +T+E +E+ + +P
Sbjct: 197 VLKGLDLGVQGMRVGGQRLLIVPPELAYGSKGVQ-----EIPPNSTIELDIELLSIKQSP 361
>TC203314 homologue to UP|FKB2_VICFA (Q41649) FK506-binding protein 2
precursor (Peptidyl-prolyl cis-trans isomerase)
(PPIase) (Rotamase) (15 kDa FKBP) (FKBP-15) , partial
(85%)
Length = 746
Score = 38.1 bits (87), Expect = 0.004
Identities = 33/95 (34%), Positives = 49/95 (50%)
Frame = +2
Query: 148 GDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKR 207
GD V + GK+ + G VF ++FE + P+ +G+ KG +G L M G KR
Sbjct: 197 GDRVKVHYRGKL-TDGTVFDSSFERNN-PIEFELGTGQVIKGWDQG----LLGMCLGEKR 358
Query: 208 KVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
K+ +P +LG+ D GS IP ATL + E+
Sbjct: 359 KLKIPAKLGY----GDQGSPPTIPGGATLIFDTEL 451
>TC203882 similar to UP|FKB2_VICFA (Q41649) FK506-binding protein 2 precursor
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) (15 kDa FKBP) (FKBP-15) , partial (85%)
Length = 944
Score = 37.7 bits (86), Expect = 0.005
Identities = 33/95 (34%), Positives = 49/95 (50%)
Frame = +2
Query: 148 GDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKR 207
GD V + GK+ + G VF ++FE + P+ +G+ KG +G L M G KR
Sbjct: 260 GDRVKVHYRGKL-TDGTVFDSSFERNN-PIEFELGTGQVIKGWDQG----LLGMCLGEKR 421
Query: 208 KVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
K+ +P +LG+ E GS IP ATL + E+
Sbjct: 422 KLKIPSKLGYGEQ----GSPPTIPGGATLIFDAEL 514
>TC218695 similar to UP|Q9LDC0 (Q9LDC0) FKBP-like protein (Arabidopsis
thaliana genomic DNA, chromosome 3, P1 clone: MIL23),
partial (79%)
Length = 1562
Score = 37.7 bits (86), Expect = 0.005
Identities = 36/128 (28%), Positives = 59/128 (45%)
Frame = +1
Query: 115 ANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDK 174
+N V+ E EV L + +K G G P E + F +T++ ++
Sbjct: 286 SNPPKVDSEVEV-LHEKVTKQIIKEGHGQKPSKYSTCFFHYRAWAEKSQHKFEDTWQ-EQ 459
Query: 175 KPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLA 234
+P+ +V+G K G+ + +MKAG + V V +LG+ E G+ S +PP A
Sbjct: 460 RPIEMVLGKE---KKEMTGLGIGVASMKAGERALVRVGWELGYGEEGS--FSFPNVPPKA 624
Query: 235 TLEYIVEV 242
L Y VE+
Sbjct: 625 DLVYEVEL 648
>BM892431 similar to GP|20453225|gb| At5g26752 {Arabidopsis thaliana},
partial (9%)
Length = 421
Score = 37.4 bits (85), Expect = 0.006
Identities = 26/82 (31%), Positives = 40/82 (48%), Gaps = 7/82 (8%)
Frame = +2
Query: 7 SPPVFSLPLTRNS--HHIASSSP-TPPPIPPQQSQSPT----ASSSQSVQGTAKAQPQNP 59
S P + PL R S H + SS P +P PIPP ++ P+ A+S+ S P NP
Sbjct: 80 SEPTNATPLGRASSPHRLLSSPPPSPRPIPPSSAKKPSKDLAATSTNSTTPPIPTPPPNP 259
Query: 60 IKPVTSSTKVDSTDWIATSLTR 81
P + + + ++SL+R
Sbjct: 260 FTPTSLTFPSSTCPRASSSLSR 325
>TC231016
Length = 851
Score = 37.4 bits (85), Expect = 0.006
Identities = 14/29 (48%), Positives = 22/29 (75%)
Frame = +3
Query: 188 KGVCEGIEYVLKTMKAGGKRKVIVPPQLG 216
KG+ G E ++ M+ GGKR++I+PP+LG
Sbjct: 288 KGLVPGFEEGIRDMRPGGKRRIIIPPELG 374
>TC204255 PDB|1OD5_A.0|33357661|1OD5_A Chain A, Crystal Structure Of Glycinin
A3b4 Subunit Homohexamer. {Glycine max;} , partial (6%)
Length = 519
Score = 37.0 bits (84), Expect = 0.008
Identities = 23/57 (40%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Frame = +1
Query: 8 PPVFSLPLTRNSHHIA--SSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKP 62
P VF P T S + S SP+PP +PP + SP AS+S S A +P +P P
Sbjct: 58 PSVFLTPPTSPSTPLTAPSPSPSPPSLPPAPAGSPGASTS-SAMACAAPKPSSPSPP 225
Score = 28.9 bits (63), Expect = 2.3
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 11/67 (16%)
Frame = +1
Query: 25 SSPTPPPIPPQQSQS------PTASSSQ-----SVQGTAKAQPQNPIKPVTSSTKVDSTD 73
SSP+PPP P S S PT S++ S +GT++ P+NP P S+ S
Sbjct: 208 SSPSPPPPTPPPSPSAATTSPPTKPSTKSKPLPSTRGTSR--PKNPTAP*PSTAPSRSPI 381
Query: 74 WIATSLT 80
TS T
Sbjct: 382 QQGTSAT 402
>TC232375 similar to UP|Q9FJL3 (Q9FJL3) Peptidylprolyl isomerase, partial
(21%)
Length = 559
Score = 37.0 bits (84), Expect = 0.008
Identities = 39/125 (31%), Positives = 57/125 (45%), Gaps = 5/125 (4%)
Frame = +3
Query: 123 EEEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFVNT-FEGDKK---PL 177
EE+ G++ LK G G +P GD V +V TG + T F+ + P
Sbjct: 213 EEKEIGKEGLKKKLLKEGEGWDTPEAGDEV------QVHYTGTLLDGTKFDSSRDRGTPF 374
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLE 237
+ +G KG +GI TMK G +P +L + E+G S IPP ATL+
Sbjct: 375 SFTLGQGQVIKGWDQGII----TMKKGENALFTIPAELAYGESG----SPPTIPPNATLQ 530
Query: 238 YIVEV 242
+ VE+
Sbjct: 531 FDVEL 545
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 36.6 bits (83), Expect = 0.011
Identities = 21/58 (36%), Positives = 29/58 (49%)
Frame = +3
Query: 23 ASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLT 80
AS SP PPP PP+ + S T +SS + + P P P S+T ++ AT T
Sbjct: 156 ASRSPAPPPPPPRSASSSTTASSPTAVTPPSSSPPRP-SPGPSATPTSTSPSPATPST 326
Score = 32.3 bits (72), Expect = 0.21
Identities = 24/84 (28%), Positives = 30/84 (35%), Gaps = 6/84 (7%)
Frame = +3
Query: 3 TFFGSPPVFSLPLTRNSHHIASSS------PTPPPIPPQQSQSPTASSSQSVQGTAKAQP 56
TF PP S P + +SS+ P+PPP PP P S A P
Sbjct: 387 TFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYPPTSPPPPCPSLKSPNSSPNAASP 566
Query: 57 QNPIKPVTSSTKVDSTDWIATSLT 80
P T DS ++S T
Sbjct: 567 SKNSSPSVGPTPSDSLTAPSSSRT 638
Score = 28.1 bits (61), Expect = 3.9
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 7/45 (15%)
Frame = +3
Query: 22 IASSSPTPPPI-----PPQQSQSP--TASSSQSVQGTAKAQPQNP 59
++S +PTP P PP + SP A SS S A A+P +P
Sbjct: 351 LSSPAPTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSP 485
>TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light receptor
(Photolyase/blue light photoreceptor PHR2), partial
(84%)
Length = 1641
Score = 36.2 bits (82), Expect = 0.014
Identities = 21/63 (33%), Positives = 30/63 (47%), Gaps = 2/63 (3%)
Frame = +3
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPI--PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVT 64
SPP S PL+ + ++ PTPPP PP + T++SS +P P T
Sbjct: 177 SPPTPSTPLSPSLPTAPATLPTPPPSAAPPSSGSATTSASSTMSASPPPTTTPSPSSPST 356
Query: 65 SST 67
+ST
Sbjct: 357 AST 365
Score = 31.2 bits (69), Expect = 0.46
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +3
Query: 7 SPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSS 45
SP S P T S H AS+ P P P + SPT++++
Sbjct: 345 SPSTASTPPTTASRHPASTRPAPTAPPSSSTPSPTSAAA 461
Score = 26.9 bits (58), Expect = 8.8
Identities = 19/71 (26%), Positives = 34/71 (47%)
Frame = +3
Query: 2 ATFFGSPPVFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIK 61
AT PP + P + ++ ASS+ + P PP + SP++ S+ S T ++ +
Sbjct: 228 ATLPTPPPSAAPPSSGSATTSASSTMSASP-PPTTTPSPSSPSTASTPPTTASRHPASTR 404
Query: 62 PVTSSTKVDST 72
P ++ ST
Sbjct: 405 PAPTAPPSSST 437
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,502,230
Number of Sequences: 63676
Number of extensions: 175518
Number of successful extensions: 5240
Number of sequences better than 10.0: 561
Number of HSP's better than 10.0 without gapping: 3466
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4596
length of query: 250
length of database: 12,639,632
effective HSP length: 95
effective length of query: 155
effective length of database: 6,590,412
effective search space: 1021513860
effective search space used: 1021513860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0042.13


