

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0037a.10
(168 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
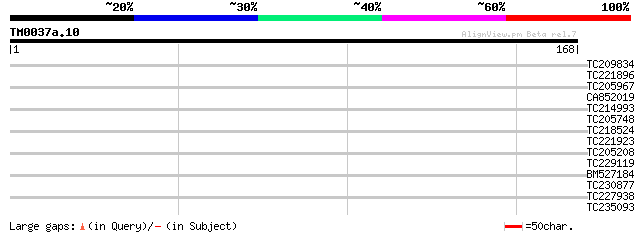
Score E
Sequences producing significant alignments: (bits) Value
TC209834 homologue to UP|Q8MRR8 (Q8MRR8) GH01240p (Fragment), pa... 31 0.24
TC221896 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhib... 28 1.6
TC205967 homologue to UP|NUIM_SOLTU (P80269) NADH-ubiquinone oxi... 28 2.7
CA852019 homologue to GP|12697486|em phosphate transporter {Sesb... 27 3.5
TC214993 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhib... 27 3.5
TC205748 similar to GB|AAR04550.1|37813554|AY337455 dual-specifi... 27 3.5
TC218524 27 4.6
TC221923 similar to UP|Q8GCY1 (Q8GCY1) Dihydrolipoamide succinyl... 27 6.0
TC205208 homologue to UP|Q84P31 (Q84P31) Aldehyde dehydrogenase ... 27 6.0
TC229119 26 7.9
BM527184 similar to PIR|T50692|T50 proline transport protein 1 [... 26 7.9
TC230877 similar to UP|Q9FHE1 (Q9FHE1) Glutathione transferase-l... 26 7.9
TC227938 homologue to UP|Q9LK77 (Q9LK77) Similarity to acyl-CoA ... 26 7.9
TC235093 similar to UP|Q8MZ21 (Q8MZ21) RE13473p, partial (8%) 26 7.9
>TC209834 homologue to UP|Q8MRR8 (Q8MRR8) GH01240p (Fragment), partial (8%)
Length = 540
Score = 31.2 bits (69), Expect = 0.24
Identities = 24/65 (36%), Positives = 29/65 (43%), Gaps = 1/65 (1%)
Frame = -1
Query: 96 TPTTHSHTSFHGECTTGSKSVTEEWIFYPCFFTTRLSPNQVSSSQL-VMNQFCYVLMTNE 154
T T HSH H EE+I C F + PN VSSS L N F +V N
Sbjct: 180 TLTAHSHIENH-----------EEFILLLC*FASFRYPNHVSSSLLSPKNYFFHVKNQNN 34
Query: 155 SVYGI 159
+V+ I
Sbjct: 33 TVFNI 19
>TC221896 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhibitor D
precursor, partial (95%)
Length = 449
Score = 28.5 bits (62), Expect = 1.6
Identities = 23/85 (27%), Positives = 32/85 (37%)
Frame = +1
Query: 64 LKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFY 123
LK S++D Y+ ++C TRS+PP + S H +C +
Sbjct: 85 LKSDQSSSYDDDEYSKPCCDLCMCTRSMPPQCSCEDIRLNSCHSDCKS------------ 228
Query: 124 PCFFTTRLSPNQVSSSQLVMNQFCY 148
TR P Q L N FCY
Sbjct: 229 --CMCTRSQPGQCRC--LDTNDFCY 291
>TC205967 homologue to UP|NUIM_SOLTU (P80269) NADH-ubiquinone oxidoreductase
23 kDa subunit, mitochondrial precursor (Complex
I-23KD) (CI-23KD) (Complex I-28.5KD) (CI-28.5KD) ,
partial (89%)
Length = 1219
Score = 27.7 bits (60), Expect = 2.7
Identities = 21/98 (21%), Positives = 37/98 (37%), Gaps = 19/98 (19%)
Frame = -2
Query: 70 SNFDLFIYNDSIEEVCYLTRS----------LPPSITPTTHSHTSFHGECTTGSKSVTEE 119
S F ++ +++ I C LT+S +P S S + +C + S+T+
Sbjct: 768 SKFKVWSFDNGINRACLLTKSTIDALGHVNIIPSCPAAAIFSLLSLNCDCLCRTYSLTKF 589
Query: 120 ---------WIFYPCFFTTRLSPNQVSSSQLVMNQFCY 148
WI C FTT + ++ FC+
Sbjct: 588 ASNTTFLPCWISPECMFTTETRAQRAFLKWIIDCNFCF 475
>CA852019 homologue to GP|12697486|em phosphate transporter {Sesbania
rostrata}, partial (21%)
Length = 793
Score = 27.3 bits (59), Expect = 3.5
Identities = 15/33 (45%), Positives = 17/33 (51%)
Frame = +3
Query: 84 VCYLTRSLPPSITPTTHSHTSFHGECTTGSKSV 116
V Y+T L S PT HS +SFH T SV
Sbjct: 156 VLYITTQLTLSSFPTQHSLSSFHLAVVTTKPSV 254
>TC214993 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhibitor D
precursor, complete
Length = 563
Score = 27.3 bits (59), Expect = 3.5
Identities = 23/90 (25%), Positives = 32/90 (35%)
Frame = +2
Query: 59 MNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTE 118
M K S++D Y+ ++C TRS+PP + S H +C +
Sbjct: 158 MELSFFKSDQSSSYDDDEYSKPCCDLCMCTRSMPPQCSCEDIRLNSCHSDCKS------- 316
Query: 119 EWIFYPCFFTTRLSPNQVSSSQLVMNQFCY 148
TR P Q L N FCY
Sbjct: 317 -------CMCTRSQPGQCRC--LDTNDFCY 379
>TC205748 similar to GB|AAR04550.1|37813554|AY337455 dual-specificity
phosphatase-like protein {Arabidopsis thaliana;} ,
partial (77%)
Length = 1419
Score = 27.3 bits (59), Expect = 3.5
Identities = 23/99 (23%), Positives = 39/99 (39%)
Frame = +3
Query: 28 SVHLVSYESNSEEPYLFEGLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYL 87
S+HLV + +L L++I++ W +L + + N S +VC +
Sbjct: 945 SLHLV--QVRLRRMWLEIPLMQIQQIQMALISKWMVLDPSPATCVIKLLDNLSCGKVCKV 1118
Query: 88 TRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCF 126
P H S G T S+ + +F+PCF
Sbjct: 1119 ---------PPPGCHISSQGAIITSDLSILHQLLFFPCF 1208
>TC218524
Length = 1324
Score = 26.9 bits (58), Expect = 4.6
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 2/37 (5%)
Frame = -3
Query: 62 FLLKYQGCS--NFDLFIYNDSIEEVCYLTRSLPPSIT 96
F+ KY C ++ L I + +IEE+ LT S PS+T
Sbjct: 395 FICKYFSCGMCSYFLLIMSIAIEEISILTMSS*PSLT 285
>TC221923 similar to UP|Q8GCY1 (Q8GCY1) Dihydrolipoamide succinyltransferase,
partial (5%)
Length = 789
Score = 26.6 bits (57), Expect = 6.0
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = -3
Query: 87 LTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEW 120
L RSL PS P HS ++ SK + +EW
Sbjct: 694 LIRSLKPSKLPCVHSQEGYYIC*PNNSKGLPKEW 593
>TC205208 homologue to UP|Q84P31 (Q84P31) Aldehyde dehydrogenase family 7
member A1 , partial (95%)
Length = 1900
Score = 26.6 bits (57), Expect = 6.0
Identities = 27/78 (34%), Positives = 33/78 (41%), Gaps = 3/78 (3%)
Frame = -1
Query: 55 SLQGMNWFLLKYQGCSNFDLFI-YNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGS 113
S+QG N L QG NFDLF+ N + +C T S T T +S H S
Sbjct: 1072 SMQGPN*SPL-LQGIPNFDLFVDSNKLV*YICVNTFMQKQSTTSCTPLTSSTHSSKQNRS 896
Query: 114 KSVTEEWIF--YPCFFTT 129
S + I Y C TT
Sbjct: 895 YSQFDVCIIHDYYCIVTT 842
>TC229119
Length = 455
Score = 26.2 bits (56), Expect = 7.9
Identities = 16/43 (37%), Positives = 27/43 (62%), Gaps = 1/43 (2%)
Frame = +2
Query: 45 EGLLEIRKFYSLQGMNWFLL-KYQGCSNFDLFIYNDSIEEVCY 86
+G + + YS+ G+ +FL+ Y+G S+F + I+N S EV Y
Sbjct: 11 DGWQDFVQHYSI-GVGYFLVFMYEGNSSFIVHIFNLSTSEVNY 136
>BM527184 similar to PIR|T50692|T50 proline transport protein 1 [imported] -
Arabidopsis thaliana, partial (31%)
Length = 421
Score = 26.2 bits (56), Expect = 7.9
Identities = 17/58 (29%), Positives = 28/58 (47%), Gaps = 4/58 (6%)
Frame = +2
Query: 18 LQWKLVDPFGSVHLVSYESNSEEPYLFEG----LLEIRKFYSLQGMNWFLLKYQGCSN 71
+ W+ +P GS +S S +P++ G ++ K +L +WFL Y CSN
Sbjct: 191 IPWRFHEPHGSYKHISPYIYSCKPHVP*GKEGQTKQLTKALAL-AQHWFLFHYVSCSN 361
>TC230877 similar to UP|Q9FHE1 (Q9FHE1) Glutathione transferase-like, partial
(5%)
Length = 785
Score = 26.2 bits (56), Expect = 7.9
Identities = 10/33 (30%), Positives = 15/33 (45%)
Frame = +2
Query: 117 TEEWIFYPCFFTTRLSPNQVSSSQLVMNQFCYV 149
T W+ + C +T L + L N FCY+
Sbjct: 482 TLSWVSFCCIYTNLLFSADIHFCCLPFNTFCYI 580
>TC227938 homologue to UP|Q9LK77 (Q9LK77) Similarity to acyl-CoA
thioesterase, partial (16%)
Length = 542
Score = 26.2 bits (56), Expect = 7.9
Identities = 7/20 (35%), Positives = 11/20 (55%)
Frame = -1
Query: 104 SFHGECTTGSKSVTEEWIFY 123
S HG C G + + W+F+
Sbjct: 68 SIHGNCVVGRRGIQIHWVFF 9
>TC235093 similar to UP|Q8MZ21 (Q8MZ21) RE13473p, partial (8%)
Length = 967
Score = 26.2 bits (56), Expect = 7.9
Identities = 10/15 (66%), Positives = 10/15 (66%)
Frame = -3
Query: 92 PPSITPTTHSHTSFH 106
PPS TP TH H S H
Sbjct: 683 PPSPTPQTHKHKSAH 639
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.136 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,547,876
Number of Sequences: 63676
Number of extensions: 156497
Number of successful extensions: 988
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 985
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 988
length of query: 168
length of database: 12,639,632
effective HSP length: 90
effective length of query: 78
effective length of database: 6,908,792
effective search space: 538885776
effective search space used: 538885776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0037a.10


