

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036c.6
(70 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
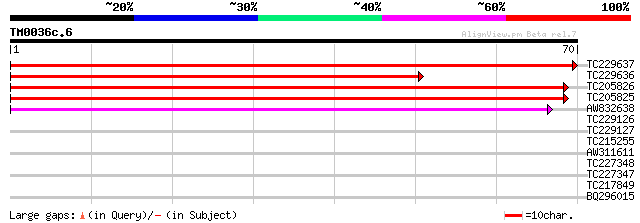
TC229637 similar to UP|KIWI_ARATH (O65154) RNA polymerase II tra... 133 2e-32
TC229636 similar to UP|KIWI_ARATH (O65154) RNA polymerase II tra... 97 2e-21
TC205826 similar to UP|KELP_ARATH (O65155) RNA polymerase II tra... 74 1e-14
TC205825 similar to UP|KELP_ARATH (O65155) RNA polymerase II tra... 74 1e-14
AW832638 weakly similar to GP|13872930|dbj P0684B02.23 {Oryza sa... 42 8e-05
TC229126 30 0.24
TC229127 30 0.31
TC215255 similar to UP|Q39087 (Q39087) DNA-binding protein, part... 28 1.2
AW311611 26 3.4
TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protei... 26 3.4
TC227347 similar to UP|O04097 (O04097) BRCA1-associated RING dom... 26 3.4
TC217849 weakly similar to UP|Q7Y0Q8 (Q7Y0Q8) Rac GDP-dissociati... 25 9.9
BQ296015 25 9.9
>TC229637 similar to UP|KIWI_ARATH (O65154) RNA polymerase II transcriptional
coactivator KIWI, partial (55%)
Length = 743
Score = 133 bits (335), Expect = 2e-32
Identities = 59/70 (84%), Positives = 68/70 (96%)
Frame = +1
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
DPDS+ +CEISKNRRV+VRNW+G I+VDIREFYVKDGKQ+PG+KGISLTMDQWNVLRNH+
Sbjct: 304 DPDSVTICEISKNRRVAVRNWKGSIMVDIREFYVKDGKQLPGRKGISLTMDQWNVLRNHV 483
Query: 61 EEIDKAVNES 70
EEIDKAVNE+
Sbjct: 484 EEIDKAVNEN 513
>TC229636 similar to UP|KIWI_ARATH (O65154) RNA polymerase II transcriptional
coactivator KIWI, partial (43%)
Length = 467
Score = 97.1 bits (240), Expect = 2e-21
Identities = 44/51 (86%), Positives = 49/51 (95%)
Frame = +3
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMD 51
DPDSI VCEISKNRRV+VRNW+G I+VDIREFYVKDGKQ+PG+KGISLTMD
Sbjct: 315 DPDSITVCEISKNRRVAVRNWKGSIMVDIREFYVKDGKQLPGRKGISLTMD 467
>TC205826 similar to UP|KELP_ARATH (O65155) RNA polymerase II transcriptional
coactivator KELP, partial (46%)
Length = 865
Score = 74.3 bits (181), Expect = 1e-14
Identities = 28/69 (40%), Positives = 51/69 (73%)
Frame = +1
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
D +++C +S RRV++++++G+ +V IRE+Y KDGK++P KGISLT +QW+ + ++
Sbjct: 358 DEGDLIICRLSDKRRVTIQDFRGKTLVSIREYYKKDGKELPTSKGISLTEEQWSTFKKNV 537
Query: 61 EEIDKAVNE 69
I+KA+ +
Sbjct: 538 PAIEKAIKK 564
>TC205825 similar to UP|KELP_ARATH (O65155) RNA polymerase II transcriptional
coactivator KELP, partial (46%)
Length = 771
Score = 74.3 bits (181), Expect = 1e-14
Identities = 28/69 (40%), Positives = 51/69 (73%)
Frame = +3
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
D +++C +S RRV++++++G+ +V IRE+Y KDGK++P KGISLT +QW+ + ++
Sbjct: 342 DEGDLIICRLSDKRRVTIQDFRGKTLVSIREYYKKDGKELPTSKGISLTEEQWSAFKKNV 521
Query: 61 EEIDKAVNE 69
I+KA+ +
Sbjct: 522 PAIEKAIKK 548
>AW832638 weakly similar to GP|13872930|dbj P0684B02.23 {Oryza sativa
(japonica cultivar-group)}, partial (12%)
Length = 271
Score = 41.6 bits (96), Expect = 8e-05
Identities = 21/67 (31%), Positives = 35/67 (51%)
Frame = +2
Query: 1 DPDSIVVCEISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRNHI 60
D +++C S R + + G V I Y DG +P KGISLT +QW+ + ++
Sbjct: 56 DECDLIICMRSYKTRATKHEYIGITFVSIL*CYN*DGTDLPTYKGISLTEEQWSPFKKNV 235
Query: 61 EEIDKAV 67
I++A+
Sbjct: 236 PAIEQAI 256
>TC229126
Length = 529
Score = 30.0 bits (66), Expect = 0.24
Identities = 13/23 (56%), Positives = 18/23 (77%)
Frame = +3
Query: 47 SLTMDQWNVLRNHIEEIDKAVNE 69
SL + NV+R H+EEIDKA++E
Sbjct: 174 SLIRFRSNVVRGHVEEIDKAISE 242
>TC229127
Length = 1013
Score = 29.6 bits (65), Expect = 0.31
Identities = 11/16 (68%), Positives = 15/16 (93%)
Frame = +2
Query: 54 NVLRNHIEEIDKAVNE 69
NV+R H+EEIDKA++E
Sbjct: 737 NVVRGHVEEIDKAISE 784
>TC215255 similar to UP|Q39087 (Q39087) DNA-binding protein, partial (67%)
Length = 732
Score = 27.7 bits (60), Expect = 1.2
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -3
Query: 12 KNRRVSVRNWQGRIVVDIREFYVKDGK 38
+NRR ++NWQ I++DIR K K
Sbjct: 676 QNRRNHIQNWQLEIILDIRPKMQKKSK 596
>AW311611
Length = 272
Score = 26.2 bits (56), Expect = 3.4
Identities = 10/13 (76%), Positives = 12/13 (91%)
Frame = +3
Query: 54 NVLRNHIEEIDKA 66
NV+R H+EEIDKA
Sbjct: 27 NVVRGHVEEIDKA 65
>TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protein (GCP1),
partial (29%)
Length = 1197
Score = 26.2 bits (56), Expect = 3.4
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +3
Query: 9 EISKNRRVSVRNWQGRIVVDIREFY 33
+I +N VS+R WQ V++RE Y
Sbjct: 948 DIPENPSVSIRIWQAVQAVNVREVY 1022
>TC227347 similar to UP|O04097 (O04097) BRCA1-associated RING domain protein
isolog; 106935-111081, partial (17%)
Length = 982
Score = 26.2 bits (56), Expect = 3.4
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +1
Query: 9 EISKNRRVSVRNWQGRIVVDIREFY 33
+I +N VS+R WQ V++RE Y
Sbjct: 292 DIPENPSVSIRIWQAVQAVNVREVY 366
>TC217849 weakly similar to UP|Q7Y0Q8 (Q7Y0Q8) Rac GDP-dissociation inhibitor
1, partial (66%)
Length = 1067
Score = 24.6 bits (52), Expect = 9.9
Identities = 18/61 (29%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Frame = -1
Query: 1 DPDSIVVCEISKNRRVS--VRNWQGRIVVDIREFYVKDGKQMPGKKGISLTMDQWNVLRN 58
+P +CE S +R+ + V++W F+ K G+ + +SL DQWN R
Sbjct: 203 NPPKKRMCEPSSSRKKTPRVQSWLVTGAKFRTFFFAKLGRIVCFVVFVSLGNDQWN*YRR 24
Query: 59 H 59
H
Sbjct: 23 H 21
>BQ296015
Length = 423
Score = 24.6 bits (52), Expect = 9.9
Identities = 18/64 (28%), Positives = 33/64 (51%), Gaps = 5/64 (7%)
Frame = +1
Query: 10 ISKNRRVSVRNWQGRIVVDIREFYVKDGKQMPGKKG-ISLTMDQWNVLRNHIEEI----D 64
I +NRR+ +++ ++V I Y+K K M ++ + L LR IE++ +
Sbjct: 94 IERNRRIHMKSLCFKLVSTIPSNYLKTSKDMLSQQDQLHLAATYIKHLRERIEKLKGEKE 273
Query: 65 KAVN 68
KA+N
Sbjct: 274 KAMN 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,899,700
Number of Sequences: 63676
Number of extensions: 27481
Number of successful extensions: 140
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 140
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 140
length of query: 70
length of database: 12,639,632
effective HSP length: 46
effective length of query: 24
effective length of database: 9,710,536
effective search space: 233052864
effective search space used: 233052864
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0036c.6


