

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.8
(562 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
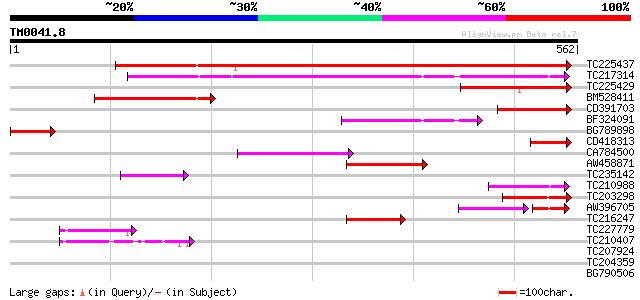
Score E
Sequences producing significant alignments: (bits) Value
TC225437 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein (Faceless... 779 0.0
TC217314 weakly similar to UP|O80938 (O80938) CER1-like protein,... 295 3e-80
TC225429 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein (Faceless... 202 4e-52
BM528411 199 3e-51
CD391703 136 2e-32
BF324091 weakly similar to GP|1418319|emb| CER1-like {Arabidopsi... 89 7e-18
BG789898 weakly similar to PIR|T02691|T026 glossy1 protein gl1 -... 84 2e-16
CD418313 83 3e-16
CA784500 weakly similar to GP|1199467|dbj possible aldehyde deca... 71 1e-12
AW458871 65 1e-10
TC235142 similar to UP|O22681 (O22681) Arabidopsis thaliana gl1 ... 62 5e-10
TC210988 similar to UP|O80938 (O80938) CER1-like protein, partia... 59 4e-09
TC203298 UP|CB23_SOYBN (P09756) Chlorophyll a-b binding protein ... 57 2e-08
AW396705 similar to GP|1209703|gb|A maize gl1 homolog {Arabidops... 39 1e-07
TC216247 similar to UP|O04077 (O04077) Sucrose transport protein... 54 2e-07
TC227779 similar to UP|Q8VWZ8 (Q8VWZ8) Sterol 4-alpha-methyl-oxi... 50 3e-06
TC210407 similar to UP|O49656 (O49656) Predicted protein, partia... 46 5e-05
TC207924 similar to UP|Q8VYI1 (Q8VYI1) At1g69640/F24J1.22, parti... 36 0.040
TC204359 similar to UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine deca... 33 0.34
BG790506 similar to GP|8809631|dbj| HSR203J protein-like protein... 30 2.2
>TC225437 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein (Faceless pollen-1)
(YORE-YORE protein), partial (70%)
Length = 1577
Score = 779 bits (2011), Expect = 0.0
Identities = 375/454 (82%), Positives = 412/454 (90%), Gaps = 2/454 (0%)
Frame = +3
Query: 106 HATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKT 165
+AT++EH ++V+IG PILGAS+MGYGSASL+YGYVL FDFLRCLGHCNVEVVPH LF+
Sbjct: 3 NATLLEHLIMTVIIGTPILGASLMGYGSASLIYGYVLIFDFLRCLGHCNVEVVPHQLFEK 182
Query: 166 FPFMKYLLYTPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCS--R 223
PF++Y++YTPTYH LHH+D KDTNFCLFMPLFD+LG+TLN SW+ H+ SSG +
Sbjct: 183 LPFLRYVIYTPTYHHLHHSD-KDTNFCLFMPLFDSLGNTLNKNSWQSHKLLSSGSGNGDM 359
Query: 224 VPDFVFLAHIVDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLF 283
VP FVFLAHIVDV+SSMHAQF RSFASLP+ TRFFL+P PI LLAMW SK FL
Sbjct: 360 VPHFVFLAHIVDVSSSMHAQFVYRSFASLPYTTRFFLLPGLPITFLVLLAMWAWSKTFLV 539
Query: 284 SFYYLRDRLHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEA 343
SFYYLR RLHQTWVVPRCGFQYFLPFA EGIN +IE+AIL ADKIGVKVISLAALNKNE+
Sbjct: 540 SFYYLRGRLHQTWVVPRCGFQYFLPFATEGINNQIEQAILRADKIGVKVISLAALNKNES 719
Query: 344 LNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQ 403
LNGGGKLFVDKHPNLRVRVVHGNTLTAAVI++EIP+DV+EVFLTGATSKLGRAIALYLCQ
Sbjct: 720 LNGGGKLFVDKHPNLRVRVVHGNTLTAAVILNEIPQDVKEVFLTGATSKLGRAIALYLCQ 899
Query: 404 KKVKVLMLTLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAP 463
KKVKVLMLTLS DRF+RIQKEAP EYQSYLVQVTKYQAAQ+CKTWIVGKWITPREQ WAP
Sbjct: 900 KKVKVLMLTLSTDRFQRIQKEAPPEYQSYLVQVTKYQAAQNCKTWIVGKWITPREQYWAP 1079
Query: 464 RGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHS 523
RGTHFHQFVVPPILPFR+DCTYG+LAAMRLPEDVEGLG CEYTM+RGVVHACHAGGVVHS
Sbjct: 1080RGTHFHQFVVPPILPFRKDCTYGDLAAMRLPEDVEGLGCCEYTMDRGVVHACHAGGVVHS 1259
Query: 524 LEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSS 557
LEGW HHEVGAIDV+RID+VW+AALKHGLRPVSS
Sbjct: 1260LEGWPHHEVGAIDVNRIDLVWEAALKHGLRPVSS 1361
>TC217314 weakly similar to UP|O80938 (O80938) CER1-like protein, partial
(50%)
Length = 1544
Score = 295 bits (756), Expect = 3e-80
Identities = 167/439 (38%), Positives = 240/439 (54%)
Frame = +1
Query: 117 VVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTP 176
++ IP L S + GYV DF+ +GHCN EVVP LF FP +KYL+YT
Sbjct: 1 LLFAIPKLTLVFTKTASVGAMLGYVTYIDFMNNMGHCNFEVVPKWLFDIFPPLKYLMYTS 180
Query: 177 TYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDV 236
++HSLHHT + TN+ LFMPL+D + T + S +LHES + +P+ V L H+
Sbjct: 181 SFHSLHHTQFR-TNYSLFMPLYDYIYGTTDKASDKLHESALKQE-EEIPNVVHLTHLTTP 354
Query: 237 TSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTW 296
S H + AS P+ ++++L WP+ +++ WV + F+ QTW
Sbjct: 355 ESIYHLRLGFAYLASKPYTSKWYLCLMWPVTAWSMILTWVYGRTFIVEGNRFDKLKLQTW 534
Query: 297 VVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHP 356
+P+ QYF+ IN IEEAIL AD+ G+KV+SL N+ E LN G L+V +HP
Sbjct: 535 AIPKYSLQYFMQSQKVAINTMIEEAILDADRKGIKVLSLGLRNQGEDLNIYGGLYVSRHP 714
Query: 357 NLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSAD 416
L+VRVV G++L AV+++ IPK +V L G +K+ A+A LCQ+ V+V L D
Sbjct: 715 KLKVRVVDGSSLVVAVVLNSIPKGTTQVLLRGKLTKIAYALAYTLCQQGVQV--AALYED 888
Query: 417 RFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPI 476
+ R++K + + Q TW+VG +T EQ AP+GT F + P
Sbjct: 889 DYVRLKKSFNSSETNLAFTKSSTQT-----TWLVGDGLTEEEQLKAPKGTLFIPYTQFPP 1053
Query: 477 LPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAID 536
+R+DC Y AM P VE + SCE + R ++ A G+VHSLEGWT HE G
Sbjct: 1054RKYRKDCFYHCTPAMLAPCSVENIHSCEDWLPRRIMSAWRIAGIVHSLEGWTEHECGH-T 1230
Query: 537 VDRIDVVWKAALKHGLRPV 555
+ ID VW + L+HG +P+
Sbjct: 1231MHNIDNVWHSTLQHGFQPL 1287
>TC225429 similar to UP|Q8H1Z0 (Q8H1Z0) Cuticle protein (Faceless pollen-1)
(YORE-YORE protein), partial (18%)
Length = 729
Score = 202 bits (513), Expect = 4e-52
Identities = 100/157 (63%), Positives = 107/157 (67%), Gaps = 47/157 (29%)
Frame = +1
Query: 448 WIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCE--- 504
WIVGKWITPREQ WAPRGTHFHQFVVPPIL FR+DCTYG+LAAMRLPEDVEGLG CE
Sbjct: 16 WIVGKWITPREQYWAPRGTHFHQFVVPPILSFRKDCTYGDLAAMRLPEDVEGLGCCEVKP 195
Query: 505 --------------------------------------------YTMERGVVHACHAGGV 520
YTM+RGVVHACHAGGV
Sbjct: 196 SSTLAVSVSFIFLLIDMI*FFSHEPITYFYLLTLINICLIRYMQYTMDRGVVHACHAGGV 375
Query: 521 VHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSS 557
VHSLEGW+HHEVGAIDV+RID+VW+AALKHGLRPVSS
Sbjct: 376 VHSLEGWSHHEVGAIDVNRIDLVWEAALKHGLRPVSS 486
>BM528411
Length = 368
Score = 199 bits (506), Expect = 3e-51
Identities = 89/120 (74%), Positives = 108/120 (89%)
Frame = +1
Query: 85 LFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGF 144
LFT+YHSLHHSSPVPES T G+AT++EH ++V+IGIPILGAS+MGYGSAS++YGYVL F
Sbjct: 1 LFTHYHSLHHSSPVPESFTAGNATLLEHLIMTVIIGIPILGASLMGYGSASMIYGYVLIF 180
Query: 145 DFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKDTNFCLFMPLFDALGHT 204
DFL CLGH NVE+VPH LF+ PF++Y++YTPTYH LHH+D KDTNFCLFMPLFD+LG+T
Sbjct: 181 DFLICLGHSNVEIVPHQLFEKLPFLRYVIYTPTYHHLHHSD-KDTNFCLFMPLFDSLGNT 357
>CD391703
Length = 358
Score = 136 bits (343), Expect = 2e-32
Identities = 61/74 (82%), Positives = 69/74 (92%)
Frame = -3
Query: 484 TYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVV 543
T + ++RLPEDVEGLG CEYTM+RGVVHACHAGGVVHSLEGW+HHEVGAIDV+RID+V
Sbjct: 311 TMSNVQSLRLPEDVEGLGCCEYTMDRGVVHACHAGGVVHSLEGWSHHEVGAIDVNRIDLV 132
Query: 544 WKAALKHGLRPVSS 557
W+AALKHGLRPVSS
Sbjct: 131 WEAALKHGLRPVSS 90
>BF324091 weakly similar to GP|1418319|emb| CER1-like {Arabidopsis thaliana},
partial (9%)
Length = 418
Score = 88.6 bits (218), Expect = 7e-18
Identities = 47/139 (33%), Positives = 80/139 (56%)
Frame = +3
Query: 330 VKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGA 389
VKV+SL N+ ++ N G+L++ ++P L++++V G++L A++++ IPK+ +V L G
Sbjct: 12 VKVLSLGLSNQGDSFNKYGELYIKRYPELKIKIVDGSSLVVAIVVNSIPKEARQVLLCGK 191
Query: 390 TSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWI 449
+K+ AIA LC++ KV T+ D + ++Q E + LV Y A K W+
Sbjct: 192 PNKVSYAIASALCERGTKV--TTMYKDEYDKLQLRISNESKKNLVFPGSYTA----KIWL 353
Query: 450 VGKWITPREQSWAPRGTHF 468
VG EQ AP+G+ F
Sbjct: 354 VGDQCDEVEQKKAPKGSLF 410
>BG789898 weakly similar to PIR|T02691|T026 glossy1 protein gl1 - maize,
partial (16%)
Length = 162
Score = 84.0 bits (206), Expect = 2e-16
Identities = 37/45 (82%), Positives = 41/45 (90%)
Frame = +2
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLP 45
M FLTRNRRI+Q+GVDFKQIDKEWDWDNFLILQ ++ASMA YM P
Sbjct: 26 MFFLTRNRRIVQKGVDFKQIDKEWDWDNFLILQALVASMACYMFP 160
>CD418313
Length = 670
Score = 83.2 bits (204), Expect = 3e-16
Identities = 37/41 (90%), Positives = 41/41 (99%)
Frame = -1
Query: 517 AGGVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPVSS 557
AGGVVHSLEGW+HHEVGAIDV+RID+VW+AALKHGLRPVSS
Sbjct: 388 AGGVVHSLEGWSHHEVGAIDVNRIDLVWEAALKHGLRPVSS 266
>CA784500 weakly similar to GP|1199467|dbj possible aldehyde decarbonylase
{Arabidopsis thaliana}, partial (18%)
Length = 396
Score = 70.9 bits (172), Expect = 1e-12
Identities = 39/115 (33%), Positives = 58/115 (49%)
Frame = +2
Query: 226 DFVFLAHIVDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSF 285
D V L H+ S H + S AS P + ++L WP + ++L W + F+
Sbjct: 32 DVVHLTHLTTPESIYHLRLGFASLASRPQSSTWYLYLMWPFTLWSVLVTWFYGQTFVMER 211
Query: 286 YYLRDRLHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNK 340
+ Q+WV+PR QY + +E +NK IEEAIL A+ VKV+SL N+
Sbjct: 212 NAFKMLNLQSWVIPRFHVQYLFKWQSETLNKLIEEAILQAELSKVKVLSLGLSNQ 376
>AW458871
Length = 411
Score = 64.7 bits (156), Expect = 1e-10
Identities = 26/80 (32%), Positives = 52/80 (64%)
Frame = +1
Query: 335 LAALNKNEALNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLG 394
L N+ ++ N G+L++ ++P L++++V G++L A++++ IPK+ +V L G +K+
Sbjct: 1 LGLSNQGDSFNKYGELYIKRYPELKIKIVDGSSLVVAIVVNSIPKEARQVLLCGKPNKVS 180
Query: 395 RAIALYLCQKKVKVLMLTLS 414
AIA LC++ KV ++ S
Sbjct: 181 YAIASALCERGTKVCLILFS 240
>TC235142 similar to UP|O22681 (O22681) Arabidopsis thaliana gl1 homolog,
partial (12%)
Length = 336
Score = 62.4 bits (150), Expect = 5e-10
Identities = 28/67 (41%), Positives = 38/67 (55%)
Frame = +3
Query: 111 EHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMK 170
EH V+ IP+ V S + GY+ DF+ LGHCN E +P +F FPF+K
Sbjct: 48 EHIAYFVLFAIPLYTTVVARTASIASYAGYLAYIDFMNNLGHCNFECIPKAIFTAFPFLK 227
Query: 171 YLLYTPT 177
YL+YTP+
Sbjct: 228 YLMYTPS 248
>TC210988 similar to UP|O80938 (O80938) CER1-like protein, partial (11%)
Length = 449
Score = 59.3 bits (142), Expect = 4e-09
Identities = 28/81 (34%), Positives = 46/81 (56%)
Frame = +1
Query: 475 PILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGA 534
P R+DC Y AM P + + SCE + R V+ A G++H+LE W +E G
Sbjct: 43 PPKKLRKDCFYHSTPAMIAPPSLVNVDSCENWLPRRVMSAWRVAGILHALECWKVNECGN 222
Query: 535 IDVDRIDVVWKAALKHGLRPV 555
+ + ++ +W+A+L+HG RP+
Sbjct: 223 V-MFSVEKIWQASLQHGFRPL 282
>TC203298 UP|CB23_SOYBN (P09756) Chlorophyll a-b binding protein 3,
chloroplast precursor (LHCII type I CAB-3) (LHCP),
complete
Length = 1518
Score = 57.0 bits (136), Expect = 2e-08
Identities = 28/69 (40%), Positives = 43/69 (61%)
Frame = -1
Query: 489 AAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVGAIDVDRIDVVWKAAL 548
AAM +P VE + SCE + R V+ A G+VHSLE W+ +E + ID VW++ L
Sbjct: 459 AAMLIPSCVENVHSCEDWLPRRVMSAWRIAGIVHSLERWSTNECN-YKMHNIDKVWRSTL 283
Query: 549 KHGLRPVSS 557
+HG +P+++
Sbjct: 282 QHGFQPLTT 256
>AW396705 similar to GP|1209703|gb|A maize gl1 homolog {Arabidopsis
thaliana}, partial (11%)
Length = 422
Score = 39.3 bits (90), Expect(2) = 1e-07
Identities = 22/69 (31%), Positives = 32/69 (45%)
Frame = +1
Query: 446 KTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEY 505
K W++G +Q AP+G+ F P R+DC Y AM P + + SCE
Sbjct: 34 KIWLLGDQCNEVDQRKAPKGSLFIPISQFPPKKLRKDCFYHSTPAMIAPPSLVNVDSCEN 213
Query: 506 TMERGVVHA 514
+ R V+ A
Sbjct: 214 WLPRRVMSA 240
Score = 35.0 bits (79), Expect(2) = 1e-07
Identities = 14/37 (37%), Positives = 27/37 (72%)
Frame = +3
Query: 519 GVVHSLEGWTHHEVGAIDVDRIDVVWKAALKHGLRPV 555
G++H+LEGW +E G ++ ++ + +A+L+HG RP+
Sbjct: 252 GILHALEGWNVNECGN-EMFSVEKIRQASLQHGFRPL 359
>TC216247 similar to UP|O04077 (O04077) Sucrose transport protein, partial
(17%)
Length = 930
Score = 53.9 bits (128), Expect = 2e-07
Identities = 25/58 (43%), Positives = 40/58 (68%)
Frame = +1
Query: 335 LAALNKNEALNGGGKLFVDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSK 392
L +N+ E LN G L+V ++PNL+V++V G++L AAV+++ IPK +V L G +K
Sbjct: 757 LGLMNQGEDLNIYGGLYVSRNPNLKVKMVDGSSLAAAVVLNNIPKGTTQVLLMGKLTK 930
>TC227779 similar to UP|Q8VWZ8 (Q8VWZ8) Sterol 4-alpha-methyl-oxidase,
partial (97%)
Length = 1131
Score = 49.7 bits (117), Expect = 3e-06
Identities = 25/78 (32%), Positives = 42/78 (53%), Gaps = 2/78 (2%)
Frame = +1
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSW V V++ ++ + + + ++YW HR H +L+ + HS+HH P LT +A
Sbjct: 373 LPSWKV--VLIQIIFYFILEDFIFYWGHRILHTKWLYKHVHSVHHEYATPFGLTSEYAHP 546
Query: 110 VEHAFI--SVVIGIPILG 125
E F+ + + G I G
Sbjct: 547 AEILFLGFATIFGPAITG 600
>TC210407 similar to UP|O49656 (O49656) Predicted protein, partial (33%)
Length = 1010
Score = 45.8 bits (107), Expect = 5e-05
Identities = 36/138 (26%), Positives = 62/138 (44%), Gaps = 4/138 (2%)
Frame = +2
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSW + ++ L+++ + + YW+HR H D+ + H +HH P +G A
Sbjct: 530 LPSW--REILSQLLVYFLVEDYTNYWIHRFLHNDWGYEKIHRVHHEYHAP----IGFAAP 691
Query: 110 VEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFP-- 167
H +++GIP S +G ++V G+++ F L +E + FP
Sbjct: 692 YAHWAEILILGIP----SFLG---PAMVPGHIITFWLWIALR--QIEAIDTHSGYDFPRS 844
Query: 168 FMKYLLY--TPTYHSLHH 183
KY+ + YH HH
Sbjct: 845 ITKYIPFYGGAEYHDYHH 898
>TC207924 similar to UP|Q8VYI1 (Q8VYI1) At1g69640/F24J1.22, partial (83%)
Length = 952
Score = 36.2 bits (82), Expect = 0.040
Identities = 38/158 (24%), Positives = 69/158 (43%), Gaps = 5/158 (3%)
Frame = +1
Query: 31 ILQTIIASMAYYMLPFL-QNLPSWNVKGVIVA--LVLHVGISEPLYYWVHRKFHGD-YLF 86
++Q ++A++ + + QN S N +++A V + I + Y++HR H + +L+
Sbjct: 97 LVQAVVATLLFALTGSDGQNTTSQNTSLLVLARQFVTAMLIMDAWQYFMHRYMHHNKFLY 276
Query: 87 TNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDF 146
+ HSLHH VP S + +E V G S + G + + F
Sbjct: 277 KHIHSLHHRLIVPYSYGALYNHPIEGLLNDTVGG----ALSFLLSGMSPRASIFFFSFAT 444
Query: 147 LRCL-GHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHH 183
++ + HC + ++P LF F YH +HH
Sbjct: 445 IKTVDDHCGL-LLPGNLFHIF-----FKNNSAYHDVHH 540
>TC204359 similar to UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase,
partial (71%)
Length = 835
Score = 33.1 bits (74), Expect = 0.34
Identities = 20/49 (40%), Positives = 24/49 (48%)
Frame = +3
Query: 431 SYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPF 479
SY +Q Y Q C T ++G ITP E S+AP H PI PF
Sbjct: 105 SYRLQTHPY*REQKCVTVVIGLRITPGEVSFAPELIHV------PIAPF 233
>BG790506 similar to GP|8809631|dbj| HSR203J protein-like protein
{Arabidopsis thaliana}, partial (11%)
Length = 421
Score = 30.4 bits (67), Expect = 2.2
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +2
Query: 445 CKTWIVGKWITPREQSWAPRGTHFHQ 470
C T KW TPR SW P T H+
Sbjct: 308 CSTIFASKWRTPRRPSWPPLTTASHR 385
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,676,976
Number of Sequences: 63676
Number of extensions: 523284
Number of successful extensions: 3461
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 3401
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3447
length of query: 562
length of database: 12,639,632
effective HSP length: 102
effective length of query: 460
effective length of database: 6,144,680
effective search space: 2826552800
effective search space used: 2826552800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0041.8


