

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
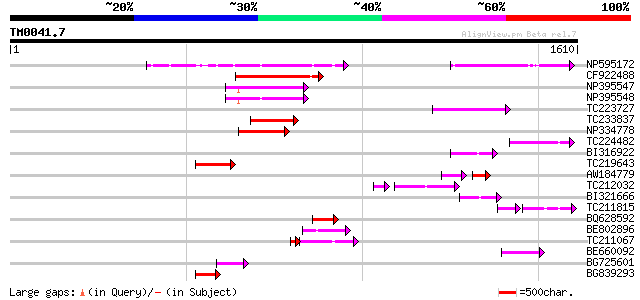
Query= TM0041.7
(1610 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 201 2e-51
CF922488 199 1e-50
NP395547 reverse transcriptase [Glycine max] 153 5e-37
NP395548 reverse transcriptase [Glycine max] 147 4e-35
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 142 2e-33
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 142 2e-33
NP334778 reverse transcriptase [Glycine max] 133 7e-31
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 115 1e-25
BI316922 98 3e-20
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 90 7e-18
AW184779 60 3e-16
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 61 2e-15
BI321666 78 3e-14
TC211815 49 2e-12
BQ628592 65 2e-10
BE802896 65 2e-10
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 54 1e-09
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 62 2e-09
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 59 2e-08
BG839293 58 4e-08
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 201 bits (511), Expect = 2e-51
Identities = 170/583 (29%), Positives = 275/583 (47%), Gaps = 10/583 (1%)
Frame = +1
Query: 388 KILVDGGAAIN-LMPRFMMKRLGKTETDLIPHDMVLSDYEGKTSTSMGAIM---LNITAG 443
KILVDGG++ N + PR + ++ K + P+ VL G+ ++ G + L+I
Sbjct: 1207 KILVDGGSSDNFIQPR--VAQVLKLPVEPAPNLRVLVG-NGQILSAEGIVQQLPLHIQGQ 1377
Query: 444 TVSRSTLFIVVPSKANYNLLLGREWIHGVGA-VPSTLHQRISIWKPDGVVE-----NVQA 497
V + + + +++LG W+ +G V + ++ D + N +A
Sbjct: 1378 EVKVPVYLLQI---SGADVILGSTWLATLGPHVADYAALTLKFFQNDKFITLQGEGNSEA 1548
Query: 498 DQSYYLAEAGYVDKKNFEKSPLVDATKMEAQDPLEKVDLGDGSKKRPTYISSLIDPELRG 557
Q+ + K+ E+ + + E + D K PT I DPEL
Sbjct: 1549 TQAQLHHFRRLQNTKSIEECFAIQLIQKEVPE--------DTLKDLPTNI----DPELA- 1689
Query: 558 RMIELLKEFKDCFA*DNDEMPGLSRDLVELQLPIKEDKKLVK*LPRRFHPDVLVKIKEEI 617
LL + FA P +D +P+K+ VK P R+ +I++ I
Sbjct: 1690 ---ILLHTYAQVFAVPASLPPQREQDHA---IPLKQGSGPVKVRPYRYPHTQKDQIEKMI 1851
Query: 618 ERLLRCKFIRAAKYVEWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDS 677
+ +L I+ + L ++ V KK+G R C D+R LNA T KD + MP + ++D
Sbjct: 1852 QEMLVQGIIQPSNSPFSLP-ILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDE 2028
Query: 678 AAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVM 737
G +Y S LD SGY+QI + ED KTAFR G YEW+VMPFGL NA AT+Q +M
Sbjct: 2029 LHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHH--GHYEWLVMPFGLTNAPATFQCLM 2202
Query: 738 NTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGD 797
N IF + F+ V+ DD+++ S S +HL HL + ++ H L KC+FG D
Sbjct: 2203 NKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVD 2382
Query: 798 FLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSL 857
+LG V G+ + K +A+LD P + KQL+ LG + RRFI + ++ P + L
Sbjct: 2383 YLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDL 2562
Query: 858 LRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQED 917
L ++D F W + AF +LK + PV+ P +P L A+ +G++L Q
Sbjct: 2563 L---QKDSFLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNG 2733
Query: 918 EDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYFSCTKLKYYI 960
+ Y S+ L + + + L + + +K ++Y+
Sbjct: 2734 -----HPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYL 2847
Score = 99.8 bits (247), Expect = 9e-21
Identities = 95/356 (26%), Positives = 150/356 (41%), Gaps = 5/356 (1%)
Frame = +1
Query: 1253 HGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSI- 1311
H G H AG + YWP + +D Y + C CQ+ +PA L +
Sbjct: 3235 HSSPIGGH-AGITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPLP 3411
Query: 1312 IKSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQS-VTQETVIEFIQNH 1370
I + A+D I + + I+V ID K+ IPL++ + V E +H
Sbjct: 3412 IPQQVWEDVAMDFITGLPNSFGLS--VIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSH 3585
Query: 1371 IVYRFGLPESLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLI 1430
IV G+P S+ +D+ +F + G L S Y+ Q++GQ E NK L +
Sbjct: 3586 IVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYL 3765
Query: 1431 KKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQE 1490
+ PK W ++L + Y + + G TPFR YG+E P + Q+ I
Sbjct: 3766 RCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGRE---PPTLTRQACSI---- 3924
Query: 1491 DIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVIL 1550
D P+EV ++L + D LTR + + + +KK + SF GD V +
Sbjct: 3925 DDPAEV-----REQLTDRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQ 4089
Query: 1551 PTDKKD---RAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKYLKAY 1603
P + R K + + GPFKV + + AY ++ S R + LK +
Sbjct: 4090 PYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHPVFHVSQLKPF 4257
>CF922488
Length = 741
Score = 199 bits (505), Expect = 1e-50
Identities = 103/250 (41%), Positives = 159/250 (63%)
Frame = +3
Query: 641 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAK 700
V+K++GK+ +C+D+RDLN A+PKD++ +P ++VD+ S +DG+SGYNQI IA
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 701 EDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKS 760
ED+ KT F GT+ + M FGLKN GATYQR M +F D + ++VY+DD++VKS
Sbjct: 183 EDMEKTTFIT--LWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKS 356
Query: 761 PSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILD 820
+ +EHL +LRK F R+R + L++NP KC F V + L F+ +GIE++ NK K IL+
Sbjct: 357 RTEEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILE 536
Query: 821 TSPPTSKKQLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELK 880
+ P ++KQ+Q LG++N++ RFI L +P + L + +W+ AF+ +K
Sbjct: 537 MAKPHTEKQVQGFLGRLNYIVRFIS*LIATCEPL--FILLCKNQFVKWDHDC*VAFERIK 710
Query: 881 NYLAIPPVMI 890
L P V++
Sbjct: 711 QCLINPHVLV 740
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 153 bits (387), Expect = 5e-37
Identities = 93/254 (36%), Positives = 128/254 (49%), Gaps = 18/254 (7%)
Frame = +1
Query: 613 IKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNG------------------KMRVCIDF 654
+++E+ +LL I W++ V V KK G + R+CID+
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 655 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGAL 714
R LN AT KD Y +P + M+ A + LDGYSGYNQI + +D KTAF C ++
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 715 GTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSF 774
Y MPFGL NA T+QR M IF D +E ++V++DD S L++L K
Sbjct: 361 FAYRR--MPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVL 534
Query: 775 ERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLL 834
+R L +N KC F V G LG + K+GIE+ K K I PP + K + S L
Sbjct: 535 QRCEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFL 714
Query: 835 GKVNFLRRFIDNLS 848
G V F RRFI + +
Sbjct: 715 GHVGFYRRFIKDFT 756
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 147 bits (371), Expect = 4e-35
Identities = 87/254 (34%), Positives = 129/254 (50%), Gaps = 18/254 (7%)
Frame = +1
Query: 613 IKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNG------------------KMRVCIDF 654
+++E+ +LL I W++ V+ V KK G ++CID+
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 655 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAFRCSGAL 714
R LN AT KD + +P + M++ AGH Y LD Y GYNQI + +D K AF C
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCP--F 354
Query: 715 GTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLSHLRKSF 774
G + + +PFGL NA T+Q M IF D +E ++V++DD V PS + L L
Sbjct: 355 GVFAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVL 534
Query: 775 ERMRIHGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLL 834
+R L +N KC F V G LG + +GIE+++ K I PP++ K ++S L
Sbjct: 535 QRCVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFL 714
Query: 835 GKVNFLRRFIDNLS 848
G+ F RRFI + +
Sbjct: 715 GQARFYRRFIKDFT 756
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 142 bits (357), Expect = 2e-33
Identities = 75/223 (33%), Positives = 110/223 (48%)
Frame = +1
Query: 1200 EYLQNPVGSTDIKVKYRALNYTIIGNELFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGA 1259
EYL + ++ A + + G+ L+K+N D L+C+ +A + H G G
Sbjct: 175 EYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGT 354
Query: 1260 HQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFRG 1319
H G M + R G YW T+ DC + + C CQ + + P L+ + WPF
Sbjct: 355 HANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSM 534
Query: 1320 WAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPE 1379
W ID+IG I P S H++I+VAIDYF KWVEA V + V+ FI+ I+ R+GLP
Sbjct: 535 WGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPR 714
Query: 1380 SLTTDQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAA 1422
+ TD GT + + E + I+ PY + N VE A
Sbjct: 715 KIITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 142 bits (357), Expect = 2e-33
Identities = 70/134 (52%), Positives = 94/134 (69%)
Frame = +2
Query: 685 SLLDGYSGYNQIFIAKEDVSKTAFRCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDF 744
S +DG+SGYNQI +A+EDV KT F GT+ + VM FGLKN GATYQR M +FHD
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVT--LWGTFSYRVMAFGLKNTGATYQRAMVALFHDM 178
Query: 745 IETFMQVYIDDLVVKSPSRDEHLSHLRKSFERMRIHGLKMNPLKCAFGVIAGDFLGFVVH 804
+ ++VY+DD++ KS + EHL +L K F R++ + LK+NP KC FGV +G LGF+V
Sbjct: 179 MHKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVS 358
Query: 805 KKGIEINKNKAKAI 818
+KGIEI+ K KA+
Sbjct: 359 QKGIEIDPEKVKAL 400
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 133 bits (334), Expect = 7e-31
Identities = 62/145 (42%), Positives = 96/145 (65%)
Frame = +3
Query: 649 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAKEDVSKTAF 708
R+C+D+RDLN A+PKD + +P ++++ + A S +DG+SGYNQI +A ED+ KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 709 RCSGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDLVVKSPSRDEHLS 768
GT+ + VM FGLKN GATY R M +F D + ++ Y+D+++ KS +EHL
Sbjct: 183 IT--LWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLV 356
Query: 769 HLRKSFERMRIHGLKMNPLKCAFGV 793
+L+ F ++R + L++NP KC FG+
Sbjct: 357 NLQNLFGQLRKYRLRLNPRKCVFGL 431
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 115 bits (289), Expect = 1e-25
Identities = 71/184 (38%), Positives = 98/184 (52%)
Frame = +1
Query: 1420 EAANKILISLIKKHVGRKPKSWHESLSQVLWAYRNSPREATGTTPFRLAYGQEAVLPAEV 1479
EAANK + +I+K + K WHE L L YR S R +TG TPF L YG EAVLP EV
Sbjct: 1 EAANKNIKKIIQK-MTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 1480 YLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLALDVLTRQKDRIAKAYNKKVKNRSF 1539
+ S RI + + + D+L ++ +R+ A+ + R+ A++KKV R F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 1540 VTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKVLSNNAYVIKELSGQRQFVTINDKY 1599
GD V K + K R GKWAPN+EGPF VK+ S A V+ + G+ +N
Sbjct: 358 HEGDLVLKKMSHAVKDHR--GKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDV 531
Query: 1600 LKAY 1603
+K Y
Sbjct: 532 VKRY 543
>BI316922
Length = 405
Score = 97.8 bits (242), Expect = 3e-20
Identities = 49/132 (37%), Positives = 72/132 (54%)
Frame = +3
Query: 1253 HGGLCGAHQAGAKMKWILFRQGMYWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSII 1312
H G+CG H M + R G Y T+ K C EY K C++C K I H+ ELH+I+
Sbjct: 9 HRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHNIV 188
Query: 1313 KSWPFRGWAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIV 1372
WPF +D++ P RQ KY++V ID F KW+E + ++ V +F+ +IV
Sbjct: 189 APWPFAI*GVDILRPF-PLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNIV 365
Query: 1373 YRFGLPESLTTD 1384
FG+P +L +D
Sbjct: 366 C*FGIPNTLISD 401
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 90.1 bits (222), Expect = 7e-18
Identities = 46/114 (40%), Positives = 71/114 (61%)
Frame = +3
Query: 528 QDPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVEL 587
Q+ E VDLG GS KR I + I +R +I LL++++D FA +MPGLS D+V+
Sbjct: 975 QEETELVDLGSGSGKREVKIGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQH 1154
Query: 588 QLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPV 641
+LP+ + VK RR P+ +KIKEE+++ F+ A+Y +W+AN+VP+
Sbjct: 1155 RLPLNPECSPVKQKLRRMKPETSLKIKEEVKK*FDAGFLAVARYPKWVANIVPI 1316
Score = 33.1 bits (74), Expect = 1.0
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = +1
Query: 468 WIHGVGAVPSTLHQRI 483
WIH VG VPSTLHQ++
Sbjct: 4 WIHSVGVVPSTLHQKL 51
>AW184779
Length = 432
Score = 59.7 bits (143), Expect(2) = 3e-16
Identities = 29/53 (54%), Positives = 34/53 (63%), Gaps = 2/53 (3%)
Frame = +3
Query: 1314 SWPFRG--WAIDLIGEIHPALSRQHKYIIVAIDYFIKWVEAIPLQSVTQETVI 1364
SW G W ID+IG I P S H +I+VAIDYF KWVEA+ SVT+ VI
Sbjct: 270 SWQHLGHMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVI 428
Score = 45.4 bits (106), Expect(2) = 3e-16
Identities = 24/71 (33%), Positives = 33/71 (45%)
Frame = +1
Query: 1225 NELFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKMKWILFRQGMYWPTIMKDC 1284
N L+K+N D LL+C+ +A + H G G H M + R G YW T+ DC
Sbjct: 10 NILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDC 189
Query: 1285 IEYAKGCQDCQ 1295
+ C CQ
Sbjct: 190 CIHVWKCHKCQ 222
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 60.8 bits (146), Expect(2) = 2e-15
Identities = 49/191 (25%), Positives = 82/191 (42%), Gaps = 6/191 (3%)
Frame = +2
Query: 1093 KNVEVKGDSELVVKQLTKEYKCISKNLAKYYVKAMSLLANFDQAGVSYIPRVSNQEANEL 1152
K ++V GDS LV+ QL E + NL Y L FD+ ++ NQ A+ L
Sbjct: 233 KLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA*EENQMADAL 412
Query: 1153 AQIASGYMIDKQKLKELIRIKEKLSPLDLEVMVIDNLTPNDWRKPIVEYLQN---PVGST 1209
A + S + + I + + P +V + W I Y+++ P+ ++
Sbjct: 413 ATLVSMFQLTPHGDLPYIEFRCRGRPAHC-CLVEEERDGKPWYFDIKRYVESKEYPLEAS 589
Query: 1210 DIKVKYR---ALNYTIIGNELFKKNVDGTLLKCLSEVDAFIAVSAAHGGLCGAHQAGAKM 1266
D + + A + + G+ L+K+N D LL C++ + + H G G H G M
Sbjct: 590 DNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSFGTHSNGHAM 769
Query: 1267 KWILFRQGMYW 1277
+ R G YW
Sbjct: 770 ARKILRAGYYW 802
Score = 41.6 bits (96), Expect(2) = 2e-15
Identities = 19/47 (40%), Positives = 28/47 (59%)
Frame = +1
Query: 1033 WKLFFDDSSHKEGSGIGMFIVSPQGIPTKFMFRIRESCSNNESEYEA 1079
W ++FD +S+ G G+G +VSP F R+ C+NN +EYEA
Sbjct: 52 WIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEA 192
>BI321666
Length = 430
Score = 78.2 bits (191), Expect = 3e-14
Identities = 41/122 (33%), Positives = 64/122 (51%)
Frame = +2
Query: 1276 YWPTIMKDCIEYAKGCQDCQKHSGIQHVPASELHSIIKSWPFRGWAIDLIGEIHPALSRQ 1335
Y P+I KD +A+ C CQ+ + LH+I++ F W ID +G P+ S
Sbjct: 32 YLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVGPFPPSFS-- 205
Query: 1336 HKYIIVAIDYFIKWVEAIPLQSVTQETVIEFIQNHIVYRFGLPESLTTDQGTMFVGRKVA 1395
++YI+V +DY KWVEA+ Q + VI+F++ I R G+P L + G+ +A
Sbjct: 206 NEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGSHLCNA*LA 385
Query: 1396 AF 1397
F
Sbjct: 386 RF 391
>TC211815
Length = 704
Score = 48.5 bits (114), Expect(2) = 2e-12
Identities = 41/153 (26%), Positives = 69/153 (44%)
Frame = +2
Query: 1456 PREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLA 1515
P+ T TPF L YG +LP EV ++ ++ + LD + + E+ V+
Sbjct: 227 PQSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIM 406
Query: 1516 LDVLTRQKDRIAKAYNKKVKNRSFVTGDYVWKVILPTDKKDRAYGKWAPNWEGPFKVKKV 1575
K R+ + +N K+ + G + + + R+ K+ N EGPFK++
Sbjct: 407 T*AF---KQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVRS--KFTTN*EGPFKIRHN 571
Query: 1576 LSNNAYVIKELSGQRQFVTINDKYLKAYKQMLH 1608
N AY ++ELSG+ N +LK Y +H
Sbjct: 572 SKNGAYKLEELSGKVVLRIWNSMHLKVYYS*VH 670
Score = 43.5 bits (101), Expect(2) = 2e-12
Identities = 22/67 (32%), Positives = 33/67 (48%)
Frame = +3
Query: 1384 DQGTMFVGRKVAAFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHE 1443
D G F RK+ F IK + + Q N + EAANK+++ +KK + W E
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLLDGANGRWVE 182
Query: 1444 SLSQVLW 1450
L ++LW
Sbjct: 183 DLVEILW 203
>BQ628592
Length = 423
Score = 65.5 bits (158), Expect = 2e-10
Identities = 29/76 (38%), Positives = 50/76 (65%), Gaps = 1/76 (1%)
Frame = -1
Query: 860 LKREDIFRWEA*HQKAFDELKNYLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDED 919
L + W + +Q+AF+++K LA P V++PP+ G+P LY++ DE++G +L Q D+
Sbjct: 423 LPKNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
Query: 920 G-IERAVFYLSRVLND 934
G E+A++YLS+ D
Sbjct: 243 GKKEQAIYYLSKKFTD 196
>BE802896
Length = 416
Score = 65.1 bits (157), Expect = 2e-10
Identities = 45/137 (32%), Positives = 69/137 (49%)
Frame = -2
Query: 832 SLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKNYLAIPPVMIP 891
S LG F RRFI + P S+LL+ + E F + + AFD LK L P++
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKC--K*AFDCLKRALITTPIIQA 242
Query: 892 PIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTMIEKLCLCLYF 951
P P L A++ +G +LAQ+ D + R ++Y SR L+ A+ YT EK L + F
Sbjct: 241 PDWTAPFELMCDASNYALGVVLAQK-IDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVF 65
Query: 952 SCTKLKYYIKPIDVMVF 968
+ K Y+ ++V+
Sbjct: 64 ALEKFHSYLLGTRIIVY 14
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 53.5 bits (127), Expect(2) = 1e-09
Identities = 43/169 (25%), Positives = 72/169 (42%), Gaps = 2/169 (1%)
Frame = +1
Query: 824 PTSKK--QLQSLLGKVNFLRRFIDNLSDKTKPFSSLLRLKREDIFRWEA*HQKAFDELKN 881
PT K ++S G +F RRF+ N S P + L+ K+ F W ++AF LK
Sbjct: 97 PTLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELV--KKNMAFTWGEKQEQAFALLKE 270
Query: 882 YLAIPPVMIPPIKGKPMRLYISATDETIGSMLAQEDEDGIERAVFYLSRVLNDAKTRYTM 941
L PV+ P K L A+ + ++L Q + Y S L+ A Y
Sbjct: 271 KLTKAPVLALPDFSKTFELECDASGVGVRAVLLQGG-----HPIAYFSEKLHSATLNYPT 435
Query: 942 IEKLCLCLYFSCTKLKYYIKPIDVMVFSHFDIIKHMLSKPILHSRIGKW 990
+K L + ++++ + ++ S +K++ K L+ R KW
Sbjct: 436 YDKELYALIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKW 582
Score = 29.3 bits (64), Expect(2) = 1e-09
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = +2
Query: 798 FLGFVVHKKGIEINKNKAKAILDTSPP 824
F GFVV + G++++ K KAI + PP
Sbjct: 23 FSGFVVGRNGVQMDPEKIKAIQEWPPP 103
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 62.0 bits (149), Expect = 2e-09
Identities = 33/122 (27%), Positives = 61/122 (49%)
Frame = -3
Query: 1396 AFAESWGIKLLTSIPYYAQANGQVEAANKILISLIKKHVGRKPKSWHESLSQVLWAYRNS 1455
A + +G+ S PY+ Q NGQ E +N+ + +++K V K W L LWA+R +
Sbjct: 370 ALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTA 191
Query: 1456 PREATGTTPFRLAYGQEAVLPAEVYLQSFRIQRQEDIPSEVYWNMMLDELVNLDEERVLA 1515
+ G +P+R+ +G+ LP E+ +++ + + + +L LDE R+ A
Sbjct: 190 YKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRLEA 11
Query: 1516 LD 1517
+
Sbjct: 10 YE 5
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 58.9 bits (141), Expect = 2e-08
Identities = 35/89 (39%), Positives = 47/89 (52%)
Frame = -3
Query: 588 QLPIKEDKKLVK*LPRRFHPDVLVKIKEEIERLLRCKFIRAAKYVEWLANVVPVIKKNGK 647
+L I D KLV R+ + + +E+ +L FIR Y L +VV V K NGK
Sbjct: 274 KLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNGK 95
Query: 648 MRVCIDFRDLNAATPKDEYHMPIAEMMVD 676
R+C D+ DLN A PKD Y +P + M D
Sbjct: 94 WRICTDYIDLN*ACPKDAYPLPNIDHMTD 8
>BG839293
Length = 781
Score = 57.8 bits (138), Expect = 4e-08
Identities = 32/72 (44%), Positives = 44/72 (60%)
Frame = +1
Query: 528 QDPLEKVDLGDGSKKRPTYISSLIDPELRGRMIELLKEFKDCFA*DNDEMPGLSRDLVEL 587
Q+ E VDLG GS KR I + I +R +I LLK+++D FA +MPGLS D+V+
Sbjct: 511 QEETELVDLGIGSGKREVKIGTGITAPIREELIILLKDYQDIFAWSYQDMPGLSSDIVQH 690
Query: 588 QLPIKEDKKLVK 599
QLP+ + VK
Sbjct: 691 QLPLNPECSPVK 726
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,831,379
Number of Sequences: 63676
Number of extensions: 951116
Number of successful extensions: 5668
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 5549
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5652
length of query: 1610
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1500
effective length of database: 5,635,272
effective search space: 8452908000
effective search space used: 8452908000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0041.7


